Featured
-
-
Article |
 Dependency-aware deep generative models for multitasking analysis of spatial omics data
Dependency-aware deep generative models for multitasking analysis of spatial omics data
Using a dependency-aware deep generative framework, spaVAE efficiently models spatially resolved transcriptomics data and advances diverse analysis tasks. Following similar strategies, spaPeakVAE and spaMultiVAE enable spatial ATAC-seq data and spatial multi-omics data modeling and analysis, respectively.
- Tian Tian
- , Jie Zhang
- & Hakon Hakonarson
-
Brief Communication |
 Scalable telomere-to-telomere assembly for diploid and polyploid genomes with double graph
Scalable telomere-to-telomere assembly for diploid and polyploid genomes with double graph
By effective and efficient integration of PacBio HiFi, Oxford Nanopore Technologies ultra-long and other sequencing data types, hifiasm (UL) enables telomere-to-telomere diploid and polyploid genome assembly at a population scale.
- Haoyu Cheng
- , Mobin Asri
- & Heng Li
-
Research Briefing |
 Quantitative profiling of regulatory DNA activity at single-cell resolution
Quantitative profiling of regulatory DNA activity at single-cell resolution
We developed a two-pronged strategy to functionally probe the enormous repertoire of noncoding DNA within genomes. Our approach markedly improved signal-to-noise ratio and successfully intersected single-cell genomics with reporter assays. The result delivers a multiplex and highly quantitative readout of regulatory sequences’ activity in dynamic and multicellular systems.
-
Article
| Open Access Scalable and unbiased sequence-informed embedding of single-cell ATAC-seq data with CellSpace
Scalable and unbiased sequence-informed embedding of single-cell ATAC-seq data with CellSpace
By learning to embed DNA k-mers and cells into a joint space, CellSpace improves single-cell ATAC-seq analysis in multiple tasks such as latent structure discovery, transcription factor activity inference and batch effect mitigation.
- Zakieh Tayyebi
- , Allison R. Pine
- & Christina S. Leslie
-
Article
| Open Access Multiplex profiling of developmental cis-regulatory elements with quantitative single-cell expression reporters
Multiplex profiling of developmental cis-regulatory elements with quantitative single-cell expression reporters
Single-cell quantitative expression reporters enable high-sensitivity quantitative characterization of cis-regulatory elements at the single-cell level in multicellular systems.
- Jean-Benoît Lalanne
- , Samuel G. Regalado
- & Jay Shendure
-
Technology Feature |
 scRNA-seq: oh, the joys
scRNA-seq: oh, the joys
To those who seek transcriptomic information at high resolution, scale and throughput, single-cell RNA sequencing brings the data. Scientists share tips and future plans as they reflect on the method’s rise to stardom.
- Vivien Marx
-
Brief Communication
| Open Access Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis
Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis
This study evaluates the performance of GPT-4 in single-cell type annotation.
- Wenpin Hou
- & Zhicheng Ji
-
Article |
 Constructing telomere-to-telomere diploid genome by polishing haploid nanopore-based assembly
Constructing telomere-to-telomere diploid genome by polishing haploid nanopore-based assembly
This work introduces two polishers for refining the draft genome generated from nanopore long reads, as well as an assembler pipeline for producing telomere-to-telomere diploid genome with low error rate.
- Joshua Casey Darian
- , Ritu Kundu
- & Wing-Kin Sung
-
Article |
 scGPT: toward building a foundation model for single-cell multi-omics using generative AI
scGPT: toward building a foundation model for single-cell multi-omics using generative AI
Pretrained using over 33 million single-cell RNA-sequencing profiles, scGPT is a foundation model facilitating a broad spectrum of downstream single-cell analysis tasks by transfer learning.
- Haotian Cui
- , Chloe Wang
- & Bo Wang
-
Article
| Open Access Genome-scale pan-cancer interrogation of lncRNA dependencies using CasRx
Genome-scale pan-cancer interrogation of lncRNA dependencies using CasRx
A CasRx-based screening platform for genome-scale long noncoding RNA transcriptome perturbation is described.
- Juan J. Montero
- , Riccardo Trozzo
- & Roland Rad
-
Perspective |
 Challenges and perspectives in computational deconvolution of genomics data
Challenges and perspectives in computational deconvolution of genomics data
This Perspective provides an overview of major challenges and associated recommendations in computational deconvolution for genomics data.
- Lana X. Garmire
- , Yijun Li
- & Andrew E. Teschendorff
-
Article
| Open Access Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN
Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN
SATURN performs cross-species integration and analysis using both single-cell gene expression and protein representations generated by protein language models.
- Yanay Rosen
- , Maria Brbić
- & Jure Leskovec
-
Brief Communication
| Open Access Correcting PCR amplification errors in unique molecular identifiers to generate accurate numbers of sequencing molecules
Correcting PCR amplification errors in unique molecular identifiers to generate accurate numbers of sequencing molecules
This study introduces a method utilizing homotrimeric nucleotide blocks to achieve accurate counts of RNA molecules in both bulk and single-cell sequencing data.
- Jianfeng Sun
- , Martin Philpott
- & Adam P. Cribbs
-
Article |
 mEnrich-seq: methylation-guided enrichment sequencing of bacterial taxa of interest from microbiome
mEnrich-seq: methylation-guided enrichment sequencing of bacterial taxa of interest from microbiome
mEnrich-seq leverages DNA methylation differences between bacteria to enrich taxa of interest from metagenomic samples for selective sequencing.
- Lei Cao
- , Yimeng Kong
- & Gang Fang
-
Method to Watch |
 Large models for genomics
Large models for genomics
Large language models are learning the language of genomics.
- Lin Tang
-
Method to Watch |
 Spatially resolved multiomics
Spatially resolved multiomics
Spatially resolved multimodal omics offers a collective way to capture molecular information in complex tissues.
- Lei Tang
-
Brief Communication
| Open Access Uniform quantification of single-nucleus ATAC-seq data with Paired-Insertion Counting (PIC) and a model-based insertion rate estimator
Uniform quantification of single-nucleus ATAC-seq data with Paired-Insertion Counting (PIC) and a model-based insertion rate estimator
This study demonstrates the need and advantage of uniformly quantifying single-nucleus ATAC-seq data using Paired-Insertion Counting.
- Zhen Miao
- & Junhyong Kim
-
Article
| Open Access Combinatorial single-cell profiling of major chromatin types with MAbID
Combinatorial single-cell profiling of major chromatin types with MAbID
MAbID offers a multiplexing approach to uncover the genomic distributions of various epigenetic markers, enabling the study of how these markers jointly direct gene expression.
- Silke J. A. Lochs
- , Robin H. van der Weide
- & Jop Kind
-
Article |
 Improved sequence mapping using a complete reference genome and lift-over
Improved sequence mapping using a complete reference genome and lift-over
By combining fast lift-over and selective re-mapping, levioSAM2 enables efficient and accurate read mapping and variant calling leveraging complete reference genomes.
- Nae-Chyun Chen
- , Luis F. Paulin
- & Ben Langmead
-
Analysis
| Open Access Comprehensive benchmarking and guidelines of mosaic variant calling strategies
Comprehensive benchmarking and guidelines of mosaic variant calling strategies
A benchmarking study evaluates 11 mosaic single-nucleotide variant and insertion-deletion mutation detection approaches.
- Yoo-Jin Ha
- , Seungseok Kang
- & Sangwoo Kim
-
Article
| Open Access Population-level integration of single-cell datasets enables multi-scale analysis across samples
Population-level integration of single-cell datasets enables multi-scale analysis across samples
By learning representations for both cells and various condition covariates, scPoli facilitates atlas-level integration and analysis of single-cell genomics datasets with improved interpretability.
- Carlo De Donno
- , Soroor Hediyeh-Zadeh
- & Fabian J. Theis
-
-
Analysis |
 Benchmarking long-read RNA-sequencing analysis tools using in silico mixtures
Benchmarking long-read RNA-sequencing analysis tools using in silico mixtures
This analysis leverages experimentally sequenced data and in silico mixtures to simulate transcript expression differences, which enables a performance assessment of long-read tools developed for isoform detection, differential transcript expression analysis and differential transcript usage analysis.
- Xueyi Dong
- , Mei R. M. Du
- & Matthew E. Ritchie
-
Article
| Open Access A new Bayesian factor analysis method improves detection of genes and biological processes affected by perturbations in single-cell CRISPR screening
A new Bayesian factor analysis method improves detection of genes and biological processes affected by perturbations in single-cell CRISPR screening
Guided sparse factor analysis (GSFA) is a powerful statistical framework to detect changes in gene expression as a result of perturbations in single-cell CRISPR screening.
- Yifan Zhou
- , Kaixuan Luo
- & Xin He
-
Brief Communication
| Open Access Fast and robust metagenomic sequence comparison through sparse chaining with skani
Fast and robust metagenomic sequence comparison through sparse chaining with skani
skani achieves fast calculation of average nucleotide identity (ANI) between metagenome-assembled genomes (MAGs), with improved robustness against incomplete and fragmented MAGs.
- Jim Shaw
- & Yun William Yu
-
Article
| Open Access Deep generative modeling of transcriptional dynamics for RNA velocity analysis in single cells
Deep generative modeling of transcriptional dynamics for RNA velocity analysis in single cells
veloVI enhances RNA velocity analysis with uncertainty quantification and extensibility by deep generative modeling of gene-specific transcriptional dynamics.
- Adam Gayoso
- , Philipp Weiler
- & Nir Yosef
-
Review Article |
 Principles and challenges of modeling temporal and spatial omics data
Principles and challenges of modeling temporal and spatial omics data
This Review discusses statistical and computational strategies for analyzing various spatial and temporal omics data types, with an emphasis on the common modeling principles.
- Britta Velten
- & Oliver Stegle
-
Article |
 Scalable Nanopore sequencing of human genomes provides a comprehensive view of haplotype-resolved variation and methylation
Scalable Nanopore sequencing of human genomes provides a comprehensive view of haplotype-resolved variation and methylation
This work introduces a wet lab and computational pipeline, Napu, for small variant calling and de novo assembly of Nanopore sequencing data, which leads to comparable performances to short-read sequencing and allows for large-scale long-read sequencing projects.
- Mikhail Kolmogorov
- , Kimberley J. Billingsley
- & Benedict Paten
-
Article |
 Recovery of missing single-cell RNA-sequencing data with optimized transcriptomic references
Recovery of missing single-cell RNA-sequencing data with optimized transcriptomic references
This paper presents an improved approach for mapping single-cell RNA-seq reads with optimized transcriptomic references, which markedly recovers previously missing gene expression data.
- Allan-Hermann Pool
- , Helen Poldsam
- & Yuki Oka
-
Research Briefing |
 scNanoHi-C uncovers single-cell high-order chromatin structures and gene regulation
scNanoHi-C uncovers single-cell high-order chromatin structures and gene regulation
Leveraging nanopore long-read sequencing, scNanoHi-C identifies multiway interactions between enhancers and their target promoters within a single cell. Compared with short-read-based single-cell Hi-C or population-based multiway sequencing methods, scNanoHi-C offers new opportunities to investigate the heterogeneities of single-cell gene regulation networks mediated by high-order 3D chromatin structures.
-
Article |
 scNanoHi-C: a single-cell long-read concatemer sequencing method to reveal high-order chromatin structures within individual cells
scNanoHi-C: a single-cell long-read concatemer sequencing method to reveal high-order chromatin structures within individual cells
scNanoHi-C combines Nanopore long-read sequencing with a proximity-ligation-based Hi-C protocol to profile high-order genome structures in individual cells, enabling the capture of multiway interactions among enhancers and promoters.
- Wen Li
- , Jiansen Lu
- & Fuchou Tang
-
Article
| Open Access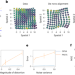 Alignment of spatial genomics data using deep Gaussian processes
Alignment of spatial genomics data using deep Gaussian processes
Gaussian Process Spatial Alignment (GPSA) aligns multiple spatially resolved genomics and histology datasets and improves downstream analysis.
- Andrew Jones
- , F. William Townes
- & Barbara E. Engelhardt
-
Article |
 UniAligner: a parameter-free framework for fast sequence alignment
UniAligner: a parameter-free framework for fast sequence alignment
Compared to other sequences, extra-long tandem repeats, such as centromeres and immunoglobulin loci, are more difficult to align. This study presents UniAligner, a computational method for efficiently and accurately aligning extra-long tandem repeats, facilitating analysis of their variation and evolution.
- Andrey V. Bzikadze
- & Pavel A. Pevzner
-
News & Views |
 Dissecting gene regulation with multimodal sequencing
Dissecting gene regulation with multimodal sequencing
Recently proposed computational approaches explore casual links between chromatin and transcriptional changes that are provided by single-cell multimodal sequencing to bridge the knowledge gap in transcriptional regulatory control.
- Ivan G. Costa
-
Article |
 Dictys: dynamic gene regulatory network dissects developmental continuum with single-cell multiomics
Dictys: dynamic gene regulatory network dissects developmental continuum with single-cell multiomics
By probabilistic modeling of gene regulation and expression kinetics, Dictys infers dynamic and context-specific gene regulatory networks using single-cell multiomics data.
- Lingfei Wang
- , Nikolaos Trasanidis
- & Luca Pinello
-
Article
| Open Access SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks
SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks
SCENIC+ is a comprehensive toolbox for inferring and analyzing enhancer-driven gene regulatory networks using single-cell multiomic data.
- Carmen Bravo González-Blas
- , Seppe De Winter
- & Stein Aerts
-
Article |
 Significance analysis for clustering with single-cell RNA-sequencing data
Significance analysis for clustering with single-cell RNA-sequencing data
This study presents a significance analysis framework for evaluating single-cell clusters. Application of the method detects cases of over-clustering in reported single-cell RNA-sequencing analysis results.
- Isabella N. Grabski
- , Kelly Street
- & Rafael A. Irizarry
-
This Month |
Grant-writing rituals
Everyone has their own methods to address the time-consuming and challenging task of grant-writing.
- Vivien Marx
-
Brief Communication |
 A comparison of single-coverage and multi-coverage metagenomic binning reveals extensive hidden contamination
A comparison of single-coverage and multi-coverage metagenomic binning reveals extensive hidden contamination
This study shows, when analyzing multi-sample metagenomic datasets, the multi-coverage binning approach outperforms the single-coverage binning alternative in generating bins with higher quality and less contamination.
- Jennifer Mattock
- & Mick Watson
-
Review Article |
 A survey of algorithms for the detection of genomic structural variants from long-read sequencing data
A survey of algorithms for the detection of genomic structural variants from long-read sequencing data
This Review provides an overview of computational methods recently developed for detecting and analyzing structural variants using long-read sequencing data.
- Mian Umair Ahsan
- , Qian Liu
- & Kai Wang
-
Article
| Open Access MultiVI: deep generative model for the integration of multimodal data
MultiVI: deep generative model for the integration of multimodal data
By learning a joint representation using deep generative modeling, MultiVI integrates multimodal and single-modality single-cell datasets, which enhances multiple functionalities.
- Tal Ashuach
- , Mariano I. Gabitto
- & Nir Yosef
-
Article
| Open Access Multiscale analysis of pangenomes enables improved representation of genomic diversity for repetitive and clinically relevant genes
Multiscale analysis of pangenomes enables improved representation of genomic diversity for repetitive and clinically relevant genes
The PanGenome Research Tool Kit (PGR-TK) achieves flexible and scalable representation, visualization and analysis of genomic variation using pangenome graphs.
- Chen-Shan Chin
- , Sairam Behera
- & Justin M. Zook
-
Article |
 Context-aware transcript quantification from long-read RNA-seq data with Bambu
Context-aware transcript quantification from long-read RNA-seq data with Bambu
Leveraging long-read RNA-seq data and machine learning, Bambu facilitates accurate transcript discovery and quantification.
- Ying Chen
- , Andre Sim
- & Jonathan Göke
-
-
-
Research Briefing |
 Capturing prime editor off-target sites within the genome
Capturing prime editor off-target sites within the genome
Prime editing systems hold tremendous promise for the precise correction of pathogenic mutations. We developed a method to tag sequences modified by a prime editor to evaluate its genome-wide precision for therapeutic applications.
-
Research Briefing |
 Photoselective sequencing harnesses microscopy to guide genomic analyses
Photoselective sequencing harnesses microscopy to guide genomic analyses
Photoselective sequencing is a new method for genomic and epigenomic profiling within specific regions of a biological specimen that are chosen using light microscopy. This combination of spatial and sequencing information preserves the connections between genomic and environmental properties and deepens our understanding of structure–function relationships in cells and tissues.
-
Article |
 Photoselective sequencing: microscopically guided genomic measurements with subcellular resolution
Photoselective sequencing: microscopically guided genomic measurements with subcellular resolution
Photoselective sequencing combines targeted illumination and photocaged fragment libraries to enable the spatial analysis of genomic sequence and chromatin accessibility profiles with subcellular resolution in the context of complex tissues.
- Sarah M. Mangiameli
- , Haiqi Chen
- & Fei Chen
-
Analysis
| Open Access Comparison of transformations for single-cell RNA-seq data
Comparison of transformations for single-cell RNA-seq data
This paper compares different transformation approaches for analysis of single-cell RNA-sequencing data and provides recommendations for method selection.
- Constantin Ahlmann-Eltze
- & Wolfgang Huber

 Large-scale foundation model on single-cell transcriptomics
Large-scale foundation model on single-cell transcriptomics Resolving epigenetic base ambiguity
Resolving epigenetic base ambiguity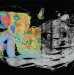 A new member of the spatial omics family
A new member of the spatial omics family