Featured
-
-
Article
| Open Access CLK-dependent exon recognition and conjoined gene formation revealed with a novel small molecule inhibitor
CLK-dependent exon recognition and conjoined gene formation revealed with a novel small molecule inhibitor
The phosphorylation of serine/arginine-rich proteins by CDC-like kinase is a central regulatory mechanism for RNA splicing reactions. Here, the authors synthesize a novel small molecule CLK inhibitor and map CLK-responsive alternative splicing events and discover an effect on conjoined gene transcription.
- Tyler Funnell
- , Shinya Tasaki
- & Samuel Aparicio
-
Article
| Open Access AKAP95 regulates splicing through scaffolding RNAs and RNA processing factors
AKAP95 regulates splicing through scaffolding RNAs and RNA processing factors
The chromatin-associated protein AKAP95 is known for its chromatin-related functions including enhancing transcription. Here the authors show that AKAP95 interacts with the splicing regulatory factors as well as RNAs to regulate the inclusion of exons and pre-mRNA splicing.
- Jing Hu
- , Alireza Khodadadi-Jamayran
- & Hao Jiang
-
Article
| Open Access Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq)
Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq)
Long non-coding RNAs are increasingly recognised to be important factors in regulating cellular processes and comprise a large faction of the transcriptome, however most are uncharacterised. Here the authors present RACE-Seq, a tool to improve and extend the annotation of low-expression transcripts.
- Julien Lagarde
- , Barbara Uszczynska-Ratajczak
- & Jennifer Harrow
-
Article
| Open Access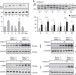 Therapeutic activity of modified U1 core spliceosomal particles
Therapeutic activity of modified U1 core spliceosomal particles
Modification of the spliceosome is being tested as a potential therapy for exon-skipping diseases, such as spinal muscular atrophy (SMA). Here the authors show that 70K and stem loop IV structural elements of a modified U1 particle are essential for splicing enhancement and effective treatment of SMA mice.
- Malgorzata Ewa Rogalska
- , Mojca Tajnik
- & Franco Pagani
-
Article
| Open Access Quaking promotes monocyte differentiation into pro-atherogenic macrophages by controlling pre-mRNA splicing and gene expression
Quaking promotes monocyte differentiation into pro-atherogenic macrophages by controlling pre-mRNA splicing and gene expression
Post-transcriptional control of RNA is important in health and disease. Here, the authors show that the RNA-binding protein Quaking guides pre-mRNA splicing and transcript abundance during monocyte to macrophage differentiation, and that Quaking depletion impairs pro-atherogenic foam cell formation.
- Ruben G. de Bruin
- , Lily Shiue
- & Eric P. van der Veer
-
Article
| Open Access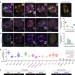 Cajal bodies are linked to genome conformation
Cajal bodies are linked to genome conformation
Nuclear bodies can nucleate at sites of active transcription and are beneficial for efficient gene expression. Here, the authors show that Cajal bodies, a prominent type of nuclear body, contribute to genome organization with global effects on gene expression and RNA splicing fidelity.
- Qiuyan Wang
- , Iain A. Sawyer
- & Miroslav Dundr
-
Article
| Open Access Structural basis of RNA recognition and dimerization by the STAR proteins T-STAR and Sam68
Structural basis of RNA recognition and dimerization by the STAR proteins T-STAR and Sam68
Sam68 and T-STAR are members of the STAR family of proteins, which regulate various aspects of RNA metabolism. Here, the authors reveal structural features required for alternative splicing regulation by these proteins.
- Mikael Feracci
- , Jaelle N. Foot
- & Cyril Dominguez
-
Article
| Open Access ESRP2 controls an adult splicing programme in hepatocytes to support postnatal liver maturation
ESRP2 controls an adult splicing programme in hepatocytes to support postnatal liver maturation
Alternative RNA splicing is important during organismal development. Here, the authors perform RNA-Seq on mouse and human liver samples to provide a comprehensive view of splicing events during liver development and growth, and identify Espr2 as a main regulator of these splicing processes.
- Amruta Bhate
- , Darren J. Parker
- & Auinash Kalsotra
-
Article
| Open Access Compound heterozygous mutations in the noncoding RNU4ATAC cause Roifman Syndrome by disrupting minor intron splicing
Compound heterozygous mutations in the noncoding RNU4ATAC cause Roifman Syndrome by disrupting minor intron splicing
Roifman Syndrome is a rare disorder whose disease manifestations include growth retardation, spondyloepiphyseal dysplasia and immunodeficiency. Here, the authors use whole-genome sequencing to discover that rare compound heterozygous variants disrupting the small nuclear RNA gene RNU4ATACcause Roifman Syndrome.
- Daniele Merico
- , Maian Roifman
- & Stephen W. Scherer
-
Article
| Open Access Abnormal splicing switch of DMD’s penultimate exon compromises muscle fibre maintenance in myotonic dystrophy
Abnormal splicing switch of DMD’s penultimate exon compromises muscle fibre maintenance in myotonic dystrophy
The splicing of the penultimate exon of the dystrophin gene is developmentally regulated. Here the authors show that the dysregulation of this exon’s splicing leads to the expression of an embryonic dystrophin form with a C-terminus distinct from the adult isoform, which leads to muscle wasting in zebrafish and mice.
- Frédérique Rau
- , Jeanne Lainé
- & Denis Furling
-
Article
| Open Access Widespread disruption of host transcription termination in HSV-1 infection
Widespread disruption of host transcription termination in HSV-1 infection
Herpes simplex virus 1 (HSV-1) efficiently shuts down host gene expression in infected cells. Here Rutkowski et al. analyse the genome-wide changes in transcription and translation in infected cells, and show that HSV-1 triggers an extensive disruption of transcription termination of cellular genes.
- Andrzej J. Rutkowski
- , Florian Erhard
- & Lars Dölken
-
Article |
 Inhibition of vemurafenib-resistant melanoma by interference with pre-mRNA splicing
Inhibition of vemurafenib-resistant melanoma by interference with pre-mRNA splicing
BRAF inhibitors have shown encouraging clinical effects in melanoma patients; however, patients rapidly develop resistance via different mechanisms including alternative splicing. Here the authors find a specific mutation affecting BRAF splicing and highlight the therapeutic potential of splicing interference.
- Maayan Salton
- , Wojciech K. Kasprzak
- & Tom Misteli
-
Article |
 PRMT9 is a Type II methyltransferase that methylates the splicing factor SAP145
PRMT9 is a Type II methyltransferase that methylates the splicing factor SAP145
Protein arginine methylation is an abundant post-translational modification often associated with RNA-binding proteins. Here the authors show that the previously uncharacterized PRMT9 enzyme catalyses the symmetrical methylation of SAP145, which promotes its association with the SMN complex and regulates splicing.
- Yanzhong Yang
- , Andrea Hadjikyriacou
- & Mark T. Bedford
-
Article |
 ALS-causative mutations in FUS/TLS confer gain and loss of function by altered association with SMN and U1-snRNP
ALS-causative mutations in FUS/TLS confer gain and loss of function by altered association with SMN and U1-snRNP
Dominant mutations in the RNA-binding protein FUS/TLS cause amyotrophic lateral sclerosis (ALS), an adult-onset motor neuron degenerative disease. Here, the authors show that ALS-causative FUS/TLS mutations directly bind the SMN and U1-snRNP complexes, producing both loss and gain of function effects on RNA processing.
- Shuying Sun
- , Shuo-Chien Ling
- & Don W. Cleveland
-
Article |
 Prevalent and distinct spliceosomal 3′-end processing mechanisms for fungal telomerase RNA
Prevalent and distinct spliceosomal 3′-end processing mechanisms for fungal telomerase RNA
In fission yeast, the telomerase RNA (TER) is produced through inhibition of the second step in splicing, resulting in spliceosomal cleavage. Here, the authors show that the inhibition of splicing is a conserved principle in fungal TER maturation that uses distinct molecular mechanisms across species.
- Xiaodong Qi
- , Dustin P. Rand
- & Julian J. -L. Chen
-
Article
| Open Access Diverse mechanisms for spliceosome-mediated 3′ end processing of telomerase RNA
Diverse mechanisms for spliceosome-mediated 3′ end processing of telomerase RNA
In fission yeast, the telomerase RNA (TER) is produced through spliceosomal cleavage. Here, Kannan et al. find that spliceosome-generated 3′ ends also occurs in other fungal TERs using distinct molecular mechanisms, suggesting multiple origins for this type of TER maturation pathway.
- Ram Kannan
- , Rachel M. Helston
- & Peter Baumann
-
Article |
 Aberrant splicing of U12-type introns is the hallmark of ZRSR2 mutant myelodysplastic syndrome
Aberrant splicing of U12-type introns is the hallmark of ZRSR2 mutant myelodysplastic syndrome
Somatic mutations in components of the core RNA splicing machinery, including ZRSR2, have been implicated in myelodysplastic syndrome (MDS). Here, Madan et al.show that ZRSR2 plays a pivotal role in splicing of the U12-type introns, while the U2-dependent splicing is largely unaffected in ZRSR2 mutant MDS bone marrow.
- Vikas Madan
- , Deepika Kanojia
- & H. Phillip Koeffler
-
Article |
 BCLAF1 and its splicing regulator SRSF10 regulate the tumorigenic potential of colon cancer cells
BCLAF1 and its splicing regulator SRSF10 regulate the tumorigenic potential of colon cancer cells
Alternative splicing often alters the biological function of proteins. Here, Zhou et al. show that the splicing factor SRSF10 directs the inclusion of exon5a in Bcl-2-associated transcription factor 1, and that this drives cell growth and tumorigenic potential in human colon cancer cells.
- Xuexia Zhou
- , Xuebing Li
- & Ying Feng
-
Article
| Open Access Phosphoregulation of Ire1 RNase splicing activity
Phosphoregulation of Ire1 RNase splicing activity
Ire1 is an effector of the unfolded protein response that is activated upon stress to maintain protein homeostasis. Here, Prischi et al. demonstrate that phosphorylation of Ire1 within its kinase activation loop increases its RNAse activity, thus identifying a regulatory cross-talk between the two domains.
- Filippo Prischi
- , Piotr R. Nowak
- & Maruf M. U. Ali
-
Article
| Open Access Widespread sex differences in gene expression and splicing in the adult human brain
Widespread sex differences in gene expression and splicing in the adult human brain
Men and women differ in terms of their neurochemistry, behaviour and susceptibility to disease. Here the authors show that sex differences in gene expression and splicing are widespread in adult human brain, and that sex-biased expression is likely to have functional consequences.
- Daniah Trabzuni
- , Adaikalavan Ramasamy
- & Mina Ryten
-
Article
| Open Access The thermodynamic patterns of eukaryotic genes suggest a mechanism for intron–exon recognition
The thermodynamic patterns of eukaryotic genes suggest a mechanism for intron–exon recognition
The thermodynamics of unwinding polynucleotide duplexes can be determined from energy changes for DNA and mRNA interactions. Here the authors show that the ratio between mRNA/DNA and DNA/DNA duplex stability upstream of the 3′- spice sites is a characteristic that can contribute to intron–exon recognition.
- Marina N. Nedelcheva-Veleva
- , Mihail Sarov
- & Stoyno S. Stoynov
-
Article |
 Splicing factor SRSF3 is crucial for hepatocyte differentiation and metabolic function
Splicing factor SRSF3 is crucial for hepatocyte differentiation and metabolic function
Splicing factors, such as the protein SRSF3, regulate mRNA metabolism but are hard to study in vivobecause genetic kockouts are usually lethal. Here, Sen and colleagues create mice with a hepatocyte-specific knockout of Srsf3 and demonstrate its role in hepatocyte differentiation and liver function.
- Supriya Sen
- , Hassan Jumaa
- & Nicholas J. G. Webster
-
Article |
 Post-transcriptional spliceosomes are retained in nuclear speckles until splicing completion
Post-transcriptional spliceosomes are retained in nuclear speckles until splicing completion
It is unclear where in the nucleus splicing takes place and how much occurs post-transcriptionally. Using antibodies raised against a phosphorylated splicing factor, Girardet al. show that the majority of splicing occurs co-transcriptionally and that post-transcriptional splicing occurs in nuclear speckles.
- Cyrille Girard
- , Cindy L. Will
- & Reinhard Lührmann
-
Article
| Open Access A segmental genomic duplication generates a functional intron
A segmental genomic duplication generates a functional intron
The appearance of a new intron that splits an exon without disrupting the corresponding peptide sequence is a rare event in vertebrate genomes. Hellstenet al.demonstrate that, under certain circumstances, a functional intron can be produced in a single step by segmental genomic duplication.
- Uffe Hellsten
- , Julie L. Aspden
- & Daniel S. Rokhsar
-
Article
| Open Access Chemical treatment enhances skipping of a mutated exon in the dystrophin gene
Chemical treatment enhances skipping of a mutated exon in the dystrophin gene
Duchenne muscular dystrophy is caused by a loss of thedystrophin gene, and control of dystrophin mRNA splicing could aid treatment of the disease. Nishida et al. show that a small molecule promotes skipping of exon 31 and increases production of a functional dystrophin protein in a patient.
- Atsushi Nishida
- , Naoyuki Kataoka
- & Masafumi Matsuo
-
Article |
 A role for TREX components in the release of spliced mRNA from nuclear speckle domains
A role for TREX components in the release of spliced mRNA from nuclear speckle domains
The pre-mRNA splicing and TREX mRNA export machineries are found in nuclear speckle domains. Diaset al. microinject CMV-DNA constructs into cells and find that transcripts containing functional splice sites accumulate in nuclear speckles and that the TREX complex is required to release the mRNA once processed.
- Anusha P. Dias
- , Kobina Dufu
- & Robin Reed

 Genome-wide identification of splicing QTLs in the human brain and their enrichment among schizophrenia-associated loci
Genome-wide identification of splicing QTLs in the human brain and their enrichment among schizophrenia-associated loci