Abstract
Ataxia-telangiectasia (A-T) is a progressive genetic disorder affecting the central nervous and immune systems, and involving chromosomal instability, cancer predisposition, radiation sensitivity and cell cycle abnormalities. Studies of the cellular phenotype of A-T have pointed to a defect in a putative system that processes a specific type of DNA damage and initiates a signal transduction pathway controlling replication and repair. A-T is genetically heterogeneous, with 4 complementation groups. While functional cloning of the A-T gene(s) using gene transfer has proven problematic, positional cloning attempts are zeroing in on a defined interval on chromosome 11q22–23 that probably harbors the mutations for all 4 complementation groups.
Similar content being viewed by others
Ataxia-Telangiectasia: A Pleiotropic Defect in an Essential Junction of Cellular Physiology
Dr. Elena Boder’s review [1] on the genetic disorder ataxia-telangiectasia (A-T) 10 years ago began with the words: ‘Ataxia-telangiectasia has been mysterious from the start.’ Since its establishment as a clinical entity in 1957 [2], this enigmatic, insidious disorder has presented a biological, medical and human challenge to clinicians and researchers. The mutations responsible for A-T seem to affect an unidentified physiological junction linking the differentiation of various tissues, essential functions in the central nervous and immune systems, genome stability, DNA replication, recombination and repair, cell cycle control, cellular aging and neoplastic transformation [3–8]. Hence, identifying the sites of these mutations is expected to have far-ranging effects in several areas of biomedical research.
A-T is inherited in an autosomal recessive manner and has been found worldwide, with patient frequencies of about 1:100,000 in the United States and Britain [9–12]. There are notable concentrations of A-T patients also in Turkey [13], Italy [14] and among Moroccan Jews in Israel [15].
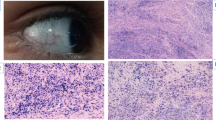
A-T makes its appearance initially as a neurological disorder [1, 4, 5]. Cerebellar ataxia begins in infancy and progresses steadily, confining the patient to a wheelchair by the beginning of the second decade of life. Other main neurological signs are involuntary movements, diminished or absent deep reflexes, apraxia of eye movements and slurred speech. The neuropathological hallmark of A-T is cerebellar degeneration involving primarily the Purkinje and granular cells; degenerative changes have also been noted in the spinal cord and ganglia, brainstem and peripheral nerves. The second clinical hallmark of A-T, which typically appears between the ages of 3 and 6 years, is telangiectases (dilation of blood vessels making them more prominent) in the eyeballs and conjunctiva, sometimes spreading over sun-exposed areas of the skin. Some 50–80% of patients show the third clinical hallmark of A-T, recurrent sinopulmonary infections signifying marked immunodeficiency. Serum levels of IgA, IgG2 and/or IgE are reduced, the number of circulating lymphocytes is diminished and mitogen response is poor. The thymus is degenerated and sometimes absent. The serum levels of two oncofetal proteins — α-fetoprotein and carcinoembryonic antigen — are consistently higher in A-T patients. Somatic growth and sexual maturation are usually retarded, with female hypogonadism being almost uniform. Progerie changes typically appear in the hair and skin, marking premature senescence. Intelligence is usually normal.
Another cardinal feature of A-T is profound cancer predisposition, which becomes evident in about 10% of patients during childhood [16–18]. Lymphomas and acute lymphocytic leukemia constitute over 85% of all cancers in A-T and appear primarily in younger patients. Other cancers, mainly epithelial, rise steadily with age [10]. Early attempts to treat these malignancies by radiotherapy resulted in acute radiation reactions, revealing another feature of A-T — a profound sensitivity to the cytotoxic effect of ionizing radiation [19–24]. The course of A-T is progressive and relentless, and patients usually die with respiratory failure or malignancy during the second or beginning of the third decade of life. There is no effective way to retard the progression of this disease.
The primary diagnostic laboratory finding in A-T is chromosomal instability, evident as high rates of chromosomal breaks, usually observed in peripheral lymphocytes or fibroblasts [25–27]. Lymphocyte cultures show cell clones containing specific chromosomal translocations involving particularly the chromosomal regions 7p14, 7q35, 14q12 and 14q32 which harbor the T-cell receptor and immunoglobulin heavy-chain genes. Such clones often precede the onset of lymphoreticular malignancies and subsequently undergo clonal expansion as malignancy progresses [27–36]. Molecular analysis of several translocation breakpoints showed that the immune system genes residing in these regions were indeed involved in these aberrations [30, 32, 33, 35, 36].
The cellular phenotype of A-T further reflects the complexity of this disorder. Besides chromosomal instability, A-T cells show a reduced life span in culture, higher requirements for unspecified serum growth factors, abnormalities in the shape and arrangement of cytoskeletal actin fibers and abnormal content of a variety of extracellular surface proteins [37, 38]. A major cellular characteristic of A-T, which has become diagnostic, is the profound sensitivity of the cells to the cytotoxic and clastogenic effects of ionizing radiation and radiomimetic chemicals. A-T cells are hypersensitive to both high- and low-energy transfer ionizing irradiation, as well as to a host of chemicals that mimic the effect of ionizing radiation on DNA by their capacity to produce radicals capable of inducing strand scissions [3, 39–51]. However, the overall kinetics of single- and double-strand break repair in A-T cells has been found normal in most studies [3, 39, 44, 49, 52, 53]. Unexpectedly, semiconservative DNA synthesis in A-T cells was found to be more resistant to the inhibitory effect of the DNA damaging agents to which A-T cells are sensitive, most notably ionizing radiation [54–56]. This phenomenon, called ‘radioresistant DNA synthesis’ (RDS), was the first evidence of a defect in cell cycle control in A-T cells (see below).
The clinical and cellular characteristics of A-T have been used to delineate phenotypic variants among patients. Patients with somewhat milder clinical signs, later age of onset and slower progression of the disease were found in several countries, and in some of them this phenotype was correlated with milder radiosensitivity and sometimes reduced or absent RDS. These parameters do not always coexist, however, demonstrating the complexity of the molecular and physiological basis of A-T [6, 8, 13, 57–65].
Another dimension is added to this complexity by other disorders with certain characteristics shared with A-T, such as immunodeficiency coupled with chromosomal instability [66–68]. The combination of microcephaly, growth retardation, immunodeficiency, chromosomal instability, radiosensitivity and RDS but no telangiectases has been particularly related to A-T. Patients with this syndrome, sometimes associated with mental retardation, were reported in several ethnic groups, and some were classified as Nijmegen breakage syndrome [69–77]. Patients with Nijmegen breakage syndrome tend to develop lymphoreticular malignancies at a higher rate than classical A-T patients [77]. One case of A-T with microcephaly and mental retardation was designated ‘ATFresno’ [78].
Attempts to delineate the possible genetic heterogeneity of A-T and its relationship with related syndromes have been done by fusing cells from different patients and measuring RDS [79–81] or radiation-induced chromosomal aberrations [82, 83] in heterokaryons. These studies revealed 4 complementation groups in classical A-T, designated A, C, D and E, and 2 complementation groups, VI and V2, among patients with Nijmegen breakage syndrome and related syndromes [77, 81]. Groups A and C accounted for 83% of cases in a sample of 50 A-T patients [81], but the size of this series precludes making generalizations to A-T worldwide. In general, no correlation was found between complementation group assignment and clinical variation in A-T. It is unclear at present whether these complementation groups represent different genes or different mutations within 1 gene. A-T researchers cautiously refer to 4 genes involved in classical A-T: ATA, ATC, ATD and ATE.
A-T heterozygotes have always received special attention. In a sense A-T is not entirely recessive since carriers mildly manifest two of the disease characteristics, cancer predisposition and radiosensitivity. Epidemiological studies consistently show that A-T heterozygotes exhibit a higher rate of certain cancers, especially breast cancer in women [9, 11, 16, 84–88]. Swift et al. [86] estimated the cancer tendency among male A-T heterozygotes to be 3.8-fold higher than that of the general population, while that of female carriers was estimated to be 3.5 higher. However, the relative risk for breast cancer in women alone was estimated to be 5.1 higher than that of a control population. Swift et al. [84] further suggested that up to 8.8% of the American white female patients with breast cancer may be A-T carriers. Easton [88] found in British and Norwegian populations a 3.9-fold risk for breast cancer in female A-T heterozygotes, while no tendency to develop other cancers was found among A-T carriers in general. These findings were accompanied by repeated reports of moderate sensitivity of cells from A-T carriers to ionizing radiation, as measured by survival and cytogenetic assays [89–108]. These results imply that A-T heterozygotes might face special hazards from routine diagnostic or therapeutic procedures involving radiation. These findings also stimulated attempts to develop laboratory assays for the detection of A-T carriers in the general population. While obligatory A-T heterozygotes could be clearly delineated from controls in some studies [89, 91, 94–98], groups of controls and A-T heterozygotes overlapped in other samples [92, 93, 101, 107]. The wide range of radiation responses among control groups makes the reliability of this parameter as an assay for carrier detection doubtful. It should also be noted that most of these methods are labor intensive and results vary between laboratories.
A-T: A Defect in Processing of a Specific DNA Lesion Affecting a Downstream Signal Transduction System
The function presumed defective in A-T is associated with the processing of a specific type of a DNA lesion. Examination of the mode of action of chemical agents to which A-T cells are hypersensitive pointed to a specific type of strand scission induced by these agents via a free radical attack on the deoxyribose moiety [40]. This high specificity should be borne in mind when attempting to make deductions about the A-T defect from the cellular phenotype. The high sensitivity of A-T cells to agents inducing this lesion has been attributed to a defect in DNA repair or in a mechanism controlling replication of damaged DNA [3, 5, 6, 39, 49]. It is noteworthy in this respect that the RDS phenomenon could be clearly dissociated from radiosensitivity in several reports [46, 59, 109–112]. Early observations that A-T cells are deficient in ‘potentially lethal damage repair’ acting in quiescent cells [113–115] seemed to support the DNA repair hypothesis. The natural candidate for the critical DNA lesion unrepaired in A-T cells was DNA breaks. Most studies based on biophysical methods have failed to reveal a defect in the overall kinetics of DNA strand break repair in A-T cells [3, 39, 44, 49, 52, 53, 110–118]. However, several investigators did notice an elevation in the residual amount of unrepaired strand breaks in irradiated A-T cells and suggested that these cells might handle the DNA strand break differently from other cells [119–122]. Cytogenetic studies also pointed to a residual amount of double-strand breaks unrepaired in A-T [112], or a higher fraction of double-strand breaks converted to chromosomal breaks [123–125]. A defect in chromatin structure was suggested to be responsible for this phenomenon [126]. Several studies pointed specifically to possible misrejoining of DNA breaks in damaged molecules introduced into A-T cells [120, 127–132] or treated with A-T cellular extracts [51, 133–135]. A recombination-based mechanism responsible for double-strand break repair and defective in A-T was suggested [51, 132] and related to in vivo observations of an increased rate of uncommon transrearrangements between T-cell receptor genes in A-T lymphocytes [136–138]. As expected, general V(D)J rejoining in A-T patients was found normal [139, 140]. Abnormal recombination in A-T was also noticed by Meyn [141], who observed an extremely high rate of intrachromosomal recombination in A-T cells, while interchromosomal recombination remained normal.
While the mechanism that directly handles the critical DNA lesion in A-T is still unclear, additional effects of the abnormality in this mechanism on downstream systems induced by DNA damage have been observed. The main observation consistently made in A-T cells is that several cell-cycle checkpoints activated by radiation damage do not function optimally. The postirradiation inhibition of the cell cycle that normally occurs at the G1 and S phases was found to be less pronounced in A-T cells, while the G2 phase in A-T cells irradiated at G1 or S was more prolonged than in normal cells. But, when A-T cells were irradiated at G2, the delay in traversing this stage into mitosis was shorter than in normal cells [102, 142–153]. Radiation-induced G1 arrest is mediated by a newly revealed signal transduction system involving a rise in the cellular level of the p53 protein, probably via a posttranslational control mechanism [153–156]. p53, in turn, induces the expression of several genes including Gadd45, p21WAF1/CIP1 and Mdm2 [155, 157–163]. The products of Gadd45 and p21WAF1/CIP1 inhibit DNA replication, while the Gadd45 protein also stimulates DNA repair [157, 159, 160, 164]. This system is perturbed in A-T cells of all complementation groups when stimulated by ionizing radiation or radiomimetic chemicals but functions normally after treatment with UV irradiation and other agents to which A-T cells are not sensitive [151, 152, 155, 164–166]. It has been proposed, therefore, that several signal transduction pathways activating p53 may be induced by specific types of DNA damage, with the one activated by strand breaks being defective in A-T cells [165, 167].
The p53 gene product mediates another radiation-induced pathway leading to programmed cell death (apoptosis) [168–170]. Meyn et al. [171] noticed that A-T cells sustain higher rates of apoptosis following irradiation, while disruption of p53 function increases their radioresistance. It was proposed that A-T cells have an unusually low threshold for triggering p53-mediated apoptosis and that their radiosensitivity stems from induction of apoptosis by usually nonlethal doses.
The DNA repair hypothesis and the DNA replication/cell cycle hypothesis could be merged by assuming that A-T cells may harbor a defect in a protein or a protein complex involved in both the initial processing of a specific DNA strand break and in triggering the signal transduction system leading to G1 arrest and enhancement of DNA repair. A defect in this system might result in unrepaired breaks leading to chromosome damage and possibly to the dominance of an error-prone repair system of lower fidelity which is usually overshadowed by the regular repair mechanism. The function of the downstream systems activated by that protein is reduced, with consequent cell cycle abnormalities. Another obvious possibility that cannot be ruled out at this point is a defect in a transcription factor responsible for the normal action of several genes controlling these systems.
Complementation Cloning Attempts: Too Many Genes Volunteering for the Same Job
The common approach to the identification of disease genes with yet unrecognized protein products is positional cloning [172, 173]. The localization of the A-T locus to chromosome 11q22–23 [174] opened the way to application of this approach to A-T (see below). But the cellular phenotype of A-T lends itself to another strategy for gene identification — functional cloning by complementation of the cellular phenotype.
Hypersensitivity to DNA-damaging agents, a prominent feature of A-T cells, calls for attempts to complement this phenotype by gene transfer. In this approach, exogenous DNA is introduced into the cells and selection applied to identify cell clones in which this sensitivity has been ‘corrected’. An attempt is then made to identify the piece of DNA supposedly responsible for this effect, which is expected to represent a normal allele of the disease gene. Functional cloning by this strategy is appealing since it circumvents the more labor-intensive positional cloning. Functional cloning proved particularly useful when rodent cell mutants sensitive to specifie radiations or chemicals were transfected with human genomic DNA or cDNA; these experiments led to isolation of a host of human genes involved in transcription and DNA repair, some of which are involved in specific complementation groups of the UV-sensitive disorders xeroderma pigmentosum, Cockayne’s syndrome and trichothiodystrophy [175]. Application of this strategy to human cells is experimentally more difficult but has been successful in isolating the genes for complementation groups A and C of xeroderma pigmentosum [176, 177] and Fanconi’s anemia group C [178].
Several laboratories have invested considerable effort during the last decade in applying this approach to A-T. Early trials to complement the radiosensitivity of A-T cells were based on transfection with genomic DNA. In this system, pieces of the exogenous DNA are expected to integrate into the cellular genome and be stably expressed. Lehmann et al. [179] and Green et al. [180] transfected the immortalized A-T(D) cell line AT5BIVA with human or mouse genomic DNA together with the selective marker gpt. One radioresistant cell clone was obtained out of 400,000 gpt+ transfectants and showed a normal level of radiation resistance but only partial correction of RDS. Attempts to rescue the responsible DNA fragment were unsuccessful. Lohrer et al. [181], using the same cell line in similar experiments, obtained no stably radioresistant transfectants. It was concluded that limiting factors, in particular the size of genomic DNA stably integrated into the genome of A-T cells, may prevent the success of experiments based on the use of such DNA.
Despite this discouraging conclusion, Kapp and Painter [182] transfected AT5BIVA cells with a genomic library in a cosmid vector and were able to identify a transfectant clone in which cellular and chromosomal radiosensitivities were corrected to an intermediate level, but RDS was retained. Subsequent isolation of the integrated DNA [183] revealed genomic sequences that mapped to the chromosomal band 11q23, to which the A-T locus had previously been linked [174]. A 3.0-kb cDNA clone corresponding to these DNA fragments was isolated and found to map near the THY-1 gene, now known to be located some 25 cM distal to the A-T locus (see below). This gene, termed ATDC (AT-D complementing), codes for 9 alternatively spliced transcripts with variable patterns of expression in different tissues [183, 184; Kapp, pers. commun.] and is not induced by ionizing radiation [184]. The ATDC protein contains several zinc finger motifs and a leucine zipper domain, indicating possible formation of a homo- or heterodimer involved in nucleic acid binding, typical of regulatory proteins [185]. Murnane et al. [186] used the two-hybrid system in yeast to demonstrate that this protein indeed forms homodimers. No mutations in this gene have been identified to date in A-T(D) cells [Kapp, pers. commun.].
In order to avoid the use of genomic DNA, Ziv et al. [187] chose to introduce a cDNA library cloned in the expression vector pCD [188] into the A-T(A) cell line AT22IJE-T [189], expecting the small size of the cDNA inserts to contribute to their stable integration and maintenance in the cellular genome. Out of 200,000 transfectants, 2 cell clones showed partial correction of radiomimetic sensitivity and RDS. Attempts to rescue the integrated cDNAs failed, however, probably due to dissociation between the inserts and the vector sequences used to identify the integrated DNA pieces.
While these results were not encouraging, the notion that phenotypic complementation can be obtained in A-T cells by gene transfer did gain support from experiments with another technique — the microcell-mediated chromosome transfer. In this system, whole chromosomes tagged with a selective marker are introduced into the cells, and the introduced genes residing in their ‘natural’ environment are expected to remain intact. The assignment of the A-T locus to chromosome 11q22-23 by Gatti et al. [174] spurred experiments with this approach. Ejima et al. [190] and Komatsu et al. [191] showed that introduction of chromosome 11 indeed restored cellular radioresistance in different A-T cell lines and that the responsible gene was distal to 11q14 [192]. Chromosomal radiosensitivity was complemented in other experiments [193], and aberrant derivatives of chromosome 11 enabled localization of the responsible gene to 11q23 [194]. Lambert et al. [195] used a chromosome 18 derivative containing translocated material from 11q22-23 to show that this chromosomal region indeed contained a gene that complements three phenotypic features of A-T(D) cells: radiomimetic sensitivity, RDS and the abnormal postirradiation cell cycle kinetics.
Functional cloning by transfer of human DNA into human cells has always suffered from two drawbacks: the low uptake of exogenous DNA by human cells and the difficulty of direct identification of the introduced DNA against the background of the recipient’s DNA. Both problems are largely eliminated when human DNA is introduced into rodent mutant cells that simulate the appropriate human mutations. In fact, most of the human genes involved in UV response identified using gene transfer were rescued from UV-sensitive rodent cells transfected with normal human DNA [175]. An extensive collection of X-ray-sensitive hamster mutants has been obtained in several laboratories [196]. Thacker and Ganesh [197] identified the mutant irs-2 obtained from V70 hamster cells, showing the same sensitivity profile and RDS like A-T cells, with no apparent defect in strand breakage repair [198]. Zdzienicka et al. [199] identified 3 V79 mutants with spontaneous chromosomal breakage, cellular and chromosomal radiosensitivity, RDS and normal strand break repair. Their sensitivity profile with regard to other DNA-damaging agents was remarkably similar to that of A-T cells [200]. Interestingly, all 3 mutants belong to the same complementation group as the irs-2 mutant [201]. Fusion of these cells with human HeLa cells resulted in full complementation of RDS [202]. However, introduction of human chromosome 11 containing an intact 11q22-23 region did not result in correction of the mutant phenotype, while the same chromosome complemented the phenotype of the A-T cell line AT5BIVA cells [203]. It was concluded that this phenotype, which so remarkably resembled the human A-T defect, was caused in the hamster cells by a gene unrelated to the human A-T gene [203]. Transfection of these mutants with HeLa genomic DNA or a human genomic cosmid library yielded transfectants that gained some radioresistance but retained RDS [204]. Microcell-mediated chromosome transfer of additional human chromosomes into these mutants showed that intermediate X-ray sensitivity could be conferred by human chromosomes 4 and 15, together with a mouse chromosome. The combination containing human chromosome 4 also fully complemented RDS [204]. These studies underscored again the dissociation between radiosensitivity and radioresistant DNA synthesis, and the genetic complexity underlying these two biological end points.
Chromosome-mediated gene transfer is advantageous as it facilitates separation between the recipient’s genome and the introduced genes. It has the drawback, however, of focussing on a chromosomal region rather than on a discrete gene. A system based on episomal expression vectors that replicate in the cells as extrachromosomal elements allows the introduction and expression of individual cDNAs without their chromosomal integration. These vectors are easy to rescue and are attractive for use with A-T cells because they are presumed less vulnerable to the inherent genomic instability of these cells. Commonly used episomal vectors contain the origin of replication and EBNA-1 antigen gene of the Epstein-Barr virus. The binding of EBNA-1 protein to the origin of replication sequence enables episomal replication of the plasmid [205–208].
Three laboratories have recently used episomal cloning systems to identify cDNAs complementing the sensitivity phenotype of A-T cells of group D [209], group A [210] and group E [211], with remarkably similar results. In all cases, stable transfectants which had acquired various degrees of radiomimetic resistance were obtained, and episomal cDNAs were rescued and identified. A total of 26 cDNA clones were obtained in these 3 studies and found to confer different degrees of resistance to ionizing radiation or radiomimetic drugs upon repeated transfection. However, in the study of Ziv et al. [210], only 1 of 13 cDNA fragments that complemented the radiomimetic sensitivity of A-T(A) cells also partly corrected RDS. These results and previous observations point to the possibility of separating between these two features of the A-T phenotype [59, 109, 110, 182, 212].
An intriguing finding of these studies was that many of the complementing cDNAs were not full length and some represented only the 3′ untranslated regions of the corresponding cDNAs [211]. Chen et al. [211] suggested that 3′ untranslated regions of certain cDNAs may be able to modify the cellular response to radiomimetic agents by unknown regulatory mechanisms. This assumption implies that the phenotypic complementation system as a tool for studying the molecular basis of certain diseases may be highly prone to background noise, at least in the case of A-T. Indeed, all the cDNAs identified in these studies represented a large variety of genes: many were previously known, and none mapped to the A-T locus at 11q22-23. Several of the previously known genes, like phospholipase A2 (obtained independently by Ziv et al. [210] and Chen et al. [211]), and heat shock cognate protein 70 [210] are involved in various stress responses. The role of others, such as ferritin H chain, cytochrome C and ribosomal proteins [210], in cellular responses to radiomimetic sensitivity is unclear.
These studies led to the conclusion that the biological end points that define the two A-T phenotypic hallmarks — radiosensitivity and RDS — can be modulated by high expression of a number of sequences, not necessarily full-length transcripts. In such a situation complementation cloning may suffer from a low signal-to-noise ratio, since some of these cDNAs may even mask the effect of a clone derived from the disease gene itself. In view of these results, attempts to identify the elusive A-T genes shifted recently to positional cloning. Phenotypic complementation may still be helpful in testing the authenticity of a candidate gene obtained by that approach.
Positional Cloning: Zeroing in on the Culprit Genes
The well-established positional cloning paradigm [172, 173] has recently had impressive success with scores of disease genes, including some high-profile ones [173, 213–218]. The basic steps in this strategy include: the localization of a disease locus to a specific chromosomal region by linkage analysis; extensive generation of highly polymorphic markers in the region and narrowing the locus by genetic analysis; long-range cloning and physical mapping of the disease locus; identification of transcribed sequences (‘gene hunting’), and, finally, a search of the candidate genes for mutations in patients. Attempts at positional cloning of the A-T genes have recently culminated in the construction of extensive transcript maps of the A-T locus and a search for mutations in a fair number of candidate genes.
The genetic heterogeneity of A-T presents a potential obstacle to linkage analysis should several A-T genes reside in different locations. This has proved to be the case in xeroderma pigmentosum and Fanconi’s anemia [175, 219]. Gatti et al. [174] conveniently skirted this problem by conducting initial linkage analysis on a 61-member Amish A-T kinship assigned to complementation group A. The first marker that gave a lod score suggestive of linkage with A-T was THY-1 localized on chromosome 11, region 11q22-23 (fig. 1a). Further analysis with additional markers at this region and additional group A families substantiated this finding, and other unassigned A-T families appeared to make the 11q22-23 localization conclusive. This milestone in A-T research opened the way to positional cloning efforts. Additional studies on American, Turkish, British, Israeli and French families [220–225] clearly showed that a major A-T locus containing the group A gene resided at 11q22–23 but proximal to THY-1. These studies utilized increasing numbers of restriction fragment length polymorphism markers and refined genetic and physical maps of this region [222, 226–230] and did not indicate locus heterogeneity, thus pointing to a possible single A-T locus at 11q22–23. This assumption gained significant support from a linkage study conducted by Ziv et al. [231] with a single Moroccan Jewish A-T family assigned to group C. Significant lod scores obtained with 11q22–23 markers indicated the close proximity of the A-T(A) and A-T(C) mutations. A consortiumbased analysis of 111 families from the United States, Turkey, England, Italy and Israel narrowed the disease locus to an 8-cM sex-averaged interval between the markers STMY and D11S132/NCAM [232] (fig. 1b). McConville et al. [233] later suggested that the group E locus may also be located in this interval, which probably spans the 11q22.3–23.1 boundary.
Positional cloning of the A-T genes. Assignment of the A-T locus to chromosome 11q22–23 [174] (a) has led to a primary linkage map of the A-T region and confinement of the A-T locus within an 8-cM interval [232] (b). Subsequent construction of a high-density microsatellite map of the region [242] and repeated-linkage analysis [245, 246] determined an interval between D11S1818 and D11S1819 probably containing the mutations for all 4 complementation groups of A-T (c). According to Vanagaite et al. [242] this interval spans about 1.5 Mb of DNA. Long-range cloning in yeast artificial chromosomes [254] (d) and cosmids (e) now enables systematic gene hunting in this region.
In the absence of a chromosomal aberration flagging the physical location of the disease gene, repeated genetic analysis was required to narrow the A-T search interval. McConville et al. [233] and Ambrose et al. [234] used a rapid method to generate biallelic markers to identify single-strand conformation polymorphisms in DNA fragments isolated from yeast artificial chromosome clones at the A-T region. With these markers they were able to narrow the A-T locus, first to 4 cM between D11S611 and D11S1897 [233] and later to 3 cM between GRIA4 and D11S1897 [234] (fig. 1c). (D11S1897 is a new locus number assigned to a marker formerly included in the locus D11S535.) A physical framework for the A-T region was emerging in parallel from extensive radiation hybrid maps of 11q with particular emphasis on 11q22–23 [235–238] and detailed pulsed-field maps of the A-T region [231, 239]. Integration of the genetic and physical maps [234, 240] narrowed the A-T locus to 3 Mb of DNA. It should be noted that A-T has never recombined with two markers within this interval, D11S384 and D11S535 (fig. 1c).
Genetic analysis of the A-T region has been based up to this point on restriction fragment length polymorphisms and single-strand conformation polymorphism markers. Such markers, usually biallelic, have a limited polymorphic information content and may potentially miss genetic information in noninformative families. Thus, the derivation and mapping of microsatellite markers at the A-T region was considered essential for further narrowing the A-T locus. A high-density microsatellite map of the A-T region was constructed by Vanagaite et al. [241, 242] and Rotman et al. [243,244] in two stages: microsatellite markers generated at random by several mapping centers and generally mapped to the distal part of 11q were physically mapped within the A-T region [241, 242], and novel markers based on polymorphic CA repeats were generated [243, 244]. A map containing 24 microsatellite markers was constructed across 6 Mb containing and flanking the A-T interval [242] (fig. 1c).
Most of these markers were used in a consortium-based study of 176 A-T families in laboratories in the United States, England and Israel [245, 246]. Lange et al. [246] drew up a comprehensive 20-point linkage map based on the A-T families and 59 CEPH families. Using a Monte Carlo linkage algorithm [247, 248], Lange et al. [246] showed that the peak of the A-T location score was under the marker D11S535, with a 2-lod support interval for A-T between D111819 and D11S1294 (fig. 1c). Haplotype analysis [245, 246] disclosed 4 recombinants which placed A-T distal to D11S1819, and 1 loss of homozygosity in patients from an inbred family, which pointed to an A-T gene proximal to D11S1818. No recombinants were found between A-T and markers in the region D11S384-D11S1294 (fig. 1c). These studies reinforced the notion that the 4 complementation groups in classical A-T are probably determined by mutations at the 11q22–23 A-T locus and may converge to the D11S1818-D11S1819 interval. According to Vanagaite et al. [242] this interval spans about 1.5 Mb (fig. 1c).
Although the evidence for a major A-T locus at D11S1818-D11S1819 is compelling, being based on a significant number of families from various ethnic groups and complementation groups, A-T families have been found that do not link to this region. In the study of Lange et al. [246], 7 out of 176 families did not show linkage to this region: 6 of them had single affected subjects that could represent new mutations, and 1 family had 2 affected individuals who shared the same haplotypes with a normal sibling. In a British A-T family, haplotype analysis in 2 affected cousins and their siblings showed no linkage to 11q22-23 [62]. The patients in this family represented a clinical variant with slower progression of the disease but otherwise had the typical clinical and cellular phenotype of A-T. Such families may represent still another locus involved in the disease in very rare cases.
Linkage analysis typically reaches its limit of resolution when the disease locus is reduced to 1–2 cM and a final number of recombinants defines its boundaries with no additional recombinants found betlcsn them. The locus can be further refined by determining the extent of linkage disequilibrium between the disease and various markers at the locus. A significant degree of allelic association between the disease and a particular marker is usually indicative of physical proximity. The degree of disequilibrium reflects the extent of recombination that has occurred between certain mutations and neighboring markers since these mutations first appeared. This analysis is especially powerful in isolated ethnic communities with a limited number of mutations. A clearly defined core haplotype retained around a particular mutation over the years may help reduce the disease locus [for examples, see 249–251].
Oskato et al. [252] noticed in Moroccan Jewish patients a significant degree of linkage disequilibrium between A-T and a series of markers extending over a wide region, between D11S384 and D11S424 (the latter is distal to DHS1647, fig. 1c). This finding was compatible with a strong founder effect in this community, which has been isolated from the rest of the Jewish people and the surrounding population for many generations. Vanagaite et al. [in preparation] used the microsatellite markers recently added to the map [242] (fig. 1c) and obtained a disequilibrium peak extending between D11S384 and D11S2105 in 16 Moroccan Jewish families. Since several Moroccan Jewish patients have been assigned to complementation group C [81; unpubl. observations], this may indicate that the ATC gene is located at the distal half of the A-T interval. A similar study by Uhrhammer et al. [253] on 27 Costa Rican A-T families showed moderate linkage disequilibrium with several markers in the D11S1816-D11S1300 interval, which did not enable reduction of the disease locus. No significant allelic association between A-T and specific markers was found by Vanagaite et al. [in preparation] in Turkish and Italian families, probably indicating a relatively large number of mutations in these populations. An intriguing allelic association was noticed by Taylor et al. [65] in 10 of 60 British A-T families in which the patients have a clinical variant of A-T, with later onset and lower chromosomal radiosensitivity than most of A-T patients. One of the chromosomes in each of these patients has a specific haplotype at the D11S1819-D11S1817 region, extending proximal to the A-T locus defined by linkage analysis (fig. 1c). It is not clear yet whether this finding flags a gene involved in the disease in a subset of patients who might harbor a mutation with a milder phenotypic effect.
In the course of these studies, as the boundaries of the A-T locus moved closer together, systematic cloning of the region in genomic contigs became essential. Rotman et al. [254, 255] constructed a yeast artificial chromosome contig encompassing the current A-T locus (fig. 1d), and similar contigs have been constructed in other laboratories involved in positional cloning of the A-T gene(s) [James, MR; McConville, CM; Gatti, RA; Concannon, P, pers. commun.]. Cosmid contigs encompassing various portions of the current A-T interval have been constructed in several laboratories (see fig. 1e for example) and used in the next step in the positional cloning scheme, gene hunting.
Gene hunting typically relies on two commonly used methods to identify transcribed sequences in genomic DNA: direct selection based on hybridization of genomic DNA with cDNA collections of various sources [256, 257] and exon trapping which identifies and clones exon sequences by virtue of their splicing capacity [258]. Transcription maps of the A-T locus indicate that this genomic region is gene rich and contains a high density of transcribed sequences, most of which are new [259; Bar-Shira et al., in preparation]. This wealth of genes within the A-T locus makes the search for mutations in patients an experimental challenge.
Conclusion
Decades of intensive research have provided numerous clues to the nature of the defect in A-T, but this defect might not be delineated at the molecular level before the culprit gene(s) are cloned. Until then, the disease remains largely a mystery, as it has been ‘from the start’ [1]. Localization of the A-T locus to chromosome 11q22-23 and subsequent application of positional cloning have, however, brought A-T researchers closer than ever to unraveling this mystery.
References
Boder E: Ataxia-telangiectasia: An overview: in Gatti RA, Swift M (eds): Ataxia-Telangiectasia: Genetics, Neuropathology and Immunology of a Degenerative Disease of Childhood. New York, Liss, 1985, pp 1–63.
Boder E, Sedgwick RP: Ataxia-telangiectasia: A familial syndrome of progressive cerebellar ataxia, oculocutaneous telangiectasia and frequent pulmonary infections. A preliminary report on 7 children, an autopsy, and a case history. USC Med Bull 1957;9:15–28
McKinnon PJ: Ataxia-telangiectasia: An inherited disorder of ionizing radiation sensitivity in man. Hum Genet 1987;75:197–208
Sedgwick RP, Boder E: Ataxia-telangiectasia; in Vinken PJ, Bruyn GW. Klawans HL (eds): Handbook of Clinical Neurology. Amsterdam, Elsevier Scientific Publishers, 1991, vol 16, pp 347–423.
Gatti RA, Boder E, Vinter HV, Sparkes RS, Norman A, Lange K: Ataxia-telangiectasia: An interdisciplinary approach to pathogenesis. Medicine 1991;70:99–117
Taylor AMR, Byrd PJ, McConville CM, Thacker S: Genetic and cellular features of ataxia telangiectasia. Int J Radiat Biol 1994;65:65–70
Harnden DG: The nature of ataxiatelangiectasia: Problems and perspectives. Int J Radiat Biol 1994,66: S13–S19.
Bundey S: Clinical and genetic features of ataxia-telangiectasia. Int J Radiat Biol 1994;66:S23–S29
Swift M, Morell D, Cromartie E, Chamberlin AR: The incidence and gene frequency of ataxia telangiectasia in the United States. Am J Hum Genet 1986;39:573–583
Swift M, Heim RA, Lench NJ: Genetic aspects of ataxia telangiectasia. Adv Neurol 1993;61:115–125
Pippard EC, Hall AJ, Barker DJP, Bridges BA: Cancer in homozygotes and heterozygotes of ataxia telangiectasia and xeroderma pigmentosum in Britain. Cancer Res 1988;48:2929–2932
Woods CG, Bundey SE, Taylor AMR: Unusual features in the inheritance of ataxia telangiectasia. Hum Genet 1990,84:555–562.
Sanal O, Berkel AI, Ersoy F, Tezcan I, Topaglu H: Clinical variants of ataxia-telangiectasia; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Berlin, Springer, 1993, vol H77, pp 183–189.
Chessa L, Lisa A, Fioram O, Zei G: Ataxia-telangiectasia in Italy: Genetic analysis. Int J Radiat Biol 1994;66:S31–S33
Levin S, Gottfried E, Cohen M: Ataxia telangiectasia: A review — With observations on 47 Israeli cases. Paediatrician 1977,6:135–146.
Morrell D, Chase CL, Swift M: Cancers in 44 families with ataxia-telangiectasia. Cancer Genet Cytogenet 1990;50:119–123
Hecht F, Heckt BK: Cancer in ataxia-telangiectasia patients. Cancer Genet Cytogenet 1990;46:9–19
Peterson RDA, Funkhouser JD, Tuck-Muller CM, Gatti RA: Cancer susceptibility in ataxia-telangiectasia. Leukemia 1992;6 supp11:8–13
Gotoff SP, Amirmokri E, Liebner EJ: Ataxia-telangiectasia. Am J Dis Child 1967;114:617–625
Morgan JL, Holcomb TM, Morrisey RW: Radiation reaction in ataxiatelangiectasia. Am J Dis Child 1968;116:557–558
Cunliffe PN, Mann JR, Cameron AH, Roberts KD: Radiosensitivity in ataxia telangiectasia. Br J Radiol 1975;48:374–376
Pritchard J, Sandland MR, Breathnach FB, Pincott JR, Cox R, Husband P: The effects of radiation therapy for Hodgkin’s disease in a child with ataxia telangiectasia. Cancer 1982;50:877–886
Abadir R, Hakami N: Ataxia telangiectasia with cancer: An indication for reduced radiotherapy and chemotherapy doses. Br J Radiol 1983;56:343–345
Hart RM, Bruce Kimler BF, Evans RG, Park CH: Radiotherapeutic management of medulloblastoma in a pediatric patient with ataxia telangiectasia. Int J Radiat Oncol Biol Phys 1987;13:1237–1240
Taylor AMR, Oxford JM, Metcalfe JA: Spontaneous cytogenetic abnormalities in lymphocytes from thirteen patients with ataxia telangiectasia. Int J Cancer 1981;27:311–319
Taylor AMR: Cytogenetics in ataxia-telangiectasia; in Bridges BA, Harnden DG (eds): Ataxia-Telangiectasia — A Cellular and Molecular Link between Cancer, Neuropathology and Immune Deficiency. Chichester, Wiley & Sons, 1982, pp 53–81.
Kojis TL, Gatti RA, Sparkes RS: The cytogenetics of ataxia telangiectasia. Cancer Genet Cytogenet 1991;56:143–156
Taylor AMR, Butterworth SV: Clonal evolution of T-cell chronic lymphocytic leukemia in a patient with ataxia-telangiectasia. Int J Cancer 1986;37:511–516
Aurias A, Croquette MF, Nuyts JP, Gnscelli C, Dutrillaux B: New data on clonal anomalies of chromosome 14 in ataxia telangiectasia: tct(14;14) and inv(14). Hum Genet 1986;72:22–24
Heppel A, Butterworth SV, Hollis RJ, Kennaugh AA, Beatty DW, Taylor AMR: Breakage of the T cell receptor a chain locus in non-malignant clones from patients with ataxia-telangiectasia. Hum Genet 1988;79:360–364
Russo G, Isobe M, Pegoraro L, Finan J, Nowell PC, Croce M: Molecular analysis of a t(7;14)(q35;32) chromosome translocation in a patient with ataxia-telangiectasia. Cell 1988;53:137–144
Stern MH, Lipkowitz S, Aurias A, Griscelli C, Thomas G, Kirsch IR: Inversion of chromosome 7 in ataxia telangiectasia is generated by a rearrangement between T-cell receptor β and T-cell receptor τ genes. Blood 1989;6:2076–2080
Metcalfe JA, Heppel-Parton A, McConville CM, Taylor AMR: Characterization of a B-lymphocyte t(2;14)(p11;q32) translocation from an ataxia telangiectasia patient conferring proliferative advantage on cells in vitro. Cytogenet Cell Genet 1991;56:91–98
Taylor AMR, Lowe PA, Stacey M, Thick J, Campbell L, Beatty D, Biggs P, Formstone CJ: Development of T-cell leukemia in an ataxia telangiectasia patient following clonal selection in t(X;14)-containmg lymphocytes. Leukemia 1992;6:961–966
Thick J, Sherrington PD, Fisch P, Taylor AMR, Rabbits TH: Molecular analysis of a new translocation, t(X;14)(q28;q11), in a premalignancy and in leukaemia associated with ataxia telangiectasia. Genes Chromosomes Cancer 1992;5:321–325
Sherrington PD, Fisch P, Taylor AMR, Rabbits TH: Clonal evolution of malignant and non-malignant T cells carrying t(14; 14) and t(X;14) in patients with ataxia telangiectasia. Oncogene 1994,9:2377–2381.
Shiloh Y, Tabor E, Becker Y: In vitro phenotype of A-T fibroblast strains: Clues to the nature of the ‘A-T DNA lesion’ and the molecular defect in A-T; in Gatti RA, Swift M (eds): Ataxia-Telangiectasia: Genetics, Neuropathology and Immunology of a Degenerative Disease of Childhood. New York, Liss, 1985, pp 111–121.
Lavin MF: Biochemical defects in ataxia-telangiectasia; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Berlin, Springer, 1993, vol H77, pp 235–255.
Lehmann AR: The cellular and molecular responses of ataxia-telangiectasia cells to DNA damage; in Bridges BA, Harnden DG (eds): Ataxia-Telangiectasia — A Cellular and Molecular Link between Cancer, Neuropathology and Immune Deficiency. Chichester, Wiley & Sons, 1982, pp 83–102.
Shiloh Y, Tabor E, Becker Y: Abnormal response of ataxia-telangiectasia cells to agents that break the deoxyribose moiety of DNA via a targeted free radical mechanism. Carcinogenesis 1983;4:1317–1322
Shiloh Y, Tabor E, Becker Y: Cells from patients with ataxia-telangiectasia are abnormally sensitive to the cytotoxic effect of a tumor promoter, phorbol-12-myristate-13-acetate. Mutat Res 1984;149:283–286
Henner WD, Blazka ME: Hypersensitivity of cultured ataxia-telangiectasia cells to etoposide. J Natl Cancer Inst 1986;76:1007–1011
Smith PJ, Makinson TA, Watson JV: Enhanced sensitivity to camptothecin in ataxia-telangiectasia cells and its relationship with expression of DNA topoisomerase I. Int J Radiat Biol 1989;555:217–231
Taylor AMR, Metcalfe JA, McConville C: Increased radiosensitivity and the basic defect in ataxia telangiectasia. Int J Radiat Biol 1989;56:677–684
Sullivan N, Lyne L: Sensitivity of fibroblasts derived from ataxia-telangiectasia patients to calicheamicin τ1. Mutat Res 1990,245:171–175.
Houldsworth J, Cohen D, Singh S, Lavin MF: The response of ataxiatelangiectasia lymphoblastoid cells to neutron irradiation. Radiat Res 1991;125:277–282
Mirzayans R, Paterson MC: Lack of correlation between hypersensitivity to cell killing and impaired inhibition of DNA synthesis in ataxia telangiectasia fibroblasts treated with 4-nitroquinoline 1-oxide. Carcinogenesis 1991;12:19–24
Caporossi D, Porfirio B, Nicoletti B, Pahtti F, Degrassi F, De Salvia R, Tanzarella C: Hypersensitivity of lymphoblastoid lines derived from ataxia telangiectasia patients to the induction of chromosomal aberrations by etoposide (VP-16). Mutat Res 1993;290:265–272
Painter RB: Radiobiology of ataxiatelangiectasia; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Berlin, Springer, 1993, vol H77, pp 257–268.
Ward AJ, Olive PL, Burr AH, Rosin MP: Response of fibroblast cultures from ataxia-telangiectasia patients to reactive oxygen species generated during inflammatory reactions. Environ Mol Mutagen 1994;24:103–111
Thacker J: Cellular radiosensitivity in ataxia-telangiectasia. Int J Radiat Biol 1994;66:S87–S96
van der Schans GP, Paterson MC, Gross WG: DNA strand break and rejoining in cultured human fibroblasts exposed to fast neutrons or gamma rays. Int J Radiat Biol 1983;44:75–85
Shiloh Y, van der Schans GP, Lohman PHM, Becker Y: Induction and repair of DNA damage in normal and ataxia-telangiectasia fibroblasts treated with neocarzinostatin. Carcinogenesis 1983;4:917–921
Painter RB, Young BR: Radiosensitivity in ataxia telangiectasia: A new explanation. Proc Natl Acad Sci USA 1980;77:7315–7317
Houlsworth J, Lavin MF: Effect of ionizing radiation on DNA synthesis in ataxia telangiectasia cells. Nucleic Acids Res 1980;8:3709–3720
Painter RB: Altered DNA synthesis in irradiated and unirradiated ataxia-telangiectasia cells; in Gatti RA, Swift M (eds): Ataxia-Telangiectasia: Genetics, Neuropathology and Immunology of a Degenerative Disease of Childhood. New York, Liss, 1985, pp 89–100.
de Jong J, Tijssen CC: Ataxia telangiectasia in a brother and sister at older age. Clin Neurol Neurosurg 1988;90:279–281
Fiorilli M, Antonelli A, Russo G, Goescenzi M, Carbonari M, Petrinelli P: Variant of ataxia-telangiectasia with low-level radiosensitivity. Hum Genet 1985;70:274–277
Taylor AM, Flude E, Laher B, Stacey M, McKay E, Wett J, Green SH, Harding AE: Variant forms of ataxia-telangiectasia. J Med Genet 1987;24:669–677
Ziv Y, Amiel A, Jaspers NGJ, Berkel AI, Shiloh Y: Ataxia telangiectasia: A variant with altered in vitro phenotype of fibroblast cells. Mutat Res 1989;210:211–219
Chessa L, Petrinelli P, Antonelli A, Fiorilli R: Heterogeneity in ataxia telangiectasia classical phenotype associated with intermediate radiosensitivity. Am J Med Genet 1992;42:741–746
Hernandez D, McConville CM, Stacey M, Woods CG, Brown MM, Shutt P, Rysiecki G, Taylor AMR: A family showing no evidence of linkage between the ataxia telangiectasia gene and chromosome 11q22-23. J Med Genet 1993;30:135–140
Taylor AMR, McConville CM, Woods CG, Byrd PJ, Hernandez D: Clinical and cellular heterogeneity in ataxia telangiectasia; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Springer, Berlin, 1993, vol H77, pp 209–234.
Lange E, Gatti RA, Sobel E, Concannon P, Lange K: How many A-T genes?; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Springer, Berlin, 1993, vol H77, pp 37–54.
Taylor AMR, Rotman G, Shiloh Y, Byrd PJ, McConville CM: A haplotype common to intermediate radiosensitivity variants of ataxia-telangiectasia. Int J Radiat Biol 1994;66:S35–S41
Ying KL, Docteau WE: Cytogenetic anomalies in a patient with ataxia, immune deficiency, and high alpha-fetoprotein in the absence of telangiectasia. Cancer Genet Cytogenet 1981;4:311–317
Maserati E, Ottolini A, Veggiotti A, Lanzi G, Pasquali F: Ataxia-without-telangiectasia in two sisters with rearrangements of chromosomes 7 and 14. Clin Genet 1988;34:283–287
Byrne E, Hallpike JF, Manson JI, Sutherland GR, Thong YH: Ataxiawithout-telangiectasia: Progressive multisystem degeneration with IgE deficiency and chromosomal instability. J Neurol Sci 1984;66:307–317
Weemaes CMR, Huustinx TWJ, Scheres JMJC, van Munster PJJ, Bakkeren JAJM, Taalman RDFM: A new chromosomal instability disorder: The Nijmegen breakage syndrome. Arch Paediatr Scand 1981;70:557–564
Taalman RDFM, Jaspers NGJ, Scheres JMJC, de Wit J, Hustinx TWJ: Hypersensitivity to ionizing radiation in vitro, a new chromosomal breakage disorder, the Nijmegen breakage syndrome. Mutat Res 1983;112:23–32
Taalman RDFM, Hustinx TWJ, Weemaes CRM, Seemanova E, Schmidt A, Passarge E, Scheres JMJC: Further delineation of the Nijmegen breakage syndrome. Am J Med Genet 1989;32:425–431
Seemanova E, Passarge E, Benesova J, Houstek J, Kasal P, Sevcikova M: Familial microcephaly with normal intelligence, immunodeficiency and risk for lymphoreticuler malignancies: A new autosomal recessive disorder. Am J Med Genet 1985;20:639–648
Seemanova E: An increased risk for malignant neoplasms in heterozygotes for a syndrome of microcephaly, normal intelligence, growth retardation, remarkable facies, immunodeficiency and chromosomal instability. Mutat Res 1990;238:321–324
Conley ME, Spinner NB, Emanuel BS, Nowell PC, Nichols WW: A chromosomal breakage syndrome with profound immunodeficiency. Blood 1986;67:1251–1256
Jaspers NGJ, Taalman RDFM, Baan C: Patients with an inherited syndrome characterized by immunodeficiency, microcephaly, and chromosomal instability: Genetic relationship to ataxia telangiectasia. Am J Hum Genet 1988;42:66–73
Wegner RD, Metzger M, Hanefeld F, Jaspers NGJ, Baan C, Magdorf K, Kunze J, Sperling K: A new chromosomal instability disorder confirmed by complementation studies. Clin Genet 1988;33:20–32
Weemaes CMR, Smeets DFCM, Burgt CJAM: Nijmegen breakage syndrome: A progress report. Int J Radiat Biol 1994;66:S185–S188
Curry CJR, O’Lague P, Tsai J, Hutchinson HT, Jaspers NGJ, Wara D, Gatti RA: ATFresno: A phenotype linking ataxia-telangiectasia with the Nijmegen breakage syndrome. Am J Hum Gent 1989;45:270–275
Murnane JP, Painter RB: Complementation of the defects in DNA synthesis in irradiated and unirradiated ataxia-telangiectasia cells. Proc Natl Acad Sci USA 1982;79:1960–1963
Jaspers NGJ, Bootsma D: Genetic heterogeneity in ataxia-telangiectasia studied by cell fusion. Proc Natl Acad Sci USA 1982;79:2641–2644
Jaspers NGJ, Gatti RA, Baan C, Linssen PCML, Bootsma D: Genetic complementation analysis of ataxia telangiectasia and Nijmegen breakage syndrome: A survey of 50 patients. Cytogenet Cell Genet 1988;49:259–263
Chen P, Imray FP, Kidson C: Gene dosage and complementation analysis of ataxia telangiectasia lymphoblastoid cell lines assayed by induced chromosome aberrations. Mutat Res 1984;129:165–172
Chen P, Kidson C: Complementation analysis in ataxia telangiectasia: Fibroblast-lymphoblastoid cell heterokaryon assay by radiation-induced chromosome aberrations. Cytogenet Cell Genet 1993;64:9–11
Swift M, Reitnauer PJ, Morrell D, Chase CL: Breast and other cancers in families with ataxia-telangiectasia. N Engl J Med 1987;316:1289–1294
Swift M, Chase CL, Morrell D: Cancer predisposition of ataxia-telangiectasia heterozygotes. Cancer Genet Cytogenet 1990;46:21–27
Swift M, Morrell D, Massay RB, Chase CL: Incidence of cancer in 161 families affected by ataxia-telangiectasia. N Engl J Med 1991;325:1831–1836
Borreson AL, Anderson TI, Tretli S, Heiberg A, Moller P: Breast cancer and other cancers in Norwegian families with ataxia-telangiectasia. Genes Chromosomes Cancer 1990;2:339–340
Easton DF: Cancer risks in A-T heterozygotes. Int J Radiat Biol 1994;66:S177–S182
Chen PC, Lavin MF, Kidson C, Moss D: Identification of ataxia telangiectasia heterozygotes, a cancer prone population. Nature 1978;274:484–486
Kidson C, Chen P, Imray P: Ataxiatelangiectasia heterozygotes: Dominant expression of ionizing radiation sensitive mutants; in Bridges BA, Harnden DG (eds): Ataxia-Telangiectasia — A Cellular and Molecular Link between Cancer, Neuropathology and Immune Deficiency. Chichester, Wiley & Sons, 1982, pp 363–372.
Shiloh Y, Tabor E, Becker Y: The response of ataxia-telangiectasia homozygous and heterozygous skin fibroblasts to neocarzinostatin. Carcinogenesis 1982;3:815–820
Paterson MC, MacFarlane SJ, Gentner N, Smith BP: Cellular hypersensitivity to chronic γ-radiation in cultured fibroblasts from ataxiatelangiectasia heterozygotes; in Gatti RA, Swift M (eds): Ataxia-Telangiectasia: Genetics, Neuropathology and Immunology of a Degenerative Disease of Childhood. New York, Liss, 1985, pp 73–87.
Arlett CF, Priestley A: An assessment of the radiosensitivity of ataxia-telangiectasia heterozygotes; in Gatti RA, Swift M (eds): Ataxia-Telangiectasia: Genetics, Neuropathology and Immunology of a Degenerative Disease of Childhood. New York, Liss, 1985, pp 101–109.
Parshad R, Sanford KK, Jones GM, Tarone R: G2 chromosomal radiosensitivity of ataxia-telangiectasia heterozygotes. Cancer Genet Cytogenet 1985;14:163–168
Shiloh Y, Parshad R, Sanford KK, Jones GM: Carrier detection in ataxia-telangiectasia. Lancet 1986;i:689–690
Nagasawa H, Latt SA, Lalande ME, Little JB: Effects of X-irradiation on cell-cycle progression, induction of chromosomal aberrations and cell killing in ataxia-telangiectasia (AT) fibroblasts. Mutat Res 1985;148:71–82
Nagasawa H, Kraemer KH, Shiloh Y, Little JB: Detection of ataxia telangiectasia heterozygous cell lines by postirradiation cumulative labeling index: Measurements with coded samples. Cancer Res 1987;47:398–402
Rosin MP, Ochs HD, Gatti RA, Boder E: Heterogeneity of chromosomal breakage levels in epithelial tissues of ataxia-telangiectasia homozygotes and heterozygotes. Hum Genet 1989;83:133–138
Rudolph NS, Nagasawa H, Little JB, Latt SA: Identification of ataxiatelangiectasia heterozygotes by flow cytometric analysis of X-ray damage. Mutat Res 1989,211:19–29.
Waghray A, Al-Sedairy S, Ozand PT, Hannan MA: Cytogenetic characterization of ataxia telangiectasia (AT) heterozygotes using lymphoblastic cell lines and chronic γ-irradiation. Hum Genet 1990;84:532–534
Weeks DE, Paterson MC, Lange K, Andrais B, Davis RC, Yoder F, Gatti RA: Assessment of chronic gamma radiosensitivity as an in vitro assay for heterozygote identification of ataxia-telangiectasia. Radiat Res 1991;128:90–99
Lavin MF, Le Poidevin P, Bates P: Enhanced levels of radiation-induced g2 phase delay in ataxia telangiectasia heterozygotes. Cancer Genet Cytogenet 1992;60:183–187
Scott D, Jones LA, Elyan SAG, Spreadborough A, Cowan R, Ribiero G: Identification of A-T heterozygotes; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Berlin, Springer, 1993, Vol H77, pp 101–116.
Scott D, Spreadborough AR, Roberts SA: Radiation-induced G2 delay and spontaneous chromosome aberrations in ataxia-telangiectasia homozygotes and heterozygotes. Int J Radiat Biol 1994;66:S157–S163
Chen P, Farrell A, Hobson K, Girjes A, Lavin M: Comparative study of radiation-induced G2 phase delay and chromatid damage in families with ataxia-telangiectasia. Cancer Genet Cytogenet 1994;76:43–46
Pandita TK, Hittelman WN: Increased initial levels of chromosome damage and heterogeneous chromosome repair in ataxia telangiectasia heterozygote cells. Mutat Res 1994;310:1–13
Huo YK, Wang Z, Hong JH, Chessa L, McBride WH, Perlman SL, Gatti RA: Radiosensitivity of ataxia-telangiectasia, X-linked agammaglobulinemia and related syndromes using a modified colony survival assay. Cancer Res 1994;54:2544–2547
Naeim A, Repinski C, Huo Y, Hong HJ, Chessa L, Naeim F, Gatti RA: Ataxia-telangiectasia: Flow cytometric cell-cycle analysis of lymphoblastoid cell lines in G2/M before and after γ-irradiation. Mod Pathol 1994;7:587–592
Mirzayans R, Paterson MC: Lack of correlation between hypersensitivity to cell killing and impaired inhibition of DNA synthesis in ataxia telangiectasia fibroblasts treated with 4-nitroquinoline-1-oxide. Carcinogenesis 1991;12:19–24
Lavin MF, Schroeder AL: Damage-resistant DNA synthesis in eukaryotes. Mutat Res 1989;193:193–206
Sasaki MS, Taylor AMR: Dissociation between radioresistant DNA replication and chromosomal radiosensitivity in ataxia telangiectasia cells. Mutat Res 1994;307:107–113
Cornforth MN, Bedford JS: On the nature of a defect in cells from individuals with ataxia-telangiectasia. Science 1985:227:1589–1591.
Weichselbaum RR, Nove J, Little JB: Deficient recovery from potentially lethal radiation damage in ataxia telangiectasia and xeroderma pigmentosum. Nature 1978;271:261–263
Cox R, Masson WK, Weichselbaum RR, Nove J, Little JB: The repair of potentially lethal damage in X-irradiated cultures of normal and ataxia telangiectasia human fibroblasts. Int J Radiat Biol 1981;39:357–365
Shiloh Y, Tabor E, Becker Y: Repair of potentially lethal and sublethal damage induced by neocarzinostatin in normal and ataxia-telangiectasia skin fibroblasts. Biochem Biophys Res Commun 1983;110:483–490
Fornace AJ, Little JB: Normal repair of DNA single-strand breaks in patients with ataxia-telangiectasia. Biochim Biophys Acta 1980;607:432–437
Thierry D, Rigaud O, Duranton I, Moustacchi E, Magdelena H: Quantitative measurement of DNA strand breaks and repair in gamma-irradiated human leukocytes from normal and ataxiatelangiectasia donors. Radiat Res 1985;102:347–358
Radford IR, Hodgson GS: A comparison of the induction of DNA double-strand breakage and lethal lesions by X-irradiation in ataxia telangiectasia and normal fibroblasts. Radiat Res 1990;124:334–335
Coquerelle TM, Weibezahn KF, Lucke-Huhle: Rejoining of double strand breaks in normal human and ataxia-telangiectasia fibroblasts after exposure to 60Co γ-rays,241 Am α-particles or bleomycin. Int J Radiat Biol 1987;51:209–218
Debenham PG, Webb MBT, Jones NJ, Cox R: Molecular studies of the nature of the repair defect in ataxia-telangiectasia and their implications for cellular radiobiology. J Cell Sci 1987;6 suppl:177–189
Mozdarani H, Bryant PE: Induction and rejoining of chromatid breaks in X-irradiated A-T and normal human G2 fibroblasts. Int J Radiat Biol 1989;56:645–650
Blocher D, Sigut D, Hannan MA: Fibroblasts from ataxia-telangiectasia (A-T) and A-T heterozygotes show an enhanced level of residual DNA double-strand breaks after low dose-rate γ-irradiation assayed by pulsed-field gel electrophoresis. Int J Radiat Biol 1991:60:791–802.
Pandita TK, Hittelman WN: Initial chromosome damage but not DNA damage is greater in ataxia telangiectasia cells. Radiat Res 1992;130:94–103
Pandita TK Hittelman WN: The contribution of DNA and chromosome repair deficiencies to the radiosensitivity of ataxia-telangiectasia. Radiat Res 1992;131:214–223
Liu N, Bryant PE: Enhanced chromosomal response of ataxia-telangiectasia cells to specific types of DNA double-strand breaks. Int J Radiat Biol 1994;66:S115–S121
Hittelman WN, Pandita TK: Possible role of chromatin alteration in the radiosensitivity of ataxiatelangiectasia. Int J Radiat Biol 1994;66:S109–S113
Cox R, Debenham PG, Masson WK, Webb MBT: Ataxia-telangiectasia: A human mutation giving high frequency misrepair of double stranded scissions. Mol Biol Med 1986;3:229–244
Liu N, Bryant PE: Response of ataxia telangiectasia cells to restriction endonuclease induced DNA double-strand breaks. I. Cytogenetic characterization. Mutagenesis 1993;8:503–510
Thacker J: The use of integrating DNA vectors to analyse the molecular defects in ionising radiationsensitive mutants of mammalian cells including ataxia telangiectasia. Mutant Res 1989:220:187–204.
Runger TM, Poot M, Kraemer KH: Abnormal processing of transfected plasmid DNA in cells from patients with ataxia-telangiectasia. Mutat Res 1992;293:47–54
Tatsumi-Miyajima J, Takahashi Y, Takebe H: Analysis of mutations caused by DNA doublestrand breaks produced by a restriction enzyme in shuttle vector Plasmids propagated in ataxia telangiectasia cells. Mutat Res 1993;294:317–323
Powell S, Whitacker S, Peacock J, McMillan T: Ataxia telangiectasia: An investigation of the repair defect in the cell line AT5BIVA by plasmid reconstruction. Mutat Res 1993;294:9–20
North P, Ganesh A, Thacker J: The rejoining of double-strand breaks in DNA by human cell extracts. Nucleic Acids Res 1990;18:6205–6210
Thacker J, Chalk J, Ganesh A, North P: A mechanism for deletion formation in DNA by human cell extracts: The involvement of short sequence repeats. Nucleic Acids Res 1992;20:6183–6188
Ganesh A, North P, Thacker J: Repair and misrepair of site-specific DNA double-strand breaks by human cell extracts. Mutat Res 1993;299:251–259
Carbonari M, Cherchi M, Paganelli R, Giannini G, Galli E, Gaetano C, Papetti C, Fiorilli M: Relative increase of T cells expressing the gamma/delta rather than the alpha/beta receptor in ataxia-telangiectasia. N Engl J Med 1990;322:73–76
Kobayashi Y, Tycko B, Soreng AL, Sklar J: Transrearragements between antigen receptor genes in normal human lymphoid tissues and in ataxia telangiectasia. J Immunol 1991;147:3201–3209
Lipkowitz S, Stern MH, Kirsch IR. Hybrid T cell receptor genes formed by interlocus recombination in normal and ataxia-telangiectasia lymphocytes. J Exp Med 1990;172:409–418.
Hsieh CL, Arlett CF, Lieber MR: V(D)J recombination in ataxia telangiectasia, Bloom’s syndrome and a DNA ligase I-associated immunodeficiency disorder. J Biol Chem 1993;268:20105–20109
Kirsch IR: V(D)J recombination and ataxia-telangiectasia: A review. Int J Radiat Biol 1994;66:S97–S108
Meyn MS: High spontaneous intrachromosomal recombination rates in ataxia-telangiectasia. Science 1993;260:1327–1330
Zampetti-Bosseler F, Scott D: Cell death, chromosome damage and mitotic delay in normal human, ataxia telangiectasia and retinoblastoma fibroblasts after X-irradiation. Int J Radiat Biol 1981;39:547–558
Scott D, Zampetti-Bosseler F: Cell cycle dependence of mitotic delay in X-irradiated normal and ataxiatelangiectasia fibroblasts. Int J Radiat Biol 1982;42:679–683
Imray FP, Kidson C: Perturbations of cell cycle progression in gamma-irradiated ataxia-telangiectasia and Huntington’s disease cells detected by DNA flow cytometric analysis. Mutat Res 1983;112:369–382
Ford MD, Martin L, Lavm MF: The effects of ionizing radiation on cell cycle progression in ataxia telangiectasia. Mutat Res 1984;125:115–122
Nagasawa H, Latt SA, Lalande M, Little JB: Effects of X-irradiation on cell-cycle progression, induction of chromosomal aberrations and cell killing in ataxia telangiectasia (AT) fibroblasts. Mutat Res 1985;148:71–82
Smith PJ, Anderson CO, Watson JV: Abnormal retention of X-irradiation ataxia-telangiectasia fibroblasts in G2 phase of the cell cycle: Cellular RNA content, chromatin stability and the effect of 3-aminobenzamide. Int J Radiat Biol 1985;47:701–712
Bates P, Lavin MF: Comparison of gamma-irradiation-induced accumulation of ataxia-telangiectasia and control cells in G2 phase. Mutat Res 1985;218:165–170
Rudolph NS, Latt SA: Flow cytometric analysis of X-ray sensitivity in ataxia-telangiectasia. Mutat Res 1989;211:31–41
Beamish H, Lavin MF: Radiosensitivity in ataxia-telangiectasia: Anomalies in radiation-induced cell cycle delay. Int J Radiat Biol 1994;65:175–184
Hong JH, Gatti RA, Huo YK, Chiang CS, McBride WH: G2/M arrest and release in ataxia telangiectasia and normal cells after exposure to ionizing radiation. Radiat Res 1994;140:17–23
Beamish H, Khanna KK, Lavin MF: Ionizing radiation and cell cycle progression in ataxia telangiectasia. Radiat Res 1994;138:S130–S133
Kastan MB, Onyekwere O, Sidransky D, Vogelstein B, Craig RW: Participation of p53 protein in the cellular response to DNA damage. Cancer Res 1991;51:6304–6311
Kuerbitz SJ, Plunkett BS, Walsh W, Kastan MB: Wild-type p53 is a cell cycle checkpoint determinant following irradiation. Proc Natl Acad Sci USA 1992;89:7491–7495
Kastan MB, Zhan Q, El-Deiry WS, Carrier F, Jacks T, Walsh WV, Plunkett BS, Vogelstein B, Fornace AJ: A mammalian cell cycle checkpoint pathway utilizing p53 and GADD45 is defective in ataxia-telangiectasia. Cell 1992;71:587–597
Nelson WG, Kastan MB: DNA Strand breaks: The DNA template alterations that trigger p53-dependent DNA damage response pathways. Mol Cell Biol 1994;14:1815–1823
Zhan Q, Lord KA, Alamo I, Hollander MC, Carrier F, Ron D, Kohn KW, Hoffman B, Lieberman DA, Fornace AJ Jr: The gadd and Myd genes define a novel set of mammalian genes encoding acidic proteins that synergistically suppress cell growth. Mol Cell Biol 1994;14:2361–2371
El Deiry WS, Harper JW, O’Conner PM, Velculescu VE, Canman CE, Jackman J, Pietenpol JA, Burrell M, Hill DE, Wang Y, Wiman KG, Mercer WE, Kastan MB, Kohn KW, Elledge SJ, Kinzler KW, Vogelstein B: WAF1/CIP1 is induced in p53-mediated G1 arrest and apoptosis. Cancer Res 1994;54:1169–1174
Dulic V, Kaufmann WK, Wilson SJ, Harper W, Elledge SJ, Reed SI: p53-dependent inhibition of cyclin-dependent kinase activities in human fibroblasts during radiation-induced Gi arrest. Cell 1994;76:1013–1023
Slebos RJC, Lee MH, Plunkett BS, Kessis TD, Williams BO, Jacks T, Hedrick L, Kastan MB, Cho KR: p53-dependent G1 arrest involves pRB-related proteins and is disrupted by the human papillomavirus 16 E7 oncoprotein. Proc Natl Acad Sci USA 1994;91:5320–5324
Barak Y, Juven T, Haffner R, Oren M: mdm2 expression is induced by wild type p53 activity. EMBO J 1993;12:461–468
Wu X, Bayle H, Olson D, Levine AJ: The p53-mdm2 autoregulatory feedback loop. Genes Dev 1993;7:1126–1132
Chen C, Oliner JD, Zhan Q, Fornace AJ, Vogelstein B, Kastan MB: Interaction between p53 and MDM2 in mammalian cell cycle checkpoint pathway. Proc Natl Acad Sci USA 1994;91:2684–2688
Smith ML, Chen IT, Zhan Q, Bae I, Chen CY, Gilmer TM, Kastan MB, O’Connor PM, Fornace AJ Jr: Interaction of the p53-regulated protein Gadd45 with proliferating cell nuclear antigen. Science 1994;266:1376–1379
Khanna KK, Lavin MF: Ionizing radiation and UV induction of p53 protein by different pathways in ataxia-telangiectasia cells. Oncogene 1993;8:3307–3312
Canman CE, Wolff AC, Chen CY, Fornace AJ, Kastan MB: The p53-dependent G1 cell cycle checkpoint pathway and ataxia-telangiectasia. Cancer Res 1994;54:5054–5058
Lavin MF, Khanna KK, Beamish H, Teale B, Hobson K, Watters D: Defect in radiation signal transduction in ataxia-telangiectasia. Int J Radiat Biol 1994;66:S151–S156
Lowe SW, Schmitt SW, Smith SW, Osborne BA, Jacks T: p53 is required for radiation-induced apoptosis in mouse thymocytes. Nature 1993;362:847–849
Clarke AR, Purdie CA, Harrison DJ, Morris RG, Bird CC, Hooper ML, Wylie AH: Thymocyte apoptosis induced by p53-dependent and independent pathways. Nature 1993;362:849–852
Lotem J, Sachs L: Hematopoietic cells from mice deficient in wildtype p53 are more resistant to induction of apoptosis by some agents. Blood 1993;82:1092–1096
Meyn MS, Strasfeld L, Allen C: Testing the role of p53 in the expression of genetic instability and apoptosis in ataxia-telangiectasia. Int J Radiat Biol 1994;66:S141–S149
Wicking C, Williamson B: From linked marker to gene. Trends in Genet 1991;7:288–293
Collins FS: Positional cloning: Let’s not call it reverse anymore. Nature Genet 1992;1:3–6
Gatti RA, Berkel I, Boder E, Braedt G, Charmley P, Concannon P, Ersoy F, Foroud T, Jaspers NGJ, Lange K, Lathrop GM, Leppert M, Nakamura Y, O’Connel P, Paterson M, Salser W, Sanal, O, Silver J, Sparkes RS, Susi E, Weeks DE, Wei S, White R, Yoder F: Localization of an ataxia-telangiectasia gene to chromosome 11q22-23. Nature 1988;336:577–580
Bootsma D, Hoeijmakers JHJ: The molecular basis of nucleotide excision repair syndromes. Mutat Res 1994;307:15–23
Tanaka K, Satokata I, Ogita Z, Uchida T, Okada Y: Molecular cloning of a mouse DNA repair gene that complements the defect of group-A xeroderma pigmentosum. Proc Natl Acad Sci USA 1989;86:5512–5516
Legerski R, Peterson C: Expression cloning of a human DNA repair gene involved in xeroderma pigmentosum group C. Nature 1992;359:70–73
Strathdee CA, Gavish H, Shannon WR, Buchwald M: Cloning of cDNAs for Fanconi’s anaemia by functional complementation. Nature 1992;356:763–767
Lehmann AR, Arlett CF, Burke JF, Green MHL, James MR, Lowe JE: A derivative of an ataxia-telangiectasia (A-T) cell line with normal radiosensitivity but A-T-like inhibition of DNA synthesis. Int J Radiat Biol 1986;49:639–643
Green MHL, Lowe J, Arlett CF, Harcourt SA, Burke JF, James MR, Lehmann AR, Povey SM: A gamma-ray-resistant derivative of an ataxia telangiectasia cell line obtained following DNA-mediated gene transfer. J Cell Sci Suppl 1987;6:127–137
Lohrer H, Blum M, Herrlich P: Ataxia telangiectasia resists gene cloning: An account of parameters determining gene transfer into human recipient cells. Mol Gen Genet 1988;212:474–480
Kapp LN. Painter RB: Stable radioresistance in ataxia-telangiectasia cells containing DNA from normal human cells. Int J Radiat Biol 1989;56:667–675
Kapp LN, Painter RB, Yu LC, van Loon N, Richard CW III, James MR, Cox DR, Murnane JP: Cloning of a candidate gene for ataxia telangiectasia group D. Am J Hum Genet 1992;51:45–54
Hosoi Y, Kapp LN: Expression of a candidate ataxia-telangiectasia group D gene in cultured fibroblast cell lines and human tissues. Int J Radiat Biol 1994;66:S71–S76
Leonhardt EA, Kapp LN, Young BR, Murnane JP: Nucleotide sequence analysis of a candidate gene for ataxia-telangiectasia group D (ATDC). Genomics 1994;19:130–136
Murnane JP, Zhu Y, Young BR, Christman MF: Expression of the candidate AT gene ATDC is not detectable in a human cell line with a normal response to ionizing radiation. Int J Radiat Biol 1994;66:S77–S84
Ziv Y, Danieli T. Rotman G, Sartiel A, Bar-Shira A, Swirski R, Schimke RT, Eddy RL, Shows TB, Shiloh Y: Complementation of the cellular A-T phenotype by gene transfer; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Berlin, Springer, 1993, vol H77, pp 65–74.
Okayama H, Berg P: A cDNA vector that permits expression of cDNA inserts in mammalian cells. Mol Cell Biol 1985;3:280–289
Ziv Y, Etkin S, Danieli T, Amiel A, Ravia Y, Jaspers NGJ, Shiloh Y: Cellular and biochemical characteristics of an immortalized ataxiatelangiectasia (group AB) cell line. Cancer Res 1989;49:2495–2501
Ejima Y, Oshimura M, Sasaki MS: Establishment of a novel immortalized cell line from ataxia telangiectasia fibroblasts and its use for the chromosomal assignment of radiosensitivity gene. Int J Radiat Biol 1990;58:989–997
Komatsu K, Kodama S, Okumura Y, Koi M, Oshimura M: Restoration of radiation resistance in ataxia-telangiectasia cells by the introduction of normal human chromosome 11. Mutat Res 1990;235:59–63
Ejima Y, Oshimura M, Sasaki MS: Determination of the chromosomal site for the human radiosensitive ataxia telangiectasia gene by chromosome transfer. Mutat Res 1991;250:337–343
Kodama S, Komatsu K, Okumura Y, Oshimura M: Suppression of X-ray-induced chromosome aberrations in ataxia telangiectasia cells by introduction of a normal human chromosome 11. Mutat Res 1992;293:31–37
Ejima Y, Oshimura M, Sasaki MS: Use of microcell hybrids for analysis of the 11q23 region and improved localization of the A-T group A/C genes; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Berlin, Springer, 1993, vol H77, pp 75–85.
Lambert C, Schultz RA, Smith M, Wagner-McPherson C, McDaniel LD, Donlon T, Stanbridge EJ, Friedberg EC: Functional complementation of ataxia-telangiectasia group D (AT-D) cells by microcellmediated chromosome transfer and mapping of the AT-D locus to the region 11q23-23. Proc Natl Acad Sci USA 1991;88:5907–5911
Collins AR: Mutant rodent cell lines sensitive to ultraviolet light, ionizing radiation and cross-linking agents: A comprehensive survey of genetic and biochemical characteristics. Mutat Res 1993;293:99–118
Thacker J, Ganesh AN: DNA-break repair, radioresistance of DNA synthesis, and camptothecin sensitivity in the radiation-sensitive irs mutants: Comparisons to ataxia-telangiectasia cells. Mutat Res 1990;235:49–58
Cheong N, Wang Y, Jackson M, Iliakis G: Radiation-sensitive irs mutants rejoin DNA doublestrand breaks with efficiency similar to that of parental V79 cells but show altered response to radiationinduced g2 delay. Mutat Res 1992;274:111–122
Zdzienicka MZ, Jaspers NGJ, van der Schans GP, Natarajan AT, Simons JWIM: Ataxia-telangiectasia-like Chinese hamster V79 cell mutants with radioresistant DNA synthesis, chromosomal instability, and normal DNA strand break repair. Cancer Res 1989;49:1481–1485
Jongmans W, Verhaegh GWCT, Sankaranarayanan K, Lohman PHM, Zdzienicka MZ: Cellular characteristics of Chinese hamster cell mutants resembling ataxia telangiectasia cells. Mutat Res 1993;294:207–214
Zdzienicka MZ, Verhaegh GWCT, Jongmans W, Jaspers NGJ, Oshimura M, James MR, Lohman PHM: AT-like radiosensitive rodent cell mutants: An alternative approach to the isolation of the AT gene(s); in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Berlin, Springer, 1993, vol H77, pp 87–97.
Verhaegh GWCT, Jaspers NGJ, Lohman PHM, Zdzienicka MZ: Co-dominance of radioresistant DNA synthesis in a group of AT-like Chinese hamster cell mutants. Cytogenet Cell Genet 1993;63:176–180
Jongmans W, Wiegart J, Oshimura M, James MR, Lohman PHM, Zdzienicka MZ: Human chromosome 11 complements ataxia-telangiectasia cells but does not complement the defect m AT-like Chinese hamster cell mutants. Hum Genet 1993;92:259–264
Zdzienicka MZ, Verhaegh GWCT, Jongmans W, Morolli B, Jaspers NGJ, Oshimura M: Functional complementation studies with X-ray-sensitive mutants of Chinese hamster cells closely resembling ataxia-telangiectasia cells. Int J Radiat Biol 1994;66:S189–S195
Belt BGM, Groeneveld H, Teubel WJ, van de Putte P, Backendorf C: Construction and properties of an Epstein-Barr-virus-derived cDNA expression vector for human cells. Gene 1989;84:407–417
Groger RK, Morrow DM, Tykochinski ML: Directional antisense and sense cDNA cloning using Epstein-Barr virus episomal expression vectors. Gene 1989;81:285–294
Peterson C, Legersky R: High-frequency transformation of human repair-deficient celll lines by an Epstein-Barr virus-based expression vector. Gene 1991:107:279–284.
Swirski RA, van den Berg D, Murphy AJM, Lambert CM, Friedberg EC, Schimke RT: Improvements in the Epstein-Barr-based shuttle vector system for direct cloning in human tissue culture cells. Methods: Companion Methods Enzy-mol 1992;4:133–142
Meyn MS, Lu-Kuo JM, Herzing LBK: Expression cloning of multiple human cDNAs that complement the phenotypic defects of ataxia-telangiectasia group D fibroblasts. Am J Hum Genet 1993;53:1206–1216
Ziv Y, Bar-Shira A, Jorgensen TJ, Russell PS, Sartiel A, Shows TB, Eddy RL, Buchwald M, Legerski R, Schimke RT, Shiloh Y: Human cDNA clones that complement the radiomimetic sensitivity of ataxiatelangiectasia (group A) cells. Submitted.
Chen P, Girjes A, Hobson K, Beamish H, Khanna KK, Farrell A, Gatel M, Teale B, Lavin MF: Genetic complementation of radiosensitivity by 3′ untranslated regions (UTR) of RNA. Submitted.
Komatsu K, Okumura Y, Kodame S, Yoshida M, Miller RC: Lack of correlation between radiosensitivity and inhibition of DNA synthesis in hybrids (A-T × Heia). Int J Radiat Biol 1989;56:863–867
Harley HG, Brook JD, Rundle SA, Crow S, Reardon W, Buckler AJ, Harper PS, Housman DE, Shaw DJ: Expansion of an unstable DNA region and phenotypic variation in myotonic dystrophy. Nature 1992;355:545–546
The Huntington’s Disease Collaborative Research Group: A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 1993;72:971–983
The European Polycystic Kidney Disease Consortium: The polycystic kidney disease 1 gene encodes a 14 kb transcript and lies within a duplicated region on chromosome 16. Cell 1994;77:881–894
Miki Y. Swensen J, Shattuck-Eidens D, et al: A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 1994;266:66–71
Lefebvre S, Burglen L, Reboullet S, Clermont O, Burlet P, Viollet L, Benichou B, Cruaud C, Millasseau P, Zeviani M, Le Paslier D, Frezal J, Cohen D, Weissenbach J, Munnich A, Melki J: Identification and characterization of a spinal muscular atrophy-determining gene. Cell 1995;80:155–165
Roy N, Mahadevan MS, McLean M, Shutler G, Yaraghi Z, Farahni R, Baird S, Besner-Johnston A, Lefebvre C, Kang X, Salih M, Aubry H, Tamai K, Guan X, Ioannou P, Crawford TO, de Jong P, Surh L, Ikeda JE, Korneluk RG, MacKenzie A: The gene for neuronal apoptosis inhibitory protein is partially deleted in individuals with spinal muscular atrophy. Cell 1995;80:167–178
Strathdee Ca, Duncan AMV, Buchwald M: Evidence for at least four Fanconi anaemia genes including FACC on chromosome 9. Nature Genet 1992;1:196–198
Sanal O, Wei S, Foroud T, Malhotra U, Concannon P, Charmley P, Salser W, Lange K, Gatti RA: Further mapping of an ataxia-telangiectasia locus to the chromosome 11q23 region. Am J Hum Genet 1990;47:860–866
McConville CM, Woods CG, Farrail M, Metcalfe JA, Taylor AMR: Analysis of 7 polymorphic markers at chromosome 11q22–23 in 35 ataxia-telangiectasia families: Further evidence of linkage. Hum Genet 1990;85:215–220
McConville CM, Formstone CJ, Hernandez D, Thick J, Taylor AMR: Fine mapping of the chromosome 11q22–23 region using PFGE, linkage and haplotype analysis: Localization of the gene for ataxia telangiectasia to a 5-cM region flanked by NCAM/DRD2 and STMY/CJ52.75, ph2.22. Nucleic Acids Res 1990;18:4335–4343
Sanal O, Lange E, Telatar M, Sobel E, Salazar-Novak J, Ersoy F, Morrison A, Concannon P, Tolun A, Gatti RA: Ataxia-telangiectasia: Linkage analysis of chromosome 11q22–23 markers in Turkish families. FASEB J 1992,6:2846–2852.
Ziv Y, Frydman M, Lange E, Zelnik N, Rotman G, Julier C, Jaspers NGJ, Dagan Y, Abeliovicz D, Dar H, Borochowitz Z, Lathrop M, Gatti RA, Shiloh Y: Ataxia-telangiectasia: Linkage analysis in highly inbred Arab and Druze families and differentiation from an ataxiamicrocephaly-cataract syndrome. Hum Genet 1992:88:619–626.
Cornells F, James M, Cherif D, Tokino T, Davies J, Girault D, Bernard C, Crocquett MF, Theau D, Avet-Loiseau H, Litt M, Berger R, Nakamura Y, Lathrop M, Julier C: Precise localization of a gene responsible for ataxia-telangiectasia on chromosome 11q; in Gatti RA, Painter RB (eds): Ataxia-Telangiectasia. NATO ASI Series. Berlin, Springer, 1993, vol H77, pp 23–36.
Charmley P, Foroud T, Wei S, Concannon P, Weeks D, Lange K, Gatti RA: A primary linkage map of the human chromosome 11q22–23 region. Genomics 1990;6:316–323
Julier C, Nakamura Y, Lathrop M, O’Connell P, Leppert M, Litt M, Mohandas T, Lalouel JM, White R: A detailed genetic map of the long arm of chromosome 11. Genomics 1990;7:335–345
Wei S, Rocchi M, Archidiacono N, Sacchi N, Romeo G, Gatti RA: Physical mapping of the human chromosome 11q23 region containing the ataxia-telangiectasia locus. Cancer Genet Cytogenet 1990;46:1–8
Savage PD, Jones C, Silver J, Geurts van Kessel AHM, Gonzalez-Sarmiento R, Palm L, Hanson CA, Kersey JH: Mapping studies and expression of genes located on human chromosome 11, band q23. Cytogenet Cell Genet 1988;49:289–292
Charmley P, Nguyen J, Wei S, Gatti RA: Genetic linkage analysis and homology relationships of genes located on human chromosome 11q. Genomics 1991;10:608–617
Ziv Y, Rotman G, Frydman M, Foroud T, Gatti RA, Shiloh Y: The ATC (ataxia-telangiectasia complementation group C) locus localizes to 11q22-q23. Genomics 1991;9:373–375
Foroud T, Sobel E, Ziv Y, Goradia T, Wei S, Charmley P, McConville C, Chao A, Chessa L, Tolun A, Sanal O, Julier C, Concannon P, Fiorilli M, Taylor M, Shiloh Y, Lange K, Gatti RA: Localization of the AT locus to an 8 cM interval defined by STMY and S132. Am J Hum Genet 1991;49:1263–1279
McConville CM, Byrd PL, Ambrose HJ, Stankovic T, Ziv Y, Bar-Shira A, Vanagaite L, Rotman G, Shiloh Y, Gillett GT, Riley JH, Taylor AMR: Paired STSs amplified from radiation hybrids, and from associated YACs, identify highly polymorphic loci flanking the ataxia-telangiectasia locus on chromosome 11q22–23. Hum Mol Genet 1993;2:969–974
Ambrose HJ, Byrd PJ, McConville CM, Cooper PR, Stankovitz T, Riley JH, Shiloh Y, McNamara JO, Fuko T, Taylor AMR: A physical map across chromosome 11q22–23 containing the major locus for ataxia telangiectasia. Genomics 1994;21:612–619
Formstone CJ, Byrd PJ, Ambrose HJ, Riley JH, Hernandez D, McConville CM, Taylor AM: The order and orientation of a cluster of metalloproteinase genes, stromelysin 2, collagenase and stromelysin together with D11s35, on chromosome 11q22-23. Genomics 1993;16:289–291
Gillett GT, McConville CM, Byrd PJ, Stankovic T, Taylor AMR, Hunt DM, West LF, Fox MF, Povey S, Benham FJ: Irradiation hybrids for human chromosome 11: Characterization and use for generating region-specific markers in 11q14-q23. Genomics 1993:15: 332–341.
Richard CW, Cox DR, Kapp L, Murnane J. Cornellis F, Julier C, Lathrop M, James MR: A radiation hybrid map of human chromosome 11q22–23 containing the ataxia-telangiectasia disease locus. Genomics 1993;17:1–5
James MR, Richard CW III, Schott JJ, Yousry C, Clark K, Bell J, Terwilliger JD, Hazan J, Dubay C, Vigal A, Agrapart M, Imai T, Nakamura Y, Polymeropoulos M, Weissenbach J, Cox DR, Lathrop GM: A radiation hybrid map of 506 STS markers spanning human chromosome 11. Nat Genet 1994;8:70–76
Uhrhammer N, Concannon P, Huo Y, Nakamura Y, Gatti RA: A pulsed-field gel electrophoresis map in the ataxia-telangiectasia region of chromosome 11q22.3. Genomics 1994;20:278–280
McConville CM, Byrd PJ, Ambrose HJ, Taylor AMR: Genetic and physical mapping of the ataxia-telangiectasia locus on chromosome 11q22–23. Int J Radiat Biol 1994;66:S45–S56
Vanagaite L, Savitsky K, Rotman G, Ziv Y, Gerken SC, White R, Weissenbach J, Gillett G, Benham FJ, Richard CW, James MR, Collins FS, Shiloh Y: Physical localization of microsatellite markers at the ataxia-telangiectasia locus at 11q22-23. Genomics 1994;22:231–233
Vanagaite L, James MR, Rotman G, Savitsky K, Bar-Shira A, Gilad S, Ziv Y, Uchenik V, Sartiel A, Collins FS, Sheffield VC, Weissenbach J, Shiloh Y: A high-density microsatellite map of the ataxiatelangiectasia locus. Hum Genet, in press.
Rotman G, Vanagaite L, Collins FS, Shiloh Y: Rapid identification of polymorphic CA-repeats in YAC clones spanning the ataxiatelangiectasia locus on chromosome 11q22–23. Mol Biotechnol, in press.
Rotman G, Vanagaite L, Collins Fs, Shiloh Y: Three dinucleotide repeat polymorphisms at the ataxia-telangiectasia locus. Hum Mol Genet 1994;3:2079.
Gatti RA, Lange E, Rotman G, Chen X, Uhrhammer N, Liang T, Chiplunkar S, Yang L, Udar N, Dandekar S, Sheikhavandi S, Wang Z, Yang HM, Polikow J, Elashoff M, Teletar M, Sanal O, Chessa L, McConville C, Taylor M, Shiloh Y, Porras O, Borreson AL, Wegner RD, Curry C, Gerken S, Lange K, Concannon P: Genetic haplotyping of ataxia-telangiectasia families localizes the major gene to an 850 kb region on chromosome 11q23.1. Int J Radiat Biol 1994;66:S57–S62
Lange E, Borreson AL, Chen X, Chessa L, Chiplunkar S, Concannon P, Dandekar S, Gerken S, Lange K, Liang T, McConville C, Polakow J, Porras O, Rotman G, Sanal O, Telatar M, Sheikhavandi S, Shiloh Y, Sobel E, Taylor M, Udar N, Uhrhammer N, Vanagaite L, Wang Z, Yang HM, Yang L, Ziv Y, Gatti RA: Localization of an ataxia-telangiectasia gene to a ∼ 500 kb interval on chromosome 11q23.1: linkage analysis of 176 families by an international consortium. Submitted.
Lange K, Sobel E: A random walk method for computing genetic location scores. Am J Hum Genet 1991;49:1320–1334
Sobel E, Lange K: Metropolis sampling in pedigree analysis. Stat Methods Med Res 1993;2:263–282
Hastbacka J, de la Chapelle A, Mahtani MM, Clines G, Reeve-Daly MP, Daly M, Hamilton BA, Kusumi K, Tnvedi B, Weaver A, Coloma A, Lovett M, Buckler A, Kaitila I, Lander ES: The diastrophic dysplasia gene encodes a novel sulfate transporter: Positional cloning by fine-structure linkage disequilibrium mapping. Cell 1994;78:1073–1087
Mitchison HM, Taschner PFM, O’Rawe AM, de Vos N, Phillips HA, Thompson AD, Kozman HM, Haines JL, Schlumpf K, D’Arigo K, Boustany RM, Callen DF, Breuning MH, Gardiner RM, Mole SE, Lerner TJ: Genetic mapping of the Batten disease locus (CLN3) to the interval D16S288-D16S383 by analysis of haplotypes and allelic association. Genomics 1994;22:465–468
Sulisalo T, Klockars J, Makitie O, Francomano CA, de la Chapelle A, Kaitila I, Sistonen P: High-resolution linkage disequilibrium mapping of the cartilage-hair-hypoplasia gene. Am J Hum Genet 1994;55:937–945
Oskato R, Bar-Shtra A, Vanagaite L, Ziv Y, Ehrlich S, Rotman G, McConville CM, Chakravarti A, Shiloh Y: Ataxia-telangiectasia: Allelic association with 11q22–23 markers in Moroccan-Jewish patients. Am J Hum Genet 1993; 53(suppl):A1055.
Uhrhammer NA, Lange E, Gatti RA: Localization of an ataxia-telangiectasia gene by linkage disequilibrium in the Costa Rican population. Am J Hum Genet 1994;55(suppl):A354.
Rotman G, Savitsky K, Ziv Y, Cole CG, Higgins MJ, Bar-Am I, Dunham I, Bar-Shira A, Vanagaite L, Shinzen Q, Zhang J, Nowak NJ, Chandrasekharappa SC, Lehrach H, Avivi L, Shows TB, Collins FS, Bentley DR, Shiloh Y: A YAC contig spanning the ataxia-telangiectasia locus (groups A and C) at 11q22–23. Genomics 1994;24:234–242
Rotman G, Savitsky K, Vanagaite L, Bar-Shira A, Ziv Y, Gilad S, Uchenik V, Smith S, Shiloh Y: Physical and genetic mapping at the ATA/ATC locus on chromosome 11q22–23. Int J Radiat Biol 1994;66:S63–66
Lovett M: Fishing for complements: Finding genes by direct selection. Trends Genet 1994;10:352–357
Parimoo S, Kolluri R, Weissman SM: cDNA selection from total yeast DNA containing YACs. Nucleic Acids Res 1993:18:4422–4423.
Church DM, Stotler CJ, Rutter JL, Murrell JR, Trofatter JA, Buckler AJ: Isolation of genes from complex sources of mammalian genomic DNA using exon amplification. Nature Genet 1994,6:98–104.
Shiloh Y, Ziv Y, Savitsky K, Bar-Shira A, Gilad S, Vanagaite L, Uchenik V, Smith S, Patanjali SR, Tagle D, Simmons A, Clines G, Collins FS, Weissman S, Lovett M, Rotman G: Genetic, physical and functional analysis of the ataxiatelangiectasia locus on chromosome 11q22–23. Am J Hum Genet 1994;55(suppl):A49.
Acknowledgements
This review is dedicated to Dr. Elena Boder, who pioneered A-T research and has inspired it for four decades.
The author wishes to acknowledge the valuable contribution of the A-T research team at Tel Aviv University and is indebted to numerous colleagues for comments and exchanges of information. Work in the author’s laboratory is supported by the A-T Children’s Project, the A-T Medical Research Foundation, the Thomas Appeal (A-T Medical Research Trust), the United States-Israel Binational Science Foundation, and the National Institute of Neurological Disorders and Stroke (NS31763).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Shiloh, Y. Ataxia-Telangiectasia: Closer to Unraveling the Mystery. Eur J Hum Genet 3, 116–138 (1995). https://doi.org/10.1159/000472285
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1159/000472285
Key Words
This article is cited by
-
The role of nuclear architecture in genomic instability and ageing
Nature Reviews Molecular Cell Biology (2007)
-
ATM Polymorphism and hereditary nonpolyposis colorectal cancer (HNPCC) ageof onset (United States)
Cancer Causes & Control (2005)
-
Molecular variants of the ATM gene in Hodgkin's disease in children
British Journal of Cancer (2004)
-
Radiation-induced apoptosis in developing mouse retina exhibits dose-dependent requirement for ATM phosphorylation of p53
Cell Death & Differentiation (2004)
-
ATM function and telomere stability
Oncogene (2002)