Abstract
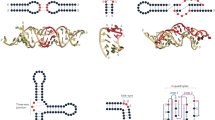
THE structure of 5S RNA has been a lively field for experiment and speculation, with no definitive resolution of the problem in sight (see refs 1 and 2 for reviews). Fox and Woese3 suggested comparative sequence analysis of prokaryotic 5S RNAs as a method for locating base paired regions, an appealing procedure because of its similarity to the method used to substantiate the cloverleaf model for tRNA. Its logic states that if all known 5S RNA sequences contain complementary bases at a site, a double helix probably exists there in the functional form. We propose that comparative sequence analysis suggests two different functional structures for prokaryotic 5S RNA. One of them has been introduced by Fox and Woese3, and its set condary structure is based on four conserved helices: the ‘molecular stalk’ (Escherichia coli residues 1–10 and 119–110), the weak ‘tuned helix’ (18–23/65–60), the ‘common arm base’ (31–34/51–48) and the ‘prokaryotic loop’ (82-86/94-90). The base paired structure that results is consistent with enzymatic digestion14 and chemical modification1,5 results on native 5S RNA, if allowance is made for protection of residues by tertiary bonding as well as secondary structure. This ‘model’, however, does not explain the apparent conservation of a double helix (33–39/88–82 in E. coli) (Table 1) which is sterically inconsistent with two of the helices listed (the common arm base and the prokaryotic loop). Fox and Woese3 noted this conservation but considered it unimportant because of the variable helix length and the conflict with the other two helices.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Monier, R. in Ribosomes (eds Nomura, M., Tissières, A. & Lengyel, P.) 141–168 (Cold Spring Harbor, New York, 1974).
Erdmann, V. A. Progr. Nucl. Acid Res. 18, 45–90 (1976).
Fox, G. E. & Woese, C. R. Nature 256, 505–507 (1975).
Jordan, B. R. J. molec. Biol. 55, 423–439 (1971).
Aubert, M., Bellemare, G. & Monier, R. Biochimie 55, 135–142 (1973).
Brownlee, G. G., Sanger, F. & Barrell, B. G. Nature 215, 735–736 (1967).
Dubuy, B. & Weissman, S. M. J. biol. Chem. 246, 747–761 (1971).
Pribula, C. D., Fox, G. E. & Woese, C. R. FEBS Lett. 64, 350–352 (1976).
Pribula, C. D., Fox, G. E., Woese, C. R., Sogin, M. & Pace, N. FEBS Lett. 44, 322–323 (1974).
Marotta, C. A., Varicchio, F., Smith, I. & Weissman, S. M. J. biol. Chem. 251, 3122–3127 (1976).
Raué, H. A., Stoof, T. J. & Planta, R. J. Eur. J. Biochem. 59, 35–42 (1975).
Corry, M. J., Payne, P. I. & Dyer, T. A. FEBS Lett. 46, 63–66 (1974).
Aubert, M., Scott, J. F., Reynier, M. & Monier, R. Proc. natn. Acad. Sci. U.S.A. 61, 292–299 (1968).
Lecanidou, R. & Richards, E. G. Eur. J. Biochem. 57, 127–133 (1975).
Hilbers, C. W. & Shulman, R. G. Proc. natn. Acad. Sci. U.S.A. 71, 3239–3242 (1974).
Wong, L. L. & Kearns, D. R. Biopolymers 13, 371–380 (1974).
Daniel, W. E. & Colby, M. Proc. natn. Acad. Sci. U.S.A. 72, 2582–2586 (1975).
Kearns, D. R. & Wong, Y. P. J. molec. Biol. 87, 755–774 (1974).
Maassen, J. A. & Möller, W. Biochem. biophys. Res. Commun. 64, 1175–1183 (1975).
Belitsina, N. V., Glukhova, M. A. & Spirin, A. S. J. molec. Biol. 108, 609–613 (1976).
Herr, W. & Noller, F. FEBS Lett. 53, 248–252 (1975).
Erdmann, V. A., Sprinzl, M. & Pongs, O. Biochem. biophys. Res. Commun. 54, 942–948 (1973).
Schwarz, U., Menzel, H. M. & Gassen, H. G. Biochemistry 15, 2484–2490 (1976).
Riddle, D. L. & Carbon, J. Nature new Biol. 242, 230–234 (1973).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
WEIDNER, H., YUAN, R. & CROTHERS, D. Does 5S RNA function by a switch between two secondary structures?. Nature 266, 193–194 (1977). https://doi.org/10.1038/266193a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/266193a0
This article is cited by
-
RNA secondary structures and their prediction
Bulletin of Mathematical Biology (1984)
-
Heterogeneity of the genes coding for 5 S RNA in three related strains of the genus Bacillus
Molecular and General Genetics MGG (1977)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.