Featured
-
-
Article
| Open Access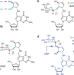 Synthesis and structure elucidation of the human tRNA nucleoside mannosyl-queuosine
Synthesis and structure elucidation of the human tRNA nucleoside mannosyl-queuosine
Mannosyl-queuosine (manQ) is a non-canonical RNA nucleoside present in the anticodon loop of certain tRNAs. Here, the authors use a combination of total synthesis and mass spectrometry to contradict the literature-reported structure and show that manQ features an alpha-allyl connectivity of its mannose moiety.
- Markus Hillmeier
- , Mirko Wagner
- & Thomas Carell
-
Article
| Open Access Atlas of RNA editing events affecting protein expression in aged and Alzheimer’s disease human brain tissue
Atlas of RNA editing events affecting protein expression in aged and Alzheimer’s disease human brain tissue
Resources reporting RNA editing sites from brain tissue have been published. Here, the authors provide an atlas of RNA editing events found in the aged and Alzheimer’s disease human brain tissue resulting in changes at protein level.
- Yiyi Ma
- , Eric B. Dammer
- & Philip L. De Jager
-
Article
| Open Access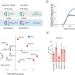 NUDT2 initiates viral RNA degradation by removal of 5′-phosphates
NUDT2 initiates viral RNA degradation by removal of 5′-phosphates
RNA of some viruses is protected from degradation by a 5′ triphosphate group. Here the authors identify nudix hydrolase 2 (NUDT2) as novel antiviral defense protein that dephosphorylates viral RNA and thereby enables its degradation.
- Beatrice T. Laudenbach
- , Karsten Krey
- & Andreas Pichlmair
-
Article
| Open Access MDA5 disease variant M854K prevents ATP-dependent structural discrimination of viral and cellular RNA
MDA5 disease variant M854K prevents ATP-dependent structural discrimination of viral and cellular RNA
MDA5 is the primary immune sensor for SARS-CoV-2 and many other viruses. Mutations in MDA5 can cause disease. Here the authors employ CryoEM and biochemical methods to show how steric constraints cause MDA5 to misrecognize endogenous RNA as viral RNA.
- Qin Yu
- , Alba Herrero del Valle
- & Yorgo Modis
-
Article
| Open Access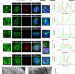 DHX15-independent roles for TFIP11 in U6 snRNA modification, U4/U6.U5 tri-snRNP assembly and pre-mRNA splicing fidelity
DHX15-independent roles for TFIP11 in U6 snRNA modification, U4/U6.U5 tri-snRNP assembly and pre-mRNA splicing fidelity
In yeast, TFIP11 and DHX15 promote the disassembly of spliceosome complex after splicing is completed. Here the authors show that human TFIP11 functions independently of DHX15 and is required for U6 snRNA 2’-O-methylation and U4/U6.U5 tri-snRNP assembly.
- Amandine Duchemin
- , Tina O’Grady
- & Denis Mottet
-
Article
| Open Access Bi-directional ribosome scanning controls the stringency of start codon selection
Bi-directional ribosome scanning controls the stringency of start codon selection
Start codon selection is commonly thought to occur through the unidirectional scanning of the mRNA by the 40 S ribosome. Here the authors provide evidence that the pre-initiation complex can backslide on the mRNA to initiate translation at upstream AUG codons.
- Yifei Gu
- , Yuanhui Mao
- & Shu-Bing Qian
-
Article
| Open Access Pseudoknot length modulates the folding, conformational dynamics, and robustness of Xrn1 resistance of flaviviral xrRNAs
Pseudoknot length modulates the folding, conformational dynamics, and robustness of Xrn1 resistance of flaviviral xrRNAs
Exoribonuclease-resistant RNAs (xrRNAs) are RNA elements that block the exoribonucleolytic degradation of RNA. Here the authors show how a long-range pseudoknot length modulates the Mg2+-dependence of flaviviral xrRNA’s folding, conformational dynamics and Xrn1 resistance.
- Xiaolin Niu
- , Ruirui Sun
- & Xianyang Fang
-
Article
| Open Access Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
The molecular events underlying the assembly and maturation of the early pre-60S particles during eukaryotic ribosome synthesis are not well understood. Here, the authors combine yeast genetics and biochemical experiments to characterise the functions of two important players of eukaryotic ribosome biogenesis, the box C/D snoRNP snR190 and the helicase Dbp7, which both interact. They show that the snR190 snoRNA acts as a RNA chaperone that assists the structuring of the 25S rRNA during the maturation of early pre-60S particles and that Dbp7 is important for facilitating remodeling events in the peptidyl transferase center region of the 25S rRNAs during the maturation of early pre-60S particles.
- Mariam Jaafar
- , Julia Contreras
- & Anthony K. Henras
-
Article
| Open Access Photoactivatable ribonucleosides mark base-specific RNA-binding sites
Photoactivatable ribonucleosides mark base-specific RNA-binding sites
RNA-protein interactions play critical roles in post-transcriptional gene regulation. Here the authors demonstrate pRBS-ID, an updated MS/MS-based method that combines the benefits of photoactivatable ribonucleosides and the chemical cleavage of RNA.
- Jong Woo Bae
- , Sangtae Kim
- & Jong-Seo Kim
-
Article
| Open Access Coupled protein synthesis and ribosome-guided piRNA processing on mRNAs
Coupled protein synthesis and ribosome-guided piRNA processing on mRNAs
Ribosome can mediate piRNA biogenesis from long non-coding RNAs following translation of the short open reading frames. Here the authors show that 80S ribosome also guides piRNA production from 3’ UTR of protein-coding genes after translation of long open reading frames, indicating a general piRNA biogenesis mechanism regardless of their precursor ORF length.
- Yu H. Sun
- , Ruoqiao Huiyi Wang
- & Xin Zhiguo Li
-
Article
| Open Access RN7SK small nuclear RNA controls bidirectional transcription of highly expressed gene pairs in skin
RN7SK small nuclear RNA controls bidirectional transcription of highly expressed gene pairs in skin
The noncoding RNA RN7SK regulates RNA polymerase II pausing and splicing. Here the authors deplete RN7SK in mouse and human during epidermal stem cell differentiation and reveal a novel role in orchestrating bidirectional transcription of highly expressed gene pairs.
- Roberto Bandiera
- , Rebecca E. Wagner
- & Michaela Frye
-
Article
| Open Access A chemical probe based on the PreQ1 metabolite enables transcriptome-wide mapping of binding sites
A chemical probe based on the PreQ1 metabolite enables transcriptome-wide mapping of binding sites
The small modified nucleotide PreQ1 binds to the PreQ1 riboswitch and regulates gene expression by inducing RNA conformation change. Here the authors design and characterize a specific preQ1-derived probe by x-ray crystallography, mass spectral analysis and transcriptome-wide using Chem-CLIP.
- Sumirtha Balaratnam
- , Curran Rhodes
- & John S. Schneekloth Jr
-
Article
| Open Access Assembly defects of human tRNA splicing endonuclease contribute to impaired pre-tRNA processing in pontocerebellar hypoplasia
Assembly defects of human tRNA splicing endonuclease contribute to impaired pre-tRNA processing in pontocerebellar hypoplasia
Mutations within subunits of the tRNA splicing endonuclease complex (TSEN) are associated with pontocerebellar hypoplasia (PCH). Here the authors show that tRNA intron excision is catalyzed by tetrameric TSEN assembled from inactive heterodimers, and provide evidence that modulation of TSEN stability may contribute to PCH phenotypes.
- Samoil Sekulovski
- , Pascal Devant
- & Simon Trowitzsch
-
Article
| Open Access In vivo structure and dynamics of the SARS-CoV-2 RNA genome
In vivo structure and dynamics of the SARS-CoV-2 RNA genome
RNA secondary structure is important for viral replication, transcription and translation. Here the authors employ SPLASH method and map in vivo RNA interactions and dynamics of SARS-CoV-2 RNA genome in different viral life cycles.
- Yan Zhang
- , Kun Huang
- & Zhihu Zhao
-
Article
| Open Access Virus detection via programmable Type III-A CRISPR-Cas systems
Virus detection via programmable Type III-A CRISPR-Cas systems
CRISPR-Cas based virus detection assays can be both rapid and sensitive. Here the authors use a Type III-A CRISPR-Cas system to detect SARS-CoV-2.
- Sagar Sridhara
- , Hemant N. Goswami
- & Hong Li
-
Article
| Open Access High-throughput RNA sequencing of paraformaldehyde-fixed single cells
High-throughput RNA sequencing of paraformaldehyde-fixed single cells
Current high-throughput single-cell transcriptomic methods are incompatible with paraformaldehyde, a common cell fixation technique. Here the authors present FD-seq, a method for droplet-based RNA sequencing of paraformaldehyde-fixed, stained and sorted single cells.
- Hoang Van Phan
- , Michiel van Gent
- & Savaş Tay
-
Article
| Open Access Molecular basis for PICS-mediated piRNA biogenesis and cell division
Molecular basis for PICS-mediated piRNA biogenesis and cell division
C. elegans piRNA biogenesis and chromosome segregation (PICS) complex is composed of TOFU-6, PICS-1, ERH-2, and two mutually exclusive factors PID-1 and TOST-1. By employing biochemical, structural, and cellular biology methods, the authors show that the PICS complex is an octamer consisting of two copies of each subunit, and functions in piRNA biogenesis and mitosis.
- Xiaoyang Wang
- , Chenming Zeng
- & Chao Xu
-
Article
| Open Access Hypoxia regulates overall mRNA homeostasis by inducing Met1-linked linear ubiquitination of AGO2 in cancer cells
Hypoxia regulates overall mRNA homeostasis by inducing Met1-linked linear ubiquitination of AGO2 in cancer cells
Met1-linked linear ubiquitination (M1-Ubi) is catalyzed by linear ubiquitin chain assembly complex (LUBAC). Here the authors show that Ago2 protein is M1-Ubi modified by LUBAC complex under hypoxia condition leading to less association of miRNA target mRNAs to Ago2 protein and de-repression of miRNA targets.
- Hailong Zhang
- , Xian Zhao
- & Jianxiu Yu
-
Article
| Open Access A quantitative model predicts how m6A reshapes the kinetic landscape of nucleic acid hybridization and conformational transitions
A quantitative model predicts how m6A reshapes the kinetic landscape of nucleic acid hybridization and conformational transitions
m6A RNA post-transcriptional modification changes RNA hybridization kinetics. Here the authors show that the methylamino group can adopt syn-conformation pairing with uridine with a mismatch-like conformation in RNA duplex. They also develop a quantitative model that predicts how m6A affects the kinetics of hybridization.
- Bei Liu
- , Honglue Shi
- & Hashim M. Al-Hashimi
-
Article
| Open Access Humans and other commonly used model organisms are resistant to cycloheximide-mediated biases in ribosome profiling experiments
Humans and other commonly used model organisms are resistant to cycloheximide-mediated biases in ribosome profiling experiments
Ribosome profiling has become the gold standard to analyze mRNA translation dynamics, and the translation inhibitor cycloheximide (CHX) is often used in its application. Here the authors systematically demonstrate that CHX does not bias the outcome of ribosome profiling experiments in most organisms.
- Puneet Sharma
- , Jie Wu
- & Sebastian A. Leidel
-
Article
| Open Access Spatiotemporally-resolved mapping of RNA binding proteins via functional proximity labeling reveals a mitochondrial mRNA anchor promoting stress recovery
Spatiotemporally-resolved mapping of RNA binding proteins via functional proximity labeling reveals a mitochondrial mRNA anchor promoting stress recovery
Proximity labeling is used to map and discover proteins in specific subcellular compartments. Here the authors combine APEX-mediated proximity labeling with organic-aqueous phase separation to identify nuclear, nucleolar, and outer mitochondrial membrane RNA binding proteins.
- Wei Qin
- , Samuel A. Myers
- & Alice Y. Ting
-
Article
| Open Access Global view on the metabolism of RNA poly(A) tails in yeast Saccharomyces cerevisiae
Global view on the metabolism of RNA poly(A) tails in yeast Saccharomyces cerevisiae
RNA polyadenosine tails are important for the export, translation and stability of mRNAs and play a role in non-coding RNA biogenesis. Here the authors measure yeast poly(A) tail lengths by direct RNA sequencing, revealing its dynamics in yeast exonuclease, deadenylase and poly(A) polymerase mutants.
- Agnieszka Tudek
- , Paweł S. Krawczyk
- & Andrzej Dziembowski
-
Article
| Open Access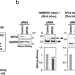 SPF45/RBM17-dependent, but not U2AF-dependent, splicing in a distinct subset of human short introns
SPF45/RBM17-dependent, but not U2AF-dependent, splicing in a distinct subset of human short introns
The length distribution of human pre-mRNA introns is very extensive. The authors demonstrate that splicing in a subset of short introns is dependent on SPF45 (RBM17), which replaces authentic U2AF-heterodimer on the truncated poly-pyrimidine tracts and interacts with the U2 snRNP protein SF3b155.
- Kazuhiro Fukumura
- , Rei Yoshimoto
- & Akila Mayeda
-
Article
| Open Access Structures of tmRNA and SmpB as they transit through the ribosome
Structures of tmRNA and SmpB as they transit through the ribosome
Trans-translation, mediated by small protein B (SmpB) and transfer-messenger RNA (tmRNA), enables recycling of the ribosomes stalled on defective mRNAs in bacteria. Here, the authors report structures of the ribosome during trans-translation that reveal a translocation intermediate and elucidate the movements of the tmRNA-SmpB complex in the ribosome.
- Charlotte Guyomar
- , Gaetano D’Urso
- & Reynald Gillet
-
Article
| Open Access m6Am-seq reveals the dynamic m6Am methylation in the human transcriptome
m6Am-seq reveals the dynamic m6Am methylation in the human transcriptome
m6Am is a dynamic and reversible RNA modification found on the mRNA cap. Here the authors report m6Am-seq to directly distinguish m6Am from m6A and identify functional methylation sites.
- Hanxiao Sun
- , Kai Li
- & Chengqi Yi
-
Article
| Open Access Structural dynamics of single SARS-CoV-2 pseudoknot molecules reveal topologically distinct conformers
Structural dynamics of single SARS-CoV-2 pseudoknot molecules reveal topologically distinct conformers
The RNA pseudoknot of SARS-CoV-2 promotes -1 programmed ribosomal frameshifting. Here the authors use single molecule force spectroscopy to study the folding of this pseudoknot, showing that it forms at least two different pseudoknot conformers with distinct fold topologies.
- Krishna Neupane
- , Meng Zhao
- & Michael T. Woodside
-
Article
| Open Access Switching at the ribosome: riboswitches need rProteins as modulators to regulate translation
Switching at the ribosome: riboswitches need rProteins as modulators to regulate translation
Translational regulation by riboswitches is an important mechanism for the modulation of gene expression in bacteria. Here the authors show that the ligand-induced allosteric switch in the adenine-sensing riboswitch from V. vulnificus is insufficient and leads only to a partial opening of the ribosome binding site and requires interaction with 30S-bound ribosomal protein S1, which acts as an RNA chaperone.
- Vanessa de Jesus
- , Nusrat S. Qureshi
- & Boris Fürtig
-
Article
| Open Access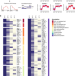 Puf6 primes 60S pre-ribosome nuclear export at low temperature
Puf6 primes 60S pre-ribosome nuclear export at low temperature
Ribosome biogenesis is crucially dependent on proper rRNA folding, a process assisted by chaperones. Here the authors reveal how Puf6 promotes correct rRNA folding at low temperature, a condition where mis-paired RNA folding intermediates frequently accumulate.
- Stefan Gerhardy
- , Michaela Oborská-Oplová
- & Vikram Govind Panse
-
Article
| Open Access A distinct assembly pathway of the human 39S late pre-mitoribosome
A distinct assembly pathway of the human 39S late pre-mitoribosome
Assembly of the mitoribosome requires assistance from numerous specialized factors. Here, structures of the human 39S late assembly intermediates identify several assembly factors which keep the 16S rRNA in immature conformations, and reveal deacylated tRNA in the ribosomal E-site, suggesting a role in 39S assembly.
- Jingdong Cheng
- , Otto Berninghausen
- & Roland Beckmann
-
Article
| Open Access A small molecule that induces translational readthrough of CFTR nonsense mutations by eRF1 depletion
A small molecule that induces translational readthrough of CFTR nonsense mutations by eRF1 depletion
Premature termination codons can cause early translation termination and lead to disease. Here the authors perform a screen to identify compounds with readthrough activity and show that these reduce eRF1 levels to suppress premature termination associated with cystic fibrosis.
- Jyoti Sharma
- , Ming Du
- & David M. Bedwell
-
Article
| Open Access Single-molecule analysis of processive double-stranded RNA cleavage by Drosophila Dicer-2
Single-molecule analysis of processive double-stranded RNA cleavage by Drosophila Dicer-2
Fly Dicer-2 is thought to use two distinct – processive or distributive – modes of cleavage by distinguishing the terminal structures of double-stranded RNA (dsRNA) substrates with the help of its cofactor LoquaciousPD (Loqs-PD). Here the authors show by single-molecule imaging that dsRNA terminal structures and Loqs-PD change the probability for Dicer to initiate processive cleavage but not the mode of cleavage action per se.
- Masahiro Naganuma
- , Hisashi Tadakuma
- & Yukihide Tomari
-
Article
| Open Access Attention-based multi-label neural networks for integrated prediction and interpretation of twelve widely occurring RNA modifications
Attention-based multi-label neural networks for integrated prediction and interpretation of twelve widely occurring RNA modifications
RNA modifications appear to play a role in determining RNA structure and function. Here, the authors develop a deep learning model that predicts the location of 12 RNA modifications using primary sequence, and show that several modifications are associated, which suggests dependencies between them.
- Zitao Song
- , Daiyun Huang
- & Jia Meng
-
Article
| Open Access A natural riboswitch scaffold with self-methylation activity
A natural riboswitch scaffold with self-methylation activity
In humans, protein methyltransferase is responsible for RNA methylation using S-adenosylmethionine as a methyl group donor. Here the authors report a self-methylation activity of a bacterial riboswitch.
- Laurin Flemmich
- , Sarah Heel
- & Ronald Micura
-
Article
| Open Access The pan-cancer lncRNA PLANE regulates an alternative splicing program to promote cancer pathogenesis
The pan-cancer lncRNA PLANE regulates an alternative splicing program to promote cancer pathogenesis
Genomic amplification of chromosome 3q often encodes proteins that contribute to cancer development. Here the authors identify a non-coding product of the 3q region, the lncRNA PLANE that promotes tumorigenesis through the deregulation of transcriptional corepressor NCOR2 pre-mRNA splicing.
- Liu Teng
- , Yu Chen Feng
- & Xu Dong Zhang
-
Article
| Open Access Structural basis for late maturation steps of the human mitoribosomal large subunit
Structural basis for late maturation steps of the human mitoribosomal large subunit
Mitochondrial ribosomes (mitoribosomes) are characterized by a distinct architecture and thus biogenesis pathway. Here, cryo-EM structures of mitoribosome large subunit assembly intermediates elucidate final steps of 16 S rRNA folding, methylation and peptidyl transferase centre (PTC) completion, as well as functions of several mitoribosome assembly factors.
- Miriam Cipullo
- , Genís Valentín Gesé
- & Joanna Rorbach
-
Article
| Open Access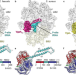 Structural basis of ABCF-mediated resistance to pleuromutilin, lincosamide, and streptogramin A antibiotics in Gram-positive pathogens
Structural basis of ABCF-mediated resistance to pleuromutilin, lincosamide, and streptogramin A antibiotics in Gram-positive pathogens
Mitochondrial ribosomes (mitoribosomes) are characterized by a distinct architecture and thus biogenesis pathway. Here, cryo-EM structures of mitoribosome large subunit assembly intermediates elucidate final steps of 16 S rRNA folding, methylation and peptidyl transferase centre (PTC) completion, as well as functions of several mitoribosome assembly factors.
- Caillan Crowe-McAuliffe
- , Victoriia Murina
- & Daniel N. Wilson
-
Article
| Open Access Large Stokes shift fluorescence activation in an RNA aptamer by intermolecular proton transfer to guanine
Large Stokes shift fluorescence activation in an RNA aptamer by intermolecular proton transfer to guanine
Fluorogenic RNA aptamers such as Chili display strong fluorescence enhancement upon aptamer–ligand complex formation. Here, the authors provide insights into the mechanism of fluorescence activation of Chili by solving the crystal structures of Chili with its bound positively charged ligands DMHBO+ and DMHBI+, and they reveal that Chili uses an excited state proton transfer mechanism based on time-resolved optical spectroscopy measurements.
- Mateusz Mieczkowski
- , Christian Steinmetzger
- & Claudia Höbartner
-
Article
| Open Access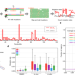 RNA structure probing reveals the structural basis of Dicer binding and cleavage
RNA structure probing reveals the structural basis of Dicer binding and cleavage
Sequencing methods such as icSHAPE were developed to probe RNA structures transcriptome-wide in cells. To probe intact RNA structures, the authors develop icSHAPE-MaP and apply to Dicer-bound substrates showing that distance measuring is important for Dicer cleavage of pre-miRNAs.
- Qing-Jun Luo
- , Jinsong Zhang
- & Qiangfeng Cliff Zhang
-
Article
| Open Access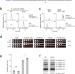 A single m6A modification in U6 snRNA diversifies exon sequence at the 5’ splice site
A single m6A modification in U6 snRNA diversifies exon sequence at the 5’ splice site
Spliceosomal U6 snRNA is modified by m6A. Here the authors show that m6A modification on U6 snRNA promotes efficient recognition of the splice site through cooperating with U5 snRNA, indicating that U6 snRNA m6A allows the 5’ exon sequence variation.
- Yuma Ishigami
- , Takayuki Ohira
- & Tsutomu Suzuki
-
Article
| Open Access Activation of Prp28 ATPase by phosphorylated Npl3 at a critical step of spliceosome remodeling
Activation of Prp28 ATPase by phosphorylated Npl3 at a critical step of spliceosome remodeling
Yeast helicase Prp28 promotes the first step of spliceosome remodeling. By placing a photoactivatable unnatural amino acid in Prp28, the authors capture Prp28 in action revealing its dynamic interactions and cofactor Npl3.
- Fu-Lung Yeh
- , Shang-Lin Chang
- & Tien-Hsien Chang
-
Article
| Open Access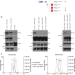 RTEL1 influences the abundance and localization of TERRA RNA
RTEL1 influences the abundance and localization of TERRA RNA
Long non coding RNA TERRA transcripts can form R-loops at chromosome ends. Here, the authors reveal a role for the helicase RTEL in affecting TERRA levels and localization.
- Fiorella Ghisays
- , Aitor Garzia
- & John H. J. Petrini
-
Article
| Open Access 40S ribosome profiling reveals distinct roles for Tma20/Tma22 (MCT-1/DENR) and Tma64 (eIF2D) in 40S subunit recycling
40S ribosome profiling reveals distinct roles for Tma20/Tma22 (MCT-1/DENR) and Tma64 (eIF2D) in 40S subunit recycling
Following the termination of translation at stop codons, the eukaryotic 60S subunit of the ribosome is removed by the ATPase ABCE1. Here using 40S ribosome footprinting the authors provide a direct demonstration that the yeast orthologs of eIF2D, MCT-1, and DENR recycle the 40S subunits.
- David J. Young
- , Sezen Meydan
- & Nicholas R. Guydosh
-
Article
| Open Access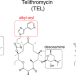 Context-specific action of macrolide antibiotics on the eukaryotic ribosome
Context-specific action of macrolide antibiotics on the eukaryotic ribosome
Macrolide antibiotics inhibit bacterial translation in a context-specific manner, arresting ribosomes at defined sites within mRNAs and selectively inhibiting synthesis of only a subset of cellular proteins. Here the authors provide a structural basis for the context-specific activity of macrolides on the eukaryotic ribosome.
- Maxim S. Svetlov
- , Timm O. Koller
- & Alexander S. Mankin
-
Article
| Open Access Structural coordination between active sites of a CRISPR reverse transcriptase-integrase complex
Structural coordination between active sites of a CRISPR reverse transcriptase-integrase complex
In some systems, a single protein comprising reverse transcriptase (RT), integrase and maturase enables concerted sequence integration and crRNA production. Here, analyses including the structure of a Cas6-RT-Cas1—Cas2 complex suggest coordination between all three active sites and capacity to acquire CRISPR sequences from RNA and DNA substrates.
- Joy Y. Wang
- , Christopher M. Hoel
- & Jennifer A. Doudna
-
Article
| Open Access Structures of flavivirus RNA promoters suggest two binding modes with NS5 polymerase
Structures of flavivirus RNA promoters suggest two binding modes with NS5 polymerase
Flaviviruses use a ~70 nucleotide stem-loop structure called stem-loop A (SLA) at the 5’ end of the RNA genome as a promoter for RNA synthesis by the viral polymerase NS5. Here the authors describe the structures of dengue and Zika virus SLAs, identify the SLA-binding site on NS5, and propose models for how NS5 recognizes the RNA promoter.
- Eunhye Lee
- , Paul J. Bujalowski
- & Kyung H. Choi
-
Article
| Open Access High-resolution view of HIV-1 reverse transcriptase initiation complexes and inhibition by NNRTI drugs
High-resolution view of HIV-1 reverse transcriptase initiation complexes and inhibition by NNRTI drugs
Initiation of HIV-1 reverse transcription occurs at the host tRNALys3, which forms a complex with the 5’ end of the HIV-1 viral RNA and reverse transcriptase (RT). Here, the authors present the 2.8 Å cryo-EM structure of a minimal HIV-1 RT–vRNA–tRNALys3 initiation complex (miniRTIC), and miniRTIC structures with the bound non-nucleoside reverse transcriptase inhibitors nevirapine and efavirenz at 3.1 and 2.9 Å resolution, respectively.
- Betty Ha
- , Kevin P. Larsen
- & Elisabetta Viani Puglisi
-
Article
| Open Access Optimized photochemistry enables efficient analysis of dynamic RNA structuromes and interactomes in genetic and infectious diseases
Optimized photochemistry enables efficient analysis of dynamic RNA structuromes and interactomes in genetic and infectious diseases
RNA crosslinking and proximity ligation methods are used to identify transcriptome-wide base pairing interactions. Here, the authors report PARIS2 (psoralen analysis of RNA interactions and structures 2), a method for RNA duplex determination in vivo with higher efficiency than the previous PARIS method.
- Minjie Zhang
- , Kongpan Li
- & Zhipeng Lu
-
Article
| Open Access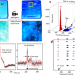 Synchronous RNA conformational changes trigger ordered phase transitions in crystals
Synchronous RNA conformational changes trigger ordered phase transitions in crystals
Time-resolved crystallography (TRX) is used for monitoring only small conformational changes of biomacromolecules within the same lattice. Here, the authors report the interplay between synchronous molecular rearrangements and lattice phase transitions in RNA crystals, providing the basis for the investigation of large conformational changes using TRX.
- Saminathan Ramakrishnan
- , Jason R. Stagno
- & Yun-Xing Wang
-
Article
| Open Access FTO-mediated cytoplasmic m6Am demethylation adjusts stem-like properties in colorectal cancer cell
FTO-mediated cytoplasmic m6Am demethylation adjusts stem-like properties in colorectal cancer cell
The demethylase FTO was shown to remove on N6-methyladenosine (m6A) and N6, 2’-O-dimethyladenosine (m6Am) modifications on RNAs. Here the authors show that FTO impedes cancer stem cell-like abilities in colorectal cancer cells through its m6Am demethylase activity, not through internal m6A demethylase activity.
- Sébastien Relier
- , Julie Ripoll
- & Alexandre David

 Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch
Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch