Abstract
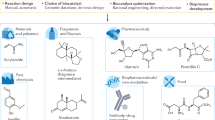
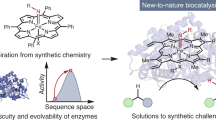
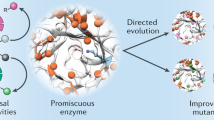
Nature provides a fantastic array of catalysts extremely well suited to supporting life, but usually not so well suited for technology. Whether biocatalysis will have a significant technological impact depends on our finding robust routes for tailoring nature's catalysts or redesigning them anew. Laboratory evolution methods are now used widely to fine-tune the selectivity and activity of enzymes. The current rapid development of these combinatorial methods promises solutions to more complex problems, including the creation of new biosynthetic pathways. Computational methods are also developing quickly. The marriage of these approaches will allow us to generate the efficient, effective catalysts needed by the pharmaceutical, food and chemicals industries and should open up new opportunities for producing energy and chemicals from renewable resources.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Adam, M. W. W. & Kelly, R. M. Finding and using hyperthermophilic enzymes. Trends Biotechnol. 16, 329– 332 (1998).
Rondon, M. R., Goodman, R. M. & Handelsman, J. The Earth's bounty: assessing and accessing soil microbial diversity. Trends Biotechnol. 17, 403– 409 (1999).
Short, J. Recombinant approaches for accessing biodiversity. Nature Biotechnol. 15, 1322–1323 ( 1997).
Bull, A. T., Ward, A. C. & Goodfellow, M. Search and discovery strategies for biotechnology: the paradigm shift. Microbiol. Mol. Biol. Rev. 64, 573–606 (2000).
Black, M. E., Newcomb, T. G., Wilson, H.-M. P. & Loeb, L. A. Creation of drug-specific Herpes Simplex Virus type 1 thymidine kinase mutants for gene therapy. Proc. Natl Acad. Sci. USA 93, 3525–3529 (1996).
Encell, L. P., Coates, M. M. & Loeb, L. A. Engineering human DNA alkyltransferases for gene therapy using random sequence mutagenesis. Cancer Res. 58, 1013–1020 (1998).
Matsumura, I., Wallingford, J. B., Surana, N. K., Vize, P. D. & Ellington, A. D. Directed evolution of the surface chemistry of the reporter enzyme β-glucuronidase. Nature Biotechnol. 17, 696–701 ( 1999).
Lanio, T., Jeltsch, A. & Pingoud, A. Towards the design of rare cutting restriction endonucleases: using directed evolution to generate variants of Eco/Rv differing in their substrate specificity by two orders of magnitude. J. Mol. Biol. 283, 59–69 ( 1998).
Liu, D. R., Magliery, T. J., Pastrnak, M. & Schultz, P. G. Engineering a tRNA and aminoacyl-tRNA synthetase for the site-specific incorporation of unnatural amino acids into proteins in vivo. Proc. Natl Acad. Sci. USA 94, 10092–10097 (1997).
Babbitt, P. New functions from old scaffolds: how nature re-engineers enzymes for new functions. Adv. Protein Chem. 55, 1– 28 (2000).
Xiang, H., Luo, L., Taylor, K. L. & Dunaway-Mariano, D. Interchange of catalytic activity within the 2-enoyl-coenzyme A hydratase/isomerase superfamily based on a common active site template. Biochemistry 38, 7638–7652 (1999).
Broun, P., Shanklin, J., Whittle, E. & Somerville, C. Catalytic plasticity of fatty acid modification enzymes underlying chemical diversity of plant lipids. Science 282, 1315–1317 (1998).
Shanklin, J. Exploring the possibilities presented by protein engineering. Curr. Opin. Plant Biol. 3, 243–248 (2000).
Shanklin, J. & Cahoon, E. B. Desaturation and related modifications of fatty acids. Annu. Rev. Plant Physiol. Plant Mol. Biol. 49, 611–641 (1998).
Wintrode, P. & Arnold, F. H. Temperature adaptation of enzymes: lessons from laboratory evolution. Adv. Protein Chem. 55, 161–226 (2000).
Jaenicke, R. Stability and stabilization of globular proteins in solution. J. Biotechnol. 79, 193–203 (2000).
Dahiyat, B. I. & Mayo, S. L. De novo protein design: fully automated sequence selection. Science 278, 82–87 (1997).
Malakaukas, S. M. & Mayo, S. L. Design, structure and stability of a hyperthermophilic protein variant. Nature Struct. Biol. 5, 470–475 ( 1998).
Cedrone, F., Ménez, A. & Quéméneur, E. Tailoring new enzyme functions by rational redesign. Curr. Opin. Struct. Biol. 10, 405–410 (2000).
Arnold, F. H. (ed.) Evolutionary Protein Design (Adv. Protein Chem. 55) (Academic, San Diego, 2000).
Affholter, J. & Arnold, F. H. Engineering a revolution. Chem. Britain 35, 48–51 (1999).
Ness, J. E., del Cardayre, S. B., Minshull, J. & Stemmer, W. P. C. Molecular breeding—the natural approach to enzyme design. Adv. Protein Chem. 55, 261–292 (2000).
Reetz, M. T. & Jaeger, K.-E. Superior biocatalysts by directed evolution. Top. Curr. Chem. 200, 31– 57 (1999).
Liebeton, K. et al. Directed evolution of an enantioselective lipase. Chem. Biol. 7, 709–718 ( 2000).
May, O., Nguyen, P. & Arnold, F. H. Inverting enantioselectivity by directed evolution of hydantoinase for improved production of l-methionine. Nature Biotechnol. 18, 317–320 (2000).
Arnold, F. H. Directed enzyme evolution 〈http://www.che.caltech.edu:80/groups/fha/Enzyme/directed.html 〉
Oue, S., Okamoto, A., Yano, T. & Kagamiyama, H. Redesigning the substrate specificity of an enzyme by cumulative effects of the mutations of non-active site residues. Biol. Chem. 274, 2344–2349 (1999).
Yano, T., Oue, S. & Kagamiyama, H. Directed evolution of an aspartate aminotransferase with new substrate specificities. Proc. Natl Acad. Sci. USA 95, 5511–5515 (1998).
Orencia, M. C. & Stevens, R. C. Structural changes in directed and immune evolved enzymes. Adv. Protein Chem. 55, 227–260 ( 2000).
Stemmer, W. P. C. Rapid evolution of a protein in vitro by DNA shuffling. Nature 370, 389–391 ( 1994).
Crameri, A., Raillard, S.-A., Bermudez, E. & Stemmer, W. P. C. DNA shuffling of a family of genes from diverse species accelerates directed evolution. Nature 391, 288– 291 (1998).
Sandmann, G., Albrecht, M., Schnurr, G., Knörzer, O. & Böger, P. The biotechnological potential and design of novel carotenoids by gene combination in Escherichia coli . Trends Biotechnol. 17, 233– 237 (2000).
Cane, D. E., Walsh, C. T. & Khosla, C. Harnessing the biosynthetic code: combinations, permutations and mutations. Science 282, 63– 68 (1998).
Schmidt-Dannert, C., Umeno, D. & Arnold, F. H. Molecular breeding of carotenoid biosynthetic pathways . Nature Biotechnol. 18, 750– 753 (2000).
Patten, P. A. et al. The immunological evolution of catalysis. Science 271, 1086–1091 ( 1996).
Barbas, C. F. III, List, B., Rader, C., Segal, D. J. & Turner, J. M. From catalytic asymmetric synthesis to the transcriptional regulation of genes: in vivo and in vitro evolution of proteins. Adv. Protein Chem. 55, 317–366 ( 2000).
Ostermeier, M. & Benkovic, S. J. Evolution of protein function by domain swapping. Adv. Protein Chem. 55, 29–78 (2000).
Sieber, V., Martinez, C. A. & Arnold, F. H. Libraries of hybrid proteins from distantly-related sequences. Nature Biotechnol. (in the press).
Riechmann, L. & Winter, G. Novel folded protein domains generated by combinatorial shuffling of polypeptide segments. Proc. Natl Acad. Sci. USA 97, 10068–10073 (2000).
Plückthun, A., Schaffitzel, C., Hanes, J. & Jermutus, L. In vitro selection and evolution of proteins. Adv. Protein Chem. 55, 367–404 ( 2000).
Roberts, R. W. Totally in vitro protein selection using mRNA-protein fusions and ribosome display. Curr. Opin. Chem. Biol. 3, 268– 273 (1999).
Olsen, M., Iverson, B. & Georgiou, G. High-throughput screening of enzyme libraries. Curr. Opin. Biotechnol. 11, 331–337 (2000).
Altamirano, M. M., Blackburn, J. M., Aguayo, C. & Fersht, A. R. Directed evolution of new catalytic activity using the α/β-barrel scaffold. Nature 403, 617– 622 (2000).
Voigt, C. A., Mayo, S. L., Arnold, F. H. & Wang, Z.-G. Computational method to reduce the search for directed protein evolution. Proc. Natl Acad. Sci. USA (in the press).
Benson, D. E., Wisz, M. S. & Hellinga, H. W. Rational design of nascent metalloenzymes. Proc. Natl Acad. Sci. USA 97, 6292– 6297 (2000).
Acknowledgements
I thank the many talented students and postdocs who have contributed to the development of new biocatalyst engineering tools in my laboratory, and the following organizations for their financial support: the US Office of Naval Research, the US National Science Foundation, the Army Research Office, Maxygen, Inc., The Biotechnology Research & Development Corporation, British Petroleum, Degussa AG and Procter & Gamble Co. I also thank C. Voigt and J. Shanklin for thoughtful comments, and J. Shanklin for Fig. 1.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Arnold, F. Combinatorial and computational challenges for biocatalyst design. Nature 409, 253–257 (2001). https://doi.org/10.1038/35051731
Issue Date:
DOI: https://doi.org/10.1038/35051731
This article is cited by
-
Autonomous Reaction Network Exploration in Homogeneous and Heterogeneous Catalysis
Topics in Catalysis (2022)
-
In silico screening and heterologous expression of soluble dimethyl sulfide monooxygenases of microbial origin in Escherichia coli
Applied Microbiology and Biotechnology (2022)
-
Biotechnological potential of psychrophilic microorganisms as the source of cold-active enzymes in food processing applications
3 Biotech (2021)
-
Non-proteinaceous hydrolase comprised of a phenylalanine metallo-supramolecular amyloid-like structure
Nature Catalysis (2019)
-
Toxicity assessment of chlorpyrifos on different organs of rat: exploitation of microbial-based enzymatic system for neutralization
Environmental Science and Pollution Research (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.