Abstract
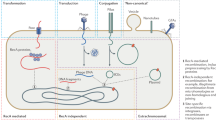
Unlike eukaryotes, which evolve principally through the modification of existing genetic information, bacteria have obtained a significant proportion of their genetic diversity through the acquisition of sequences from distantly related organisms. Horizontal gene transfer produces extremely dynamic genomes in which substantial amounts of DNA are introduced into and deleted from the chromosome. These lateral transfers have effectively changed the ecological and pathogenic character of bacterial species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lawrence, J. G. & Ochman, H. Molecular archaeology of the Escherichia coli genome. Proc. Natl Acad. Sci. USA 95, 9413–9417 ( 1998).
Davies, J. Origins and evolution of antibiotic resistance. Microbiologia 12, 9–16 (1996).
Doolittle, W. F. Phylogenetic classification and the universal tree. Science 284, 2124–2129 (1999).
Swofford, D. L., Olsen, G. J., Wadell, P. J. & Hillis, D. M. in Molecular Systematics (eds Hillis, D. M., Moritz, C. & Mable, B. K.) 407–514 (Sinauer Associates, Sunderland, Massachusetts, 1996).
Sueoka, N. On the genetic basis of variation and heterogeneity in base composition. Proc. Natl Acad. Sci. USA 48, 582– 592 (1962).
Muto, A. & Osawa, S. The guanine and cytosine content of genomic DNA and bacterial evolution. Proc. Natl Acad. Sci. USA 84, 166–169 ( 1987).
Karlin, S., Campbell, A. M. & Mrázek, J. Comparative DNA analysis across diverse genomes. Annu. Rev. Genet. 32, 185–225 (1998).
Groisman, E. A., Saier, M. H. Jr & Ochman, H. Horizontal transfer of a phosphatase gene as evidence for the mosaic structure of the Salmonella genome. EMBO J. 11, 1309–1316 ( 1992).
Lawrence, J. G. & Ochman, H. Amelioration of bacterial genomes: rates of change and exchange. J. Mol. Evol. 44, 383–397 ( 1997).
Lan, R. & Reeves, P. R. Gene transfer is a major factor in bacterial evolution. Mol. Biol. Evol. 13, 47–55 (1996).
Maynard Smith, J. The detection and measurement of recombination from sequence data. Genetics 153, 1021–1027 (1999).
Vulic, M., Dionisio, F., Taddei, F. & Radman, M. Molecular keys to speciation: DNA polymorphism and the control of genetic exchange in Enterobacteria. Proc. Natl Acad. Sci. USA 94, 9763– 9767 (1997).
Rayssiguier, C., Thaler, D. S. & Radman, M. The barrier to recombination between Escherichia coli and Salmonella typhimurium is disrupted in mismatch-repair mutants. Nature 342, 396– 399 (1989).
Medigue, C., Rouxel, T., Vigier, P., Henaut, A. & Danchin, A. Evidence for horizontal gene transfer in Escherichia coli speciation. J. Mol. Biol. 222, 851–856 (1991).
Whittam, T. S. & Ake, S. in Mechanisms of Molecular Evolution (eds Takahata, N. & Clark, A. G.) 223– 246 (Japan Scientific Society Press, Tokyo, 1992).
Riley, M. & Anilionis, A. Evolution of the bacterial genome. Annu. Rev. Microbiol. 32, 519– 560 (1978).
Wolf, Y. I., Aravind, L., Grishin, N. V. & Koonin, E. V. Evolution of aminoacyl-tRNA synthetases—analysis of unique domain architectures and phylogenetic trees reveals a complex history of horizontal gene transfer events. Genome Res. 9, 689– 710 (1999).
Aravind, L., Tatusov, R. L., Wolf, Y. I., Walker, D. R. & Koonin, E. V. Evidence for massive gene exchange between archaeal and bacterial hyperthermophiles. Trends Genet. 14, 442–444 ( 1998).
Nelson, K. E. et al. Evidence for lateral gene transfer between Archaea and Bacteria from genome sequence of Thermotoga maritima. Nature 399, 323–329 (1999).
Logsdon, J. M. Jr & Fuguy, D. M. Thermotoga heats up lateral gene transfer. Curr. Biol. 9, R747–R751 (1999).
Gaasterland, T. Archaeal genomics. Curr. Opin. Microbiol. 2, 542–547 (1999).
Dubnau, D. DNA uptake in bacteria. Annu. Rev. Microbiol. 53, 217–244 (1999).
Goodman, S. D. & Scocca, J. J. Identification and arrangement of the DNA sequence recognized in specific transformation of Neisseria gonorrhoeae. Proc. Natl Acad. Sci. USA 85, 6982–6986 (1988).
Elkins, C., Thomas, C. E., Seifert, H. S. & Sparling, P. F. Species-specific uptake of DNA by gonococci is mediated by a 10-base-pair sequence. J. Bacteriol. 173, 3911– 3913 (1991).
Smith, H. O., Tomb, J. -F., Dougherty, B. A., Fleischmann, R. D. & Venter, J. C. Frequency and distribution of DNA uptake signal sequences in the Haemophilus influenzae Rd genome. Science 269, 538– 540 (1985).
Davison, J. Genetic exchange between bacteria in the environment. Plasmid 42, 73–91 (1999).
Jiang, S. C. & Paul, J. H. Gene transfer by transduction in the marine environment. Appl. Environ. Microbiol. 64 , 2780–2787 (1998).
Schicklmaier, P. & Schmieger, H. Frequency of generalized transducing phages in natural isolates of the Salmonella typhimurium complex. Appl. Environ. Microbiol. 61, 1637–1640 (1995).
Buchanan-Wollaston, V., Passiatore, J. E. & Canon, F. The mob and oriT mobilization functions of a bacterial plasmid promote its transfer to plants. Nature 328, 170–175 (1987).
Heinemann, J. A. & Sprague, G. F. J. Bacterial conjugative plasmids mobilize DNA transfer between bacteria and yeast. Nature 340, 205–209 ( 1989).
Ricchetti, M., Fairhead, C. & Dujon, B. Mitochondrial DNA repairs double-strand breaks in yeast chromosomes. Nature 402, 96– 100 (1999).
Kleckner, N. in Mobile DNA (eds Berg, D. E. & Howe, M. M.) 227– 268 (American Society for Microbiology, Washington DC, 1989).
Berg, D. E. in Mobile DNA (eds Berg, D. E. & Howe, M. M.) 185– 210 (American Society for Microbiology, Washington DC. 1989).
Hall, R. M. Mobile gene cassettes and integrons: moving antibiotic resistance genes in Gram-negative bacteria. CIBA Found. Symp. 207, 192–205 (1997).
Rowe-Magnus, D. A. & Mazel, D. Resistance gene capture. Curr. Opin. Microbiol. 2, 483– 488 (1999).
Hall, R. M. & Collis, C. M. Mobile gene cassettes and integrons: capture and spread of genes by site-specific recombination. Mol. Microbiol. 15, 593–600 ( 1995).
Mazel, D., Dychinco, B., Webb, V. A. & Davies, J. A distinctive class of integron in the Vibrio cholerae genome. Science 280, 605–608 ( 1998).
Portnoy, D. A., Moseley, S. L. & Falkow, S. Characterization of plasmids and plasmid-associated determinants of Yersinia enterocolitica pathogenesis. Infect. Immun. 31, 775–782 ( 1981).
Maurelli, A. T., Baudry, B., d’Hauteville, H., Hale, T. L. & Sansonetti, P. J. Cloning of plasmid DNA sequences involved in invasion of HeLa cells by Shigella flexneri. Infect. Immun. 49, 164–171 (1985).
Sasakawa, C. et al. Virulence-associated genetic regions comprising 31 kilobases of the 230-kilobase plasmid in Shigella flexneri 2a. J. Bacteriol. 170, 2480–2484 (1988).
Gemski, P., Lazere, J. R., Casey, T. & Wohlhieter, J. A. Presence of virulence-associated plasmid in Yersinia pseudotuberculosis. Infect. Immun. 28, 1044–1047 (1980).
Isberg, R. R. & Falkow, S. A single genetic locus encoded by Yersinia pseudotuberculosis permits invasion of cultured animal cells by Escherichia coli K-12. Nature 317, 19–25 (1985).
McDaniel, T. K. & Kaper, J. B. A cloned pathogenicity island from enteropathogenic Escherichia coli confers the attaching and effacing phenotype on E. coli K-12. Mol. Microbiol. 23, 399–407 ( 1997).
Groisman, E. A. & Ochman, H. Pathogenicity islands: bacterial evolution in quantum leaps. Cell 87, 791–794 (1996).
Hacker, J., Blum-Oehler, G., Muhldorfer, I. & Tschape, H. Pathogenicity islands of virulent bacteria: structure, function and impact on microbial evolution. Mol. Microbiol. 23, 1089–1097 (1997).
Ritter, A. et al. tRNA genes and pathogenicity islands: influence on virulence and metabolic properties of uropathogenic Escherichia coli. Mol. Microbiol. 17, 109–121 (1995).
Blum, G et al. Excision of large DNA regions termed pathogenicity islands from tRNA-specific loci in the chromosome of an Escherichia coli wild-type pathogen. Infect. Immun. 62, 606– 614 (1994).
Moss, J. E., Cardozo, T. J., Zychlinsky, A. & Groisman, E. A. The selC-associated SHI-2 pathogenicity island of Shigella flexneri . Mol. Microbiol. 33, 74– 83 (1999).
Vokes, S. A., Reeves, S. A., Torres, A. G. & Payne, S. M. The aerobactin iron transport system genes in Shigella flexneri are present within a pathogenicity island. Mol. Microbiol. 33, 63–73 (1999).
Blanc-Potard, A. B. & Groisman, E. A. The Salmonella selC locus contains a pathogenicity island mediating intramacrophage survival. EMBO J. 16, 5376– 5385 (1997).
Sun, J., Inouye, M. & Inouye, S. Association of a retroelement with a P4-like cryptic prophage (retronphage Theta;R73) integrated into the selenocystyl-tRNA gene of Escherichia coli. J. Bacteriol. 173, 171–181 (1991).
Cheetham, B. F. & Katz, M. E. A role for bacteriophages in the evolution and transfer of bacterial virulence determinants. Mol. Microbiol. 18, 201–208 (1995).
Lindsay, J. A., Ruzin, A., Ross, H. F., Kurepina, N. & Novick, R. P. The gene for toxic shock toxin is carried by a family of mobile pathogenicity islands in Staphylococcus aureus. Mol. Microbiol. 29, 527–543 (1998).
Weeks, C. R. & Ferretti, J. J. The gene for type A streptococcal exotoxin (erythrogenic toxin) is located in bacteriophage T12. Infect. Immun. 46, 531–536 (1984).
Jackson, M. P., Neill, R. J., O'Brien, A. D., Holmes, R. K. & Newland, J. W. Nucleotide sequence analysis and comparison of the structural genes for Shiga-like toxin-I and Shiga-like toxin II encoded by bacteriophages from Escherichia coli. FEMS Microbiol. Lett. 44, 109–114 (1987).
Mirold, S. Isolation of a temperate bacteriophage encoding the type III effector protein SopE from an epidemic Salmonella typhimurium strain. Proc. Natl Acad. Sci. USA 96, 9845–9850.
Waldor, M. K. & Mekalanos, J. J. Lysogenic conversion by a filamentous phage encoding cholera toxin. Science 272, 1910–1914 (1996).
Karaolis, D. K., Somara, S., Maneval, D. R. J., Johnson, J. A. & Kaper, J. B. A bacteriophage encoding a pathogenicity island, a type-IV pilus and a phage receptor in cholera bacteria. Nature 399, 675–679 ( 1999).
Nakata, N. et al. The absence of a surface protease, OmpT, determines the intercellular spreading ability of Shigella: the relationship between the ompT and kcpA loci. Mol. Microbiol. 9, 459–468 (1993).
Maurelli, A. T., Fernández, R. E., Bloch, C. A., Rode, C. K. & Fasano, A. “Black holes” and bacterial pathogenicity: a large genomic deletion that enhances the virulence of Shigella spp. and enteroinvasive Escherichia coli. Proc. Natl Acad. Sci. USA 95, 3943– 3948 (1998).
Riley, M. & Sanderson, K. E. in The Bacterial Chromosome (eds Riley, M. & Drlica, K.) 85–96 (American Society for Microbiology, Washington DC, 1990).
Lawrence, J. G. & Roth, J. R. Selfish operons: horizontal transfer may drive the evolution of gene clusters. Genetics 143, 1843–1860 (1996).
Wolf, Y. I., Aravind, L. & Koonin, E. V. Rickettsiae and Chlamydia: evidence of horizontal gene transfer and gene exchange. Trends Genet. 14, 442–444 (1998).
Groisman, E. A. & Ochman, H. How Salmonella became a pathogen. Trends Microbiol. 5, 343–349 (1997).
Groisman, E. A. The ins and outs of virulence gene expression: Mg2+ as a regulatory signal. Bioessays 20, 96– 101 (1998).
Deiwick, J., Nikolaus, T., Erdogan, S. & Hensel, M. Environmental regulation of Salmonella pathogenicity island 2 gene expression. Mol. Microbiol. 31, 1759– 1773 (1999).
Casjens, S. The diverse and dynamic structure of bacterial genomes. Annu. Rev. Genet. 32, 339–377 ( 1998).
Andersson, S. G. E. & Kurland, C. G. Reductive evolution of resident genomes. Trends Microbiol. 6, 263–268 (1998).
Andersson, J. O. & Andersson, S. G. E. Insights into the evolutionary process of genome degradation. Curr. Opin. Genet. Dev. 9, 664–671 ( 1999).
Lawrence, J. G. Gene transfer, speciation, and the evolution of bacterial genomes. Curr. Opin. Microbiol. 2, 519–523 (1999).
Lawrence, J. G. & Roth, J. R. in Organization of the Prokaryotic Genome (ed. Charlebois, R. L.) 263– 289 (American Society for Microbiology, Washington, DC, 1999).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ochman, H., Lawrence, J. & Groisman, E. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000). https://doi.org/10.1038/35012500
Issue Date:
DOI: https://doi.org/10.1038/35012500
This article is cited by
-
Pangenome analysis of Shewanella xiamenensis revealed important genetic traits concerning genetic diversity, pathogenicity and antibiotic resistance
BMC Genomics (2024)
-
Health impacts of an extreme dust event: a case and risk assessment study on airborne bacteria in Beijing, China
Environmental Sciences Europe (2024)
-
Duplicated antibiotic resistance genes reveal ongoing selection and horizontal gene transfer in bacteria
Nature Communications (2024)
-
Horizontal gene transfer in eukaryotes: aligning theory with data
Nature Reviews Genetics (2024)
-
Engineering intelligent chassis cells via recombinase-based MEMORY circuits
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.