Abstract
Salmonella enterica subspecies I, serovar Typhimurium (S. typhimurium), is a leading cause of human gastroenteritis, and is used as a mouse model of human typhoid fever1. The incidence of non-typhoid salmonellosis is increasing worldwide2,3,4, causing millions of infections and many deaths in the human population each year. Here we sequenced the 4,857-kilobase (kb) chromosome and 94-kb virulence plasmid of S. typhimurium strain LT2. The distribution of close homologues of S. typhimurium LT2 genes in eight related enterobacteria was determined using previously completed genomes of three related bacteria, sample sequencing of both S. enterica serovar Paratyphi A (S. paratyphi A) and Klebsiella pneumoniae, and hybridization of three unsequenced genomes to a microarray of S. typhimurium LT2 genes. Lateral transfer of genes is frequent, with 11% of the S. typhimurium LT2 genes missing from S. enterica serovar Typhi (S. typhi), and 29% missing from Escherichia coli K12. The 352 gene homologues of S. typhimurium LT2 confined to subspecies I of S. enterica—containing most mammalian and bird pathogens5—are useful for studies of epidemiology, host specificity and pathogenesis. Most of these homologues were previously unknown, and 50 may be exported to the periplasm or outer membrane, rendering them accessible as therapeutic or vaccine targets.
Similar content being viewed by others
Main
The genus Salmonella comprises two species: S. enterica, which is subdivided into over 2,000 serovars, and Salmonella bongori. Some serovars of S. enterica, such as S. typhi, cause systemic infections and typhoid fever, whereas others, such as S. typhimurium, cause gastroenteritis. Some serovars, such as S. typhi, are host specialists that infect only humans, whereas others such as S. typhimurium, are host generalists that occur in humans and many other mammalian species. Domestic animals act as a reservoir for the food-borne spread of host-generalist serovars, which accounts for the high incidence of non-typhoid Salmonella infections worldwide. Estimated costs of food-borne diseases in the United States (with salmonellosis a major component) range from 4.8 to 23 billion dollars3.
Salmonella typhimurium strain LT2, the principal strain for cellular and molecular biology in Salmonella, was isolated in the 1940s and used in the first studies on phage-mediated transduction1. Attenuated mutants of S. enterica may be used as live oral vaccines against Salmonella infection, to express antigens from other pathogens, and to deliver proteins to solid tumours6.
The general characteristics of the S. typhimurium LT2 genome are summarized in Table 1. The full dataset is presented as Supplementary Information and is available also at http://genome.wustl.edu/gsc/Projects/S.typhimurium.
The overall similarity of S. typhimurium LT2 to eight other enterobacterial genomes is summarized in Table 2. We compared S. typhimurium LT2 to three other fully sequenced genomes: S. typhi7, the principal cause of typhoid in humans; E. coli K12 (ref. 8), a non-pathogen; and E. coli O157:H7, an enterohaemorrhagic strain9. (Escherichia coli is a member of the closest known genus to Salmonella.) We sequenced genome samples over 97% coverage from S. enterica serovar S. paratyphi A, a frequent cause of typhoid, and K. pneumoniae, an opportunistic human pathogen closely related to Salmonella. Visual representations of the comparisons of S. typhimurium LT2 to the sequenced and sampled genomes are available at both gene and sequence resolution using STM-Enteric and STM-Menteric web tools (http://galapagos.cse.psu.edu/enterix)10,11. A total of 4,330 complete open reading frames (ORFs) from the S. typhimurium LT2 genome were amplified using specific PCR primers with appended, common 5′ ends. These ORFs were microarrayed on glass and probed with fluorescently labelled genomic DNA12 from S. bongori and from S. enterica serovars S. paratyphi A, S. paratyphi B and S. arizonae, to determine the presence of homologues of S. typhimurium LT2 genes. The microarray data for S. paratyphi A was over 98% concordant with data from the S. paratyphi A genomic sequence (see Supplementary Information). Salmonella typhimurium LT2, S. typhi, S. paratyphi A and S. paratyphi B are all in subspecies I of S. enterica, which colonizes mammals and birds and causes 99% of Salmonella infections in humans5,13. Salmonella arizonae is in subspecies IIIa, which colonizes reptiles and also, rarely, humans; S. bongori, the only other species of Salmonella, does not cause disease in mammals.
Genomic comparisons among the four completed genomes (S. typhimurium LT2, S. typhi, E. coli K12 and E. coli O157:H7) reveal that they are collinear for most genes except for inversions over the terminus of replication (TER)9,14. Salmonella typhimurium LT2 has a 588-kb inversion compared with E. coli K12. Rearrangements between the rrn operons are common in S. typhi 7,15, but are not found in E. coli K12, E. coli O157:H7 or S. typhimurium LT2, or in most other Salmonella strains. Large, stable duplications are rare in enterobacterial genomes other than structural RNA genes and transposable elements. In S. typhimurium LT2 the largest duplicated region of coding sequences (CDS) is the cytochrome c biogenesis locus (ccm, 7.5 kb). The two copies are 99% identical at the DNA level and 100% identical for amino acids.
The chromosomes of enteric bacteria are mosaics, composed of collinear regions interspersed with ‘loops’ or ‘islands’ unique to certain species; the islands sometimes encode pathogenicity functions (called Salmonella pathogenicity islands, SPIs)9,16. Of the 4,489 CDS and pseudogenes annotated in the S. typhimurium LT2 chromosome, at least 2,466 (55%) have a close homologue in all eight other enterobacterial genomes that we examined. These homologous genes (shown in red in Fig. 1) along with structural RNAs constitute 2.5 Mb of the S. typhimurium LT2 genome.
a, The chromosome. Base pairs are indicated outside the outer circle. The outer two circles represent the coding orientation, with the forward strand on the outside and the reverse strand on the inside. Red indicates close homologues in all eight genomes. Green indicates genes with a close homologue in at least one other Salmonella (S. typhi, S. paratyphi A, S. paratyphi B, S. arizonae or S. bongori) but not in E. coli K12, E. coli O157:H7 and K. pneumoniae. Blue indicates genes present only in S. typhimurium LT2. Grey indicates other combinations. The black inner circle is the G+C content; the purple/yellow innermost circle is the GC bias. The positions of the origin of replication (ORI) and terminus (TER) are shown. b, The plasmid pSLT. Base pairs are indicated outside the outer circle. The plasmid is not to scale. The colouring scheme is the same as for a.
Table 3 summarizes the distribution of the larger islands among the eight genomes, on the basis of sequence and microarray analysis. SPIs 1–5, previously described in S. typhimurium (see refs 1, 17), are listed along with many newly detected large islands and the subset of smaller islands that encode fimbriae or potential virulence factors. Fifteen of the islands are adjacent to a transfer RNA. In fact, almost half of the individually encoded tRNAs are adjacent to an island. At least seven islands encode integrase-like proteins, which are often found in islands16. Salmonella typhimurium LT2 contains four functional prophages: Gifsy-1 and -2 (known to have a role in infection18,19) and Fels-1 and -2 (ref. 12). The insertion site of these prophages was predicted from the sequence and was confirmed by microarray-based comparison of labelled DNA from strains that were cured of prophage. These phage are not present in the other eight genomes, although homologues of a few genes exist, presumably in related phage. A previously unknown phage, or phage remnant, that included the S. typhimurium LT2 genes STM4201 to STM4219 was detected in S. typhimurium LT2 by homology to other phage.
Most strains of S. typhimurium contain a plasmid of about 90 kb. The plasmid of strain LT2 is called pSLT20. Out of 108 annotated CDS and pseudogenes in pSLT, only three have a close homologue in S. typhi, S. paratyphi A or S. paratyphi B, as expected owing to these strains lacking the plasmid. A search through GenBank revealed 50 pSLT genes that have a close homologue in plasmids from other Salmonella serovars. Many homologues of genes in the tra operon of the F-factor of E. coli K12 were identified, which presumably are responsible for the self-transmissibility of pSLT at rates of up to 3 × 10-4 (ref. 21). The copy number of the pSLT plasmid was estimated as 2.75, using the relative sequence coverage of the plasmid versus the chromosome in the shotgun phase of sequencing, and estimated as 1.4–3.1 under a variety of growth conditions, when measured by average signal intensity on microarrays (data not shown).
Fimbriae on the cell surface mediate adhesion to host cells22. Salmonella typhimurium LT2 contains 12 putative operons of the chaperone–usher assembly class: stc (called yehABCD in E. coli K12), bcf, fim, lpf, saf, stb, std, stf, sth, sti, stj, all of which are located on the chromosome, and pef, which is located on the plasmid. Operons bcf, fim, lpf and pef were reported earlier for S. typhimurium LT2 and shown to be functional; saf, stb, std and sth were detected only by hybridization22. The sti and stj operons were previously undetected; thus, six of the operons were not previously sequenced and two were not previously detected. Salmonella typhimurium LT2 also has the csg operon (originally called agf) for curli fimbriae, from the nucleator-dependent assembly pathway, but genes for type IV fimbriae7 are not detected. Table 3 describes the taxonomic distribution of close homologues of the fimbriae gene clusters in the other eight enterobacterial genomes, but does not include information on the presence of more divergent homologues or orthologues in the other genomes.
Complete sequencing of many closely related genomes, such as E. coli K12, E. coli O157:H7, S. typhi and S. typhimurium LT2 (see Table 2), aids the detection of pseudogenes, because a frameshift or stop codon is recognizable only if the gene is collinear with a functional, homologous gene in another genome. This allowed the detection of at least 204 pseudogenes in S. typhi7, whereas the S. typhimurium LT2 chromosome has only about 39. The large number of pseudogenes in S. typhi may contribute to or be a consequence of the restriction of S. typhi to growth in humans alone7, whereas S. typhimurium LT2, with a broad host range, has far fewer pseudogenes. Pseudogenes may be unrecognized when a close, intact homologue is unavailable, as is true for 11% or more of the S. typhimurium LT2 and S. typhi genomes (Table 2). Other potential pseudogenes are encoded across insertion/deletion events that distinguish S. typhimurium LT2 and S. typhi, such as S. typhimurium LT2 gene STM0098, which is split by an insertion of about 2,000 bp in S. typhi and S. paratyphi (or a deletion of this size in S. typhimurium LT2). Such genes have been annotated as insertion/deletion events rather than pseudogenes (see Supplementary Information). Finally, pseudogenes might actually have at least one functional fragment. Nevertheless, these caveats do not alter the conclusion that S. typhimurium LT2 has far fewer pseudogenes than its close relative S. typhi.
The consequences of loss of function in pseudogenes in S. typhimurium LT2 is usually unclear, as the function of the known, intact homologue in another organism is often unknown. However, there are pseudogenes in S. typhimurium LT2 for maltose regulation (malXY23) and for trehalose metabolism (treB), where the normal allele can substitute in the maltose pathway24; thus, S. typhimurium LT2 has disrupted maltose pathways with multiple, independent mutations. Histidine is not used as a carbon source in LT2, although all of the genes are present; this may be explained by a pseudogene mutation in hutU. Some of the pseudogenes are in potentially redundant systems: the dcoA pseudogene shows over 98% amino-acid identity to another putative sodium ion pump, oxaloacetate decarboxylase alpha chain (STM3352); cutF (STM0241), which encodes the copper homeostasis protein, is a pseudogene, but cutC (which remains intact) is sufficient for the same function25.
Genes found only in Salmonella (see Table 3 for examples) may have been recruited since the divergence from Escherichia coli and Klebsiella, or they may have been lost from these genera. There are 1,106 S. typhimurium LT2 CDS in this class (marked in green in Fig. 1) that have a close homologue in at least one of the other five Salmonella examined. Many S. typhimurium LT2 genes associated with pathogenesis are in this class, including invasion genes such as inv and prg; proteins exported by the type III secretion system, such as SopB, SopD, SopE2, SipA, SipB, SipC and SptP; and some secretory system genes, such as members of ssa and sse. Most of these genes have homologues in S. bongori, indicating that these characteristics are maintained by even the most divergent of Salmonella. A subset of 352 S. typhimurium LT2 CDS (8%) have close homologues in one or more of the other three subspecies I genomes (S. typhi, S. paratyphi A and S. paratyphi B) but not in S. arizonae, S. bongori, E. coli K12, E. coli O157:H7 or K. pneumoniae. These genes, most of which were unknown previously, may include genes for specialization of subspecies I to warm-blooded hosts. Among these 352 S. typhimurium LT2 CDS are bigA, envF, shdA, sifAB, sinH, srfJ, srgAB (homologues of ail and clpB), and some genes in the fimbrial clusters, saf, stb, stc, std, stf and sti. Only 70 genes (20%) are named, whereas 49% of all CDS are named, indicating that the group has remained largely unstudied. There are 121 S. typhimurium LT2 CDS that have no close homologues in any of the other eight genomes, only a few of which are named, including individual genes from the pef and stj fimbriae, spvR, and a homologue of mvpA.
CDS rich in A+T are almost threefold over-represented among genes that have no close homologues outside subspecies I, confirming previous observations made with a smaller set of such genes26. It is not obvious why CDS confined to only some Salmonella should so often be A+T-rich. There is no such bias among G+C-rich genes. If some of these genes are of recent foreign origin then perhaps there is a preferential A+T-rich source for recruited genes.
Attenuated S. typhimurium has been used both as a vaccine and for expression of heterologous proteins for vaccines against other organisms27. Proteins that are predicted to be exported to the periplasm or outer membrane, or beyond, are of interest for vaccines and therapy, as they would be exposed for targeting. PSORT28, which predicts cellular location from protein sequence, predicts 251 outer-membrane proteins and 347 periplasmic proteins in S. typhimurium LT2, 182 of which are missing from E. coli K12, E. coli O157:H7 or K. pneumoniae, and about 50 of which are confined to subspecies I.
Antigens from all of the predicted surface and secreted CDS can be cloned, expressed on cells, and tested for immunological response, as was done recently for CDS conserved among Neisseria meningitidis29. As a first step, we have amplified by PCR almost all the S. typhimurium LT2 CDS in a manner suitable for expression, and the distribution of close homologues among eight other enterobacterial genomes has been determined for each gene.
Methods
Genome sequencing
The complete 4.8-Mb genome of S. enterica serovar Typhimurium strain LT2 (American Type Culture Collection, ATCC, number 700720) was sequenced using a combination of two approaches. The first approach was fivefold, whole-genome shotgun sampling, and the second was three- to tenfold sampling of gel-purified restriction fragments ranging in size from 43.3 to 570 kb14. Sonicated and size-fractionated DNA (approximately 1.5 kb) was cloned into M13 and plasmid vectors. Subclones were sequenced using dye-primer and dye-terminator chemistry on ABI 377 and 3700 sequencing machines. We assembled 92,648 sequence reads, representing ∼7.7-fold final coverage, using the PHRAP assembly program (P. Green, unpublished data). The assembled genome sequence is in agreement with the physical map of the S. typhimurium LT2 genome14, with reads from both ends of about 372 lambda DASH-II clones with inserts of 13–20 kb, and with a S. typhimurium LT2 bacterial artificial chromosome library (prepared by R. Wing) restriction fingerprinted at the Genome Sequencing Center. Two other genomes were partially sequenced using whole-genome, shotgun sample sequencing: 27,670 sequence reads of S. enterica serovar Paratyphi A strain RKS4993 (ATCC 9510) yielded a 3.7-fold coverage (97.5%) for this ∼4.6 Mb genome, having 894 contigs with a predicted average gap size of ∼200 bp; 36,848 sequence reads of K. pneumoniae strain MGH78578 (ATCC 700721), a clinical isolate from sputum, yielded a 3.9-fold depth of coverage (98%) for this ∼5.6 Mb genome, resulting in 711 contigs with a predicted average gap size of ∼160 bp.
Annotation
The primary annotation database was Acedb (http://www.acedb.org). Protein gene predictions were performed using GeneMark (http://opal.biology.gatech.edu/GeneMark) and Glimmer (http://www.tigr.org/softlab/glimmer/glimmer.html). Predicted proteins were searched against the protein family database, Pfam 5.5 (http://pfam.wustl.edu), against the COG database to find putative orthologues in other completed genomes30, and against the protein localization prediction software, PSORT28. The annotation was compared with the S. typhi annotation from the Sanger Centre and all differences in CDS predictions and start predictions were reassessed, on the basis of choice of start codon, length, conservation in Enterobacteriaceae and presence of identifiable motifs. The details of microarray construction and hybridization with genomic probes is described elsewhere12. Strains, lambda and BAC clones are available from the Salmonella Genetic Stock Centre, http://www.ucalgary.ca/~kesander.
References
Neidhardt, F. C. (ed. in chief) Escherichia coli and Salmonella: Cellular and Molecular Biology (ASM, Washington DC, 1996).
Chalker, R. B. & Blaser, M. J. A review of human salmonellosis: III. Magnitude of Salmonella infection in the United States. Rev. Infect. Dis. 10, 111–124 (1988).
Todd, E. Epidemiology of foodborne illness: North America. Lancet 336, 788–790 (1990).
Cooke, E. M. Epidemiology of foodborne illness: UK. Lancet 336, 790–793 (1990).
Selander, R. K., Li, J. & Nelson, K. in Escherichia coli and Salmonella: Cellular and Molecular Biology (eds Neidhardt, F. C. et al.) 2691–2707 (ASM, Washington DC, 1996).
Bermudes, D., Low, B. & Pawelek, J. Tumor-targeted Salmonella. Highly selective delivery vectors. Adv. Exp. Med. Biol. 465, 57–63 (2000).
Parkhill, J. et al. Complete genome sequence of a multiple drug resistant Salmonella enterica serovar Typhi CT18. Nature 413, 848–852 (2001).
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1474 (1997).
Perna, N. T. et al. Genome sequence of enterohaemorrhagic Escherichia coli O157:H7. Nature 409, 529–533 (2001).
Florea, L. et al. Web-based visualization tools for bacterial genome alignments. Nucleic Acids Res. 28, 3486–3496 (2000).
McClelland, M. et al. Comparison of the Escherichia coli K-12 genome with sampled genomes of a Klebsiella pneumoniae and three Salmonella enterica serovars, Typhimurium, Typhi and Paratyphi. Nucleic Acids Res. 28, 4974–4986 (2000).
Porwollik, S. et al. The Δ-uvrB mutations in the Ames strains of Salmonella span 15 to 119 genes. Mut. Res. 483, 1–11 (2001).
Popoff, M. Y., Bockemuhl, J. & Brenner, F. W. Supplement 1999 (no. 43) to the Kauffmann-White scheme. Res. Microbiol. 151, 893–896 (2000).
Liu, S. L., Hessel, A. & Sanderson, K. E. The XbaI-BlnI-CeuI genomic cleavage map of Salmonella typhimurium LT2 determined by double digestion, end labelling, and pulsed-field gel electrophoresis. J. Bacteriol. 175, 4104–4120 (1993).
Liu, S. L. & Sanderson, K. E. Highly plastic chromosomal organization in Salmonella typhi. Proc. Natl Acad. Sci. USA 93, 10303–10308 (1996).
Ochman, H., Lawrence, J. G. & Groisman, E. A. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000).
Blanc-Potard, A. B., Solomon, F., Kayser, J. & Groisman, E. A. The SPI-3 pathogenicity island of Salmonella enterica. J. Bacteriol. 181, 998–1004 (1999).
Stanley, T. L., Ellermeier, C. D. & Slauch, J. M. Tissue-specific gene expression identifies a gene in the lysogenic phage Gifsy-1 that affects Salmonella enterica serovar Typhimurium survival in Peyer's patches. J. Bacteriol. 182, 4406–4413 (2000).
Figueroa-Bossi, N., Uzzau, S., Maloriol, D. & Bossi, L. Variable assortment of prophages provides a transferable repertoire of pathogenic determinants in Salmonella. Mol. Microbiol. 39, 260–272 (2001).
Matsui, H. et al. Virulence plasmid borne spvB and spvC genes can replace the 90-kilobase plasmid in conferring virulence to Salmonella enterica serovar Typhimurium in subcutaneously inoculated mice. J. Bacteriol. 183, 4652–4658 (2001).
Ahmer, B. M., Tran, M. & Heffron, F. The virulence plasmid of Salmonella typhimurium is self-transmissible. J. Bacteriol. 181, 1364–1368 (1999).
Townsend, S. M. et al. Salmonella enterica serovar Typhi possesses a unique repertoire of fimbrial gene sequences. Infect. Immun. 69, 2894–2901 (2001).
Reidl, J. & Boos, W. The malX malY operon of Escherichia coli encodes a novel enzyme II of the phosphotransferase system recognizing glucose and maltose and an enzyme abolishing the endogenous induction of the maltose system. J. Bacteriol. 173, 4862–4876 (1991).
Decker, K., Gerhardt, F. & Boos, W. The role of the trehalose system in regulating the maltose regulon of Escherichia coli. Mol. Microbiol. 32, 777–788 (1999).
Gupta, S. D., Lee, B. T., Camakaris, J. & Wu, H. C. Identification of cutC and cutF (nlpE) genes involved in copper tolerance in Escherichia coli. J. Bacteriol. 177, 4207–4215 (1995).
Lan, R. & Reeves, P. R. Gene transfer is a major factor in bacterial evolution. Mol. Biol. Evol. 13, 47–55 (1996).
Bumann, D., Hueck, C., Aebischer, T. & Meyer, T. F. Recombinant live Salmonella spp. for human vaccination against heterologous pathogens. FEMS Immunol. Med. Microbiol. 27, 357–364 (2000).
Nakai, K. & Horton, P. PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization. Trends Biochem. Sci. 24, 34–36 (1999).
Pizza, M. et al. Identification of vaccine candidates against serogroup B meningococcus by whole-genome sequencing. Science 287, 1816–1820 (2000).
Tatusov, R. L. et al. The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res. 29, 22–28 (2001).
Acknowledgements
This work was supported by National Institutes of Health grants to R.W., M.M. and W.M. Nine ORFs were added to the annotation courtesy of L. Umayam and O. White of the Comprehensive Microbial Resource. We thank T. Lynch, M. Becker, G. Duckels and P. Minx for library production; J. Welsh, E. Koonin, R. Tatusov, H. Salgado, J. Collado-Vides and K. Nakai for assistance with computer analysis; S. Sims for assistance with primer design; M. Sekhon and the GSC Mapping Group for BAC mapping; S.-L. Liu for large restriction fragments of S. typhimurium LT2; A. McKay, B. Pearson, J. Roth and M. Riley for advice; and J. Parkhill and G. Dougan for sharing data from the S. typhi project.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Definition of close homologue
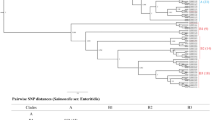
The genome sequences of STY, SPA, ECO, ECH and KPN all differ in their divergence from STM and two of these genomes are incomplete; SPA and KPN. To obtain a qualitative measure of shared gene content among these genomes a similarity threshold was devised and applied consistently to all the genomes, complete and incomplete. This threshold was determined by examining the genes that were most clearly shared by all the genomes, namely those that were syntenic symmetrical best hits in the complete genomes and which also had good homologues (>50% DNA sequence identity) in the incomplete genomes. Next, a DNA sequence threshold that would correctly identify 95% of these genes as being "present" was determined for each of the genomes. One cannot use 100% capture as the threshold because a few genes are syntenic symmetric best hits but highly divergent. The rounded identity thresholds that fit this criterion were STY, 93%; SPA, 90%; ECO and ECH, 70%; KPN, 65%. The resulting estimates of similarity are approximate. Close paralogues and some recently derived pseudogenes were also identified as "present" at these thresholds.
In the estimate of genome similarity any issues that might arise due to annotation differences between the genomes were avoided by searching the STM annotation against the unannotated genomes of the other organisms. For example, at least some of the genes annotated in STM appear to be genes that may have been missed in the ECO annotation, including STM0556, 1872, 2484, 2682, 3411, 3841, 4008, and 4562.
To compensate for missing data in the SPA and KPN samples, matches that terminated at the end of a contig in the sampled genome were accepted if they spanned more than 100 bases at one end of an STM gene [McClelland, et al., NAR 2000]. Genes that occur between aligned contigs in the reference genome (annotated in grey in Enterix, http://galapagos.cse.psu.edu/enterix/) are marked with a question mark, to indicate missing data, in the data file at the web site and in the supplement.
Figures
Rights and permissions
About this article
Cite this article
McClelland, M., Sanderson, K., Spieth, J. et al. Complete genome sequence of Salmonella enterica serovar Typhimurium LT2. Nature 413, 852–856 (2001). https://doi.org/10.1038/35101614
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35101614
This article is cited by
-
“Sharing the matrix” – a cooperative strategy for survival in Salmonella enterica serovar Typhimurium
BMC Microbiology (2023)
-
Programming bacteria for multiplexed DNA detection
Nature Communications (2023)
-
Differential regulatory control of curli (csg) gene expression in Salmonella enterica serovar Typhi requires more than a functional CsgD regulator
Scientific Reports (2023)
-
Genomic analysis of Anderson typing phages of Salmonella Typhimrium: towards understanding the basis of bacteria-phage interaction
Scientific Reports (2023)
-
The Prophylactic Protection of Salmonella Typhimurium Infection by Lentilactobacillus buchneri GX0328-6 in Mice
Probiotics and Antimicrobial Proteins (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.