Abstract
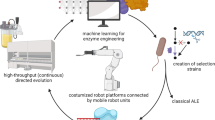
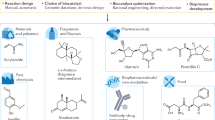
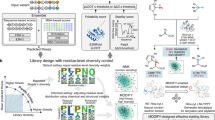
Biocatalysis, the application of enzymes to solve synthetic problems of human import, has blossomed into a powerful technology for chemical innovation. In the past decade, a threefold partnership, where nature provides blueprints for enzymatic catalysis, chemists introduce innovative activity modes with abiological substrates, and protein engineers develop new tools and algorithms to tune and improve enzymatic function, has unveiled the frontier of new-to-nature enzyme catalysis. In this Perspective, we highlight examples of interdisciplinary studies, which have helped to expand the scope of biocatalysis, including concepts of enzymatic versatility explored through the lens of biomimicry, to achieve activities and selectivities beyond those currently possible with chemocatalysis. We indicate how modern tools, such as directed evolution, computational protein design and machine learning-based protein engineering methods, have already impacted and will continue to influence enzyme engineering for new abiological transformations. A sustained collaborative effort across disciplines is anticipated to spur further advances in biocatalysis in the coming years.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Sun, H., Zhang, H., Ang, E. L. & Zhao, H. Biocatalysis for the synthesis of pharmaceuticals and pharmaceutical intermediates. Bioorg. Med. Chem. 26, 1275–1284 (2018).
Sheldon, R. A., Brady, D. & Bode, M. L. The Hitchhiker’s guide to biocatalysis: recent advances in the use of enzymes in organic synthesis. Chem. Sci. 11, 2587–2605 (2020).
Winkler, C. K., Schrittwieser, J. H. & Kroutil, W. Power of biocatalysis for organic synthesis. ACS Cent. Sci. 7, 55–71 (2021).
Renata, H., Wang, Z. J. & Arnold, F. H. Expanding the enzyme universe: accessing non-natural reactions by mechanism-guided directed evolution. Angew. Chem. Int. Ed. 54, 3351–3367 (2015).
Chen, K. & Arnold, F. H. Engineering new catalytic activities in enzymes. Nat. Catal. 3, 203–213 (2020).
Breslow, R. Biomimetic chemistry: biology as an inspiration. J. Biol. Chem. 284, 1337–1342 (2009).
Swiegers, G. F. (ed.) Bioinspiration and Biomimicry in Chemistry: Reverse-Engineering Nature (Wiley, 2012)
Flanigan, D. M., Romanov-Michailidis, F., White, N. A. & Rovis, T. Organocatalytic reactions enabled by N-heterocyclic carbenes. Chem. Rev. 115, 9307–9387 (2015).
Miller, D. C., Tarantino, K. T. & Knowles, R. R. Proton-coupled electron transfer in organic synthesis: fundamentals, applications, and opportunities. Top. Curr. Chem. 374, 30 (2016).
Breslow, R. & Dong, S. D. Biomimetic reactions catalyzed by cyclodextrins and their derivatives. Chem. Rev. 98, 1997–2012 (1998).
Prier, C. K. & Arnold, F. H. Chemomimetic biocatalysis: exploiting the synthetic potential of cofactor-dependent enzymes to create new catalysts. J. Am. Chem. Soc. 137, 13992–14006 (2015).
Huang, X. & Groves, J. T. Beyond ferryl-mediated hydroxylation: 40 years of the rebound mechanism and C–H activation. J. Biol. Inorg. Chem. 22, 185–207 (2017).
The Porphyrin Handbook: Bioinorganic and Bioorganic Chemistry Vol. 11 (eds Kadish, K. M. et al.) (Academic, 2003).
Mansuy, D. A brief history of the contribution of metalloporphyrin models to cytochrome P450 chemistry and oxidation catalysis. C. R. Chim. 10, 392–413 (2007).
Breslow, R. & Gellman, S. H. Tosylamidation of cyclohexane by a cytochrome P-450 model. J. Chem. Soc. Chem. Commun. 1982, 1400–1401 (1982).
Breslow, R. & Gellman, S. H. Intramolecular nitrene carbon–hydrogen insertions mediated by transition-metal complexes as nitrogen analogs of cytochrome P-450 reactions. J. Am. Chem. Soc. 105, 6728–6729 (1983).
Svastits, E. W., Dawson, J. H., Breslow, R. & Gellman, S. H. Functionalized nitrogen atom transfer catalyzed by cytochrome P-450. J. Am. Chem. Soc. 107, 6427–6428 (1985).
Maynard Smith, J. Natural selection and the concept of a protein space. Nature 225, 563–564 (1970).
McCullum, E. O., Williams, B. A. R., Zhang, J. & Chaput, J. C. in In Vitro Mutagenesis Protocols Vol. 634 (ed. Braman, J.) 103–109 (Humana, 2010).
Arnold, F. H. Innovation by evolution: bringing new chemistry to life (Nobel lecture). Angew. Chem. Int. Ed. 58, 14420–14426 (2019).
Kuchner, O. & Arnold, F. H. Directed evolution of enzyme catalysts. Trends Biotechnol. 15, 523–530 (1997).
Zeymer, C. & Hilvert, D. Directed evolution of protein catalysts. Annu. Rev. Biochem. 87, 131–157 (2018).
Khersonsky, O. & Tawfik, D. S. Enzyme promiscuity: a mechanistic and evolutionary perspective. Annu. Rev. Biochem. 79, 471–505 (2010).
McIntosh, J. A. et al. Enantioselective intramolecular C–H amination catalyzed by engineered cytochrome P450 enzymes in vitro and in vivo. Angew. Chem. Int. Ed. 52, 9309–9312 (2013).
Steck, V., Kolev, J. N., Ren, X. & Fasan, R. Mechanism-guided design and discovery of efficient cytochrome P450-derived C–H amination biocatalysts. J. Am. Chem. Soc. 142, 10343–10357 (2020).
Jia, Z.-J., Gao, S. & Arnold, F. H. Enzymatic primary amination of benzylic and allylic C(sp3)–H bonds. J. Am. Chem. Soc. 142, 10279–10283 (2020).
Athavale, S. V. et al. Biocatalytic, intermolecular C–H bond functionalization for the synthesis of enantioenriched amides. Angew. Chem. Int. Ed. 60, 24864–24869 (2021).
Knight, A. M. et al. Diverse engineered heme proteins enable stereodivergent cyclopropanation of unactivated alkenes. ACS Cent. Sci. 4, 372–377 (2018).
Bordeaux, M., Tyagi, V. & Fasan, R. Highly diastereoselective and enantioselective olefin cyclopropanation using engineered myoglobin-based catalysts. Angew. Chem. Int. Ed. 54, 1744–1748 (2015).
Yang, Y. & Arnold, F. H. Navigating the unnatural reaction space: directed evolution of heme proteins for selective carbene and nitrene transfer. Acc. Chem. Res. 54, 1209–1225 (2021).
Chen, K., Huang, X., Kan, S. B. J., Zhang, R. K. & Arnold, F. H. Enzymatic construction of highly strained carbocycles. Science 360, 71–75 (2018).
Emmanuel, M. A., Greenberg, N. R., Oblinsky, D. G. & Hyster, T. K. Accessing non-natural reactivity by irradiating nicotinamide-dependent enzymes with light. Nature 540, 414–417 (2016).
Biegasiewicz, K. F. et al. Photoexcitation of flavoenzymes enables a stereoselective radical cyclization. Science 364, 1166–1169 (2019).
Grosheva, D. & Hyster, T. K. in Flavin‐Based Catalysis: Principles and Applications (eds. Cibulka, R. & Fraaije, M.) 291–313 (Wiley, 2021).
Sandoval, B. A., Meichan, A. J. & Hyster, T. K. Enantioselective hydrogen atom transfer: discovery of catalytic promiscuity in flavin-dependent ‘ene’-reductases. J. Am. Chem. Soc. 139, 11313–11316 (2017).
Huang, X. et al. Photoenzymatic enantioselective intermolecular radical hydroalkylation. Nature 584, 69–74 (2020).
Fu, H. et al. Ground-state electron transfer as an initiation mechanism for biocatalytic C–C bond forming reactions. J. Am. Chem. Soc. 143, 9622–9629 (2021).
Ji, P., Park, J., Gu, Y., Clark, D. S. & Hartwig, J. F. Abiotic reduction of ketones with silanes catalysed by carbonic anhydrase through an enzymatic zinc hydride. Nat. Chem. 13, 312–318 (2021).
Mukherjee, D., Ellern, A. & Sadow, A. D. Conversion of a zinc disilazide to a zinc hydride mediated by LiCl. J. Am. Chem. Soc. 132, 7582–7583 (2010).
Sattler, W. & Parkin, G. Zinc catalysts for on-demand hydrogen generation and carbon dioxide functionalization. J. Am. Chem. Soc. 134, 17462–17465 (2012).
Gao, X., Turek-Herman, J., Choi, Y. J., Cohen, R. D. & Hyster, T. K. Photoenzymatic Synthesis of α-tertiary amines by engineered flavin-dependent “ene”-reductases. J. Am. Chem. Soc. 143, 19643–19647 (2021).
Page, C. G. et al. Quaternary charge-transfer complex enables photoenzymatic intermolecular hydroalkylation of olefins. J. Am. Chem. Soc. 143, 97–102 (2021).
Mondal, D., Fisher, B. F., Jiang, Y. & Lewis, J. C. Flavin-dependent halogenases catalyze enantioselective olefin halocyclization. Nat. Commun. 12, 3268 (2021).
Huffman, M. A. et al. Design of an in vitro biocatalytic cascade for the manufacture of islatravir. Science 366, 1255–1259 (2019).
Kiefer, A., Liu, Y.-C., Gummerer, R., Jäger, C. & Deska, J. A fully biocatalytic approach to angiopterlactone B based on a chemoinspired artificial in vitro metabolism. Preprint at https://doi.org/10.26434/chemrxiv.14679738.v1 (2021).
Cai, T. et al. Cell-free chemoenzymatic starch synthesis from carbon dioxide. Science 373, 1523–1527 (2021).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2018).
Huang, P.-S., Boyken, S. E. & Baker, D. The coming of age of de novo protein design. Nature 537, 320–327 (2016).
Röthlisberger, D. et al. Kemp elimination catalysts by computational enzyme design. Nature 453, 190–195 (2008).
Siegel, J. B. et al. Computational design of an enzyme catalyst for a stereoselective bimolecular Diels–Alder reaction. Science 329, 309–313 (2010).
Bjelic, S. et al. Computational design of enone-binding proteins with catalytic activity for the Morita–Baylis–Hillman reaction. ACS Chem. Biol. 8, 749–757 (2013).
Blomberg, R. et al. Precision is essential for efficient catalysis in an evolved Kemp eliminase. Nature 503, 418–421 (2013).
Savile, C. K. et al. Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science 329, 305–309 (2010).
Wittmann, B. J., Yue, Y. & Arnold, F. H. Informed training set design enables efficient machine learning-assisted directed protein evolution. Cell Syst. 12, 1026–1045 (2021).
Wu, Z., Kan, S. B. J., Lewis, R. D., Wittmann, B. J. & Arnold, F. H. Machine learning-assisted directed protein evolution with combinatorial libraries. Proc. Natl Acad. Sci. USA 116, 8852–8858 (2019).
Goldman, S., Das, R., Yang, K. K. & Coley, C. W. Machine learning modeling of family wide enzyme-substrate specificity screens. Preprint at https://arxiv.org/abs/2109.03900 (2021).
Acknowledgements
This report was prepared as an account of work sponsored by an agency of the United States Government. Neither the United States Government nor any agency thereof, nor any of their employees, makes any warranty, express or implied, or assumes any legal liability or responsibility for the accuracy, completeness, or usefulness of any information, apparatus, product, or process disclosed, or represents that its use would not infringe on privately owned rights. Reference herein to any specific commercial product, process, or service by trade name, trademark, manufacturer, or otherwise does not necessarily constitute or imply its endorsement, recommendation, or favouring by the United States Government or any agency thereof. The views and opinions of the authors expressed herein do not necessarily state or reflect those of the United States Government or any agency thereof. D.C.M. was supported by a Ruth Kirschstein NIH Postdoctoral Fellowship (F32GM128247). This work was sponsored by the US Army Research Office and accomplished under cooperative agreement W911NF-19-2-0026 for the Institute of Collaborative Biotechnologies. This material is based on work sponsored by the US Department of Energy, Office of Basic Energy Sciences, under award number DE-SC0021141. We wish to thank E. Alfonzo, R. Mao and K. E. Johnston for helpful discussions. All protein structures were prepared using PyMOL (The PyMOL Molecular Graphics System, v.2.0 Schrödinger, LLC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review information
Nature Synthesis thanks Gerard Roelfes and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Alison Stoddart was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Miller, D.C., Athavale, S.V. & Arnold, F.H. Combining chemistry and protein engineering for new-to-nature biocatalysis. Nat Synth 1, 18–23 (2022). https://doi.org/10.1038/s44160-021-00008-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44160-021-00008-x