Abstract
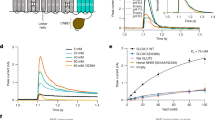
In most multicellular organisms direct cell–cell communication is mediated by the intercellular channels of gap junctions. These channels allow the exchange of ions and molecules that are believed to be essential for cell signalling during development and in some differentiated tissues. Proteins called connexins, which are products of a multigene family, are the structural components of vertebrate gap junctions1,2. Surprisingly, molecular homologues of the connexins have not been described in any invertebrate. A separate gene family, which includes the Drosophila genes shaking-B and l(1)ogre, and the Caenorhabditis elegans genes unc-7 and eat-5, encodes transmembrane proteins with a predicted structure similar to that of the connexins3,4,5,6,7,8,9. shaking-B and eat-5 are required for the formation of functional gap junctions8,10. To test directly whether Shaking-B is a channel protein, we expressed it in paired Xenopus oocytes. Here we show that Shaking-B localizes to the membrane, and that its presence induces the formation of functional intercellular channels. To our knowledge, this is the first structural component of an invertebrate gap junction to be characterized.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bruzzone, R., White, T. W. & Paul, D. L. Connections with connexins: the molecular basis of direct intercellular signalling. Eur. J. Biochem. 238, 1–27 (1996).
Kumar, N. M. & Gilula, N. B. The gap junction communication channel. Cell 84, 381–388 (1996).
Crompton, D., Todman, M., Wilkin, M., Ji, S. & Davies, J. Essential and neural transcripts from the Drosophila shaking-B locus are differentially expressed in the embryonic mesoderm and pupal nervous system. Dev. Biol. 170, 142–158 (1995).
Krishnan, S. N., Frei, E., Swain, G. P. & Wyman, R. J. Passover: a gene required for synaptic connectivity in the giant fibre system of Drosophila. Cell 73, 967–977 (1993).
Krishnan, S. N., Frei, E., Schalet, A. P. & Wyman, R. J. Molecular basis of intracistronic complementation in the Passover locus of Drosophila. Proc. Natl Acad. Sci. USA 92, 2021–2025 (1995).
Watanabe, T. & Kankel, D. R. Molecular cloning and analysis of l (1) ogre, a locus of Drosophila melanogaster with prominent effects on the postembryonic development of the central nervous system. Genetics 126, 1033–1044 (1990).
Starich, T. A., Herman, R. K. & Shaw, J. E. Molecular and genetic analysis of unc-7, a Caenorhabditis elegans gene required for coordinated locomotion. Genetics 133, 527–541 (1993).
Starich, T. A., Lee, R. Y. N., Panzarella, C., Avery, L. & Shaw, J. E. eat-5 and unc-7 represent a multigene family in Caenorhabditis elegans involved in cell–cell coupling. J. Cell Biol. 134, 537–548 (1996).
Barnes, T. M. OPUS: a growing family of gap junction proteins? Trends Genet. 10, 303–305 (1994).
Phelan, P. et al. Mutations in shaking-B prevent electrical synapse formation in the Drosophila giant fibre system. J. Neurosci. 16, 1101–1113 (1996).
Homyk, T., Szidonya, J. & Suzuki, D. T. Behavioral mutants of Drosophila melanogaster III. Isolation and mapping of mutations by direct visual observations of behavioral phenotypes. Mol. Gen. Genet. 177, 553–565 (1980).
Thomas, J. B. & Wyman, R. J. Mutations altering synaptic connectivity between identified neurons in Drosophila. J. Neurosci. 4, 530–538 (1984).
Willecke, K. & Haubrich, S. Connexin expression systems: To what extent do they reflect the situation in the animal? J. Bioenerg. Biomembr. 28, 319–326 (1996).
Barrio, L. C. et al. Gap junctions formed by connexins 26 and 32 alone and in combination are differently affected by applied voltage. Proc. Natl Acad. Sci. USA 88, 8410–8414 (1991).
Hennemann, H. et al. Two gap junction genes, connexin 31.1 and 30.3, are closely linked on mouse chormosome 4 and preferentially expressed in skin. J. Biol. Chem. 267, 17225–17233 (1992).
Swenson, K. I., Jordan, J. R., Beyer, E. C. & Paul, D. L. Formation of gap junctions by expression of connexins in Xenopus oocyte pairs. Cell 57, 145–155 (1989).
Bruzzone, R., White, T. W. & Paul, D. L. Expression of chimeric connexins reveals new properties of the formation and gating behavior of gap junction channels. J. Cell Sci. 107, 955–967 (1994).
Wilders, R. & Jongsma, H. J. Limitations of the dual voltage clamp method in assaying conductance and kinetics of gap junction channels. Biophys. J. 63, 942–953 (1992).
Ebihara, L., Beyer, E. C., Swenson, K. I., Paul, D. L. & Goodenough, D. A. Cloning and expression of a Xenopus embryonic gap junction protein. Science 243, 1194–1195 (1989).
Verselis, V. K., Bennett, M. V. L. & Bargiello, T. A. Avoltage-dependent gap junction in Drosophila melanogaster. Biophys. J. 59, 114–126 (1991).
Gho, M. Voltage-clamp analysis of gap junctions between embryonic muscles in Drosophila. J. Physiol. (Lond.) 481, 371–383 (1994).
Bruzzone, R., White, T. W., Yoshizaki, G., Patino, R. & Paul, D. L. Intercellular channels in teleosts: functional characterization of two connexins from Atlantic croaker. FEBS Lett. 385, 301–304 (1995).
Chevalier, S., Tassan, J.-P., Cox, R., Phillipe, M. & Ford, C. Both cdk2 and cdc2 proteins promote S phase initiation in frog egg extracts. J. Cell Sci. 108, 1831–1841 (1995).
Colman, A. in Transcription and Translation—A Practical Approach (ed. Hames, B. D. & Higgins, S. J.) 271–302 (IRL, Oxford, (1984)).
Hausen, P. & Dreyer, C. The use of polyacrylamide as an embedding medium for immunohistochemical studies of embryonic tissues. Stain Technol. 56, 287–293 (1981).
Spray, D. C., Harris, A. L. & Bennett, M. V. L. Equilibrium properties of a voltage-dependent junctional conductance. J. Gen. Physiol. 77, 77–93 (1981).
Acknowledgements
We thank H. Woodland for the pSPJC2L vector, and E. Jones, S. King, M. O'Shea, D. Paul, M. Stern, M. Todman and M. Yeoman for help, advice and discussion. This work was funded by the BBSRC, UK.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Phelan, P., Stebbings, L., Baines, R. et al. Drosophila Shaking-B protein forms gap junctions in paired Xenopus oocytes. Nature 391, 181–184 (1998). https://doi.org/10.1038/34426
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/34426
This article is cited by
-
Are there gap junctions without connexins or pannexins?
BMC Evolutionary Biology (2019)
-
Gap junction gene innexin3 being highly expressed in the nervous system and embryonic stage of the mud crab Scylla paramamosain
Journal of Oceanology and Limnology (2019)
-
Drosophila seizure disorders: genetic suppression of seizure susceptibility
Frontiers in Biology (2016)
-
Innexin gap junctions in nerve cells coordinate spontaneous contractile behavior in Hydra polyps
Scientific Reports (2014)
-
Cell-to-cell communication in plants, animals, and fungi: a comparative review
Naturwissenschaften (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.