Abstract
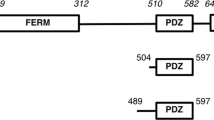
Papillomaviruses cause warts and proliferative lesions in skin and other epithelia. In a minority of papillomavirus types (‘high risk’, including human papillomaviruses 16, 18, 31, 33, 45 and 56), further transformation of the wart lesions can produce tumours1. The papillomavirus E2 protein controls primary transcription and replication of the viral genome2. Both activities are governed by a ∼200 amino-acid amino-terminal module (E2NT) which is connected to a DNA-binding carboxy-terminal module by a flexible linker. Here we describe the crystal structure of the complete E2NT module from human papillomavirus 16. The E2NT module forms a dimer both in the crystal and in solution. Amino acids that are necessary for transactivation are located at the dimer interface, indicating that the dimer structure may be important in the interactions of E2NT with viral and cellular transcription factors. We propose that dimer formation may contribute to the stabilization of DNA loops3 which may serve to relocate distal DNA-binding transcription factors to the site of human papillomavirus transcription initiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
zur Hausen, H. Molecular pathogenesis of cancer of the cervix and its causation by specific human papillomavirus types. Curr. Top. Microbiol. Immunol. 186, 131–156 (1994).
McBride, A. & Myers, G. in Human Papillomaviruses 1997 (eds Myers, G. et al.) III-54–III-73 (Theoretical Biology and Biophysics, Los Alamos National Laboratory, Los Alamos, New Mexico, 1997).
Knight, J. D., Li, R. & Botchan, M. The activation domain of the bovine papillomavirus E2 protein mediates association of DNA-bound dimers to form DNA loops. Proc. Natl Acad. Sci. USA 88, 3204–3208 ( 1991).
Mohr, I. J. et al. Targeting the E1 replication protein to the papillomavirus origin of replication by complex formation with the E2 transactivator. Science 250, 1694–1699 ( 1990).
Breiding, D. E. et al. Functional interaction of a novel cellular protein with the papillomavirus E2 transactivation domain. Mol. Cell. Biol. 17, 7208–7219 (1997).
Yao, J. M., Breiding, D. E. & Androphy, E. J. Functional interaction of the bovine papillomavirus E2 transactivation domain with TFIIB. J. Virol. 72, 1013–1019 (1998).
Hegde, R. S., Grossman, S. R., Laimins, L. A. & Sigler, P. B. Crystal structure at 1.7 Å of the bovine papillomavirus-1 E2 DNA-binding domain bound to its DNA target. Nature 359, 505–512 (1992).
Harris, S. F. & Botchan, M. R. Crystal structure of the human papillomavirus type 18 E2 activation domain. Science 284, 1673–1677 (1999).
Bernstein, F. C. et al. The protein data bank: a computer based archival file for macromolecular structures. J. Mol. Biol. 112, 535–542 (1977).
Cooper, C. S., Upmeyer, S. N. & Winokur, P. L. Identification of single amino acids in the human papillomavirus 11 E2 protein critical for the transactivation or replication functions. Virol. 241, 312– 322 (1998).
Mok, Y. K., Gay, G. D., Butler, P. J. & Bycroft, M. Equilibrium dissociation and unfolding of the dimeric human papillomavirus strain-16 E2 DNA-binding domain. Protein Sci. 5, 310– 319 (1996).
Foguel, D., Silva, J. L. & de Prat-Gay, G. Characterization of a partially folded monomer of the DNA-binding domain of human papillomavirus E2 protein obtained at high pressure. J. Biol. Chem. 273, 9050– 9057 (1998).
Estojak, J., Brent, R. & Golemis, E. Correlation of two-hybrid affinity with in vitro measurements. Mol. Cell. Biol. 15, 5820– 5829 (1995).
Sengchanthalangsy, L. et al. Characterisation of the dimer interface of transcription factor NKkB p50 homodimer. J. Mol. Biol. 289, 1029–1040 (1999).
Chao, S.-F et al. Subunit affinities and stoichiometries of the human papillomavirus type 11 E1:E2:DNA complex. Biochemistry 38, 4586–4594 (1999).
Gauthier, J.-M., Dostatni, N., Lusky, M. & Yaniv, M. Two DNA-bound E2 dimers are required for strong transcriptional activation and for cooperation with cellular factors in most cells. The New Biologist 3, 498–509 (1991).
Abroi, A., Kurg, R. & Ustav, M. Transcriptional and replicational activation functions in the bovine papillomavirus type 1 E2 protein are encoded by different structural determinants. J. Virol. 70, 6169–6179 (1996).
Brokaw, J. L., Blanco, M. & McBride, A. A. Amino acids critical for the functions of the bovine papillomavirus type 1 E2 transactivator. J. Virol. 70, 23–29 (1996).
Desaintes, C. & Demeret, C. Control of papillomavirus DNA replication and transcription. Semin. Cancer Biol. 7, 339–347 (1996).
Burns, J. E. et al. Expression, crystallization and preliminary X-ray analysis of the E2 transactivation domain from papillomavirus type 16. Acta Crystallogr. D 54, 1471–1474 (1998).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276 , 307–326 (1997).
Collaborative Computational Project N4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760– 763 (1994).
Sheldrick, G. M. & Schneider, T. R. SHELXL: high-resolution refinement. Methods Enzymol. 277, 319– 343 (1997).
Cowtan, K. D. & Main, P. Phase combination and cross-validation in iterated density-modification calculations. Acta Crystallogr. D 52, 43–48 ( 1996).
Oldfield, T.J. in Proceedings of the CCP4 Study Weekend (eds Bailey, S., Hubbard, R. & Waller D.) 15–18 (Daresbury Laboratory, Warrington, UK, 1994).
Laue, T.M., Shah, B.D., Ridgeway, T.M. & Pelletier, S.L. Computer-aided interpretation of analytical sedimentation data for proteins. In Analytical Ultracentrifugation in Biochemistry and Polymer Science (eds Harding, S.E., Rowe, A.J. & Horton, J.C.) 90–125 (Royal Society of Chemistry, London, 1992).
Kraulis, P.J. MOLSCRIPT—a program to produce both detailed and schematic plots of proteins structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Esnouf, R.M. An extensively modified version of Molscript that includes greatly enhanced coloring capabilities. J. Mol. Graphics 15, 133–138 (1997).
Merritt, E.A & Bacon, D.J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Acknowledgements
K.S.W., G.G.D., A.A.A and O.V.M. thank the BBSRC for infrastructure support and the EC for supporting work at EMBL, Hamburg, through the Access to Large Installations Project, Contract Number CHGE-CT93-0040. We also thank Yorkshire Cancer Research for project and program grant support to N.J.M., J.E.B., I.B.B. and O.V.M. A.A.A. is supported by a Wellcome Trust Career Development Fellowship.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Antson, A., Burns, J., Moroz, O. et al. Structure of the intact transactivation domain of the human papillomavirus E2 protein. Nature 403, 805–809 (2000). https://doi.org/10.1038/35001638
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35001638
This article is cited by
-
Transgenic HPV11-E2 protein modulates URR activity in vivo
Transgenic Research (2023)
-
Evolutionary variation of papillomavirus E2 protein and E2 binding sites
Virology Journal (2011)
-
Oncolytic adenovirus armed with human papillomavirus E2 gene in combination with radiation demonstrates synergistic enhancements of antitumor efficacy
Cancer Gene Therapy (2011)
-
Triggering of death receptor apoptotic signaling by human papillomavirus 16 E2 protein in cervical cancer cell lines is mediated by interaction with c-FLIP
Apoptosis (2011)
-
Direct activation of caspase 8 by the proapoptotic E2 protein of HPV18 independent of adaptor proteins
Cell Death & Differentiation (2008)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.