Article
|
Open Access
Featured
-
-
Article
| Open Access Highly pathogenic avian influenza A (H5N1) in marine mammals and seabirds in Peru
Highly pathogenic avian influenza A (H5N1) in marine mammals and seabirds in Peru
Highly pathogenic avian influenza (HPAI) A/H5N1 has recently emerged in the Americas and has been implicated in mass die-off events of pelicans and sea lions. Here, the authors report sampling and characterisation of HPAI A/H5N1 genomes from five marine mammal and seabird species in Peru.
- Mariana Leguia
- , Alejandra Garcia-Glaessner
- & Jesus Lescano
-
Article
| Open Access Genomes of cultivated and wild Capsicum species provide insights into pepper domestication and population differentiation
Genomes of cultivated and wild Capsicum species provide insights into pepper domestication and population differentiation
Existing genetics and genomics studies of peppers mainly focus on single species. Here, the authors report a pepper graph pan-genome and a genome variation map of 500 accessions from five domesticated species and close wild relatives to reveal their domestication, introgression and population differentiation.
- Feng Liu
- , Jiantao Zhao
- & Xuexiao Zou
-
Article
| Open Access Multi-trait discovery and fine-mapping of lipid loci in 125,000 individuals of African ancestry
Multi-trait discovery and fine-mapping of lipid loci in 125,000 individuals of African ancestry
There have been few genome-wide association studies that analyze multiple lipid traits simultaneously, especially in individuals of African ancestry. To address this, the authors performed a multi-trait analysis of GWAS along with fine-mapping to find new genetic loci associated with lipid traits in individuals of African ancestry.
- Abram Bunya Kamiza
- , Sounkou M. Touré
- & Segun Fatumo
-
Article
| Open Access Screening non-conventional yeasts for acid tolerance and engineering Pichia occidentalis for production of muconic acid
Screening non-conventional yeasts for acid tolerance and engineering Pichia occidentalis for production of muconic acid
Baker’s yeast is a workhorse of industrial biotechnology, but it is not suited to overproduce many bulk bioproducts, especially organic acids. Here, the authors identify Pichia occidentalis as an acid tolerant yeast and engineer it for the production of muconic acid using a newly developed genome editing toolkit.
- Michael E. Pyne
- , James A. Bagley
- & Vincent J. J. Martin
-
Article
| Open Access Barcoded multiple displacement amplification for high coverage sequencing in spatial genomics
Barcoded multiple displacement amplification for high coverage sequencing in spatial genomics
Spatial genomics offers insights into cellular interactions within tissues. Here, the authors develop barcoded multiple displacement amplification, achieving high-coverage sequencing to map complex genomic variations within cellular landscapes.
- Jinhyun Kim
- , Sungsik Kim
- & Sunghoon Kwon
-
Article
| Open Access Comparative genomic analyses reveal the genetic basis of the yellow-seed trait in Brassica napus
Comparative genomic analyses reveal the genetic basis of the yellow-seed trait in Brassica napus
Yellow-seed trait is preferred in rapeseed breeding as it can greatly improve seed oil yield and quality. Here, the authors assemble the genome of two rapeseed lines with yellow-seed and black-seed phenotypes, and clone an R2R3-MYB-type transcription factor encoding gene as a key regulator of seed color.
- Cunmin Qu
- , Meichen Zhu
- & Jiana Li
-
Article
| Open Access Whole genomic analysis reveals atypical non-homologous off-target large structural variants induced by CRISPR-Cas9-mediated genome editing
Whole genomic analysis reveals atypical non-homologous off-target large structural variants induced by CRISPR-Cas9-mediated genome editing
The safety of CRISPR-Cas9 editing is a concern. Here the authors use whole genomic analysis by 10x linked-read sequencing and optical genome mapping to interrogate the genome integrity after editing: they see large structural variants at on-target sites and unexpected large chromosomal deletions.
- Hsiu-Hui Tsai
- , Hsiao-Jung Kao
- & John Yu
-
Article
| Open Access Long-read whole-genome analysis of human single cells
Long-read whole-genome analysis of human single cells
Here the authors introduce a new method to study DNA in single cells by long-read sequencing. Their method gives a more complete view of the genomic structure of individual cells and allows to study genetic differences at the single-cell level.
- Joanna Hård
- , Jeff E. Mold
- & Adam Ameur
-
Article
| Open Access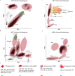 A genome-wide association study of blood cell morphology identifies cellular proteins implicated in disease aetiology
A genome-wide association study of blood cell morphology identifies cellular proteins implicated in disease aetiology
The authors identify genetic variation associated with properties of the internal biological structures of blood cells in hundreds of genes and show how such discoveries can be used to improve understanding of cellular mechanisms causing disease.
- Parsa Akbari
- , Dragana Vuckovic
- & William J. Astle
-
Perspective
| Open Access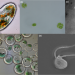 Extreme environments offer an unprecedented opportunity to understand microbial eukaryotic ecology, evolution, and genome biology
Extreme environments offer an unprecedented opportunity to understand microbial eukaryotic ecology, evolution, and genome biology
The ecology and evolution of eukaryotic microbes in extreme environments are poorly understood. In this Perspective, Rappaport and Oliverio summarize data from over 80 studies of protists in extreme environments and identify lineages of particular interest as targets for future research.
- Hannah B. Rappaport
- & Angela M. Oliverio
-
Article
| Open Access The genomic history of the indigenous people of the Canary Islands
The genomic history of the indigenous people of the Canary Islands
Here, the authors use paleogenomic data from the indigenous people of the Canary Islands to shed light on the Prehistory of North Africa, and on how insularity and resources availability shaped the genetic composition of this isolated population.
- Javier G. Serrano
- , Alejandra C. Ordóñez
- & Rosa Fregel
-
Article
| Open Access Ontogenetically distinct neutrophils differ in function and transcriptional profile in zebrafish
Ontogenetically distinct neutrophils differ in function and transcriptional profile in zebrafish
Neutrophil ontogeny in zebrafish may be a continuum or consist of distinct lineages. Here the authors characterise neutrophils derived from rostral blood island and caudal haematopoietic tissue lineages and show differential gene expression and function in steady state and during wound healing.
- Juan P. García-López
- , Alexandre Grimaldi
- & Carmen G. Feijoo
-
Article
| Open Access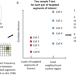 SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
Multiplexed DNA FISH technologies are powerful tools to reveal chromatin spatial organisation. Here, the authors developed SnapFISH, a computational pipeline to identify chromatin loops from multiplexed DNA FISH data.
- Lindsay Lee
- , Hongyu Yu
- & Ming Hu
-
Article
| Open Access Subtelomeric 5-enolpyruvylshikimate-3-phosphate synthase copy number variation confers glyphosate resistance in Eleusine indica
Subtelomeric 5-enolpyruvylshikimate-3-phosphate synthase copy number variation confers glyphosate resistance in Eleusine indica
Resistance to herbicide glyphosate can be evolved trough copy number variation (CNV) of its target gene EPSPS in goosegrass. Here, the authors assemble the genomes of glyphosate susceptible and resistance lines and provide evidence of sub-telomeric-repeat driven CNV of EPSPS could lead to glyphosate resistance.
- Chun Zhang
- , Nicholas A. Johnson
- & Eric L. Patterson
-
Article
| Open Access Evolutionary genomics of camouflage innovation in the orchid mantis
Evolutionary genomics of camouflage innovation in the orchid mantis
Camouflage is a widespread phenomenon in nature, and the orchid mantis is a particularly striking example. Here the authors use evolutionary genomics to uncover the genetic mechanisms behind the colour and morphology that produce innovative camouflage in the orchid mantis and dead leaf mantis.
- Guangping Huang
- , Lingyun Song
- & Fuwen Wei
-
Article
| Open Access TEQUILA-seq: a versatile and low-cost method for targeted long-read RNA sequencing
TEQUILA-seq: a versatile and low-cost method for targeted long-read RNA sequencing
The authors report TEQUILA-seq, a versatile, easy-to-implement, and low-cost method for targeted long-read RNA sequencing. TEQUILA-seq uncovers transcript isoforms and RNA mechanisms associated with human health and disease.
- Feng Wang
- , Yang Xu
- & Lan Lin
-
Article
| Open Access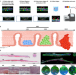 Spatial transcriptomics analysis of esophageal squamous precancerous lesions and their progression to esophageal cancer
Spatial transcriptomics analysis of esophageal squamous precancerous lesions and their progression to esophageal cancer
Understanding the molecular changes in the transition from esophageal squamous precancerous lesions (ESPL) to esophageal squamous cell carcinoma (ESCC) remains essential. Here, the authors analyze ESPL samples using spatial transcriptomics and reveal expression changes in TAGLN2 and CRNN during progression to ESCC.
- Xuejiao Liu
- , Simin Zhao
- & Zigang Dong
-
Article
| Open Access Droplet-based bisulfite sequencing for high-throughput profiling of single-cell DNA methylomes
Droplet-based bisulfite sequencing for high-throughput profiling of single-cell DNA methylomes
Single-cell DNA methylomic studies offer high resolution to differentiate cell subsets based on their epigenomic features. Here, the authors demonstrate Drop-BS, a droplet-based single-cell bisulfite sequencing library preparation method, for DNA methylome profiling.
- Qiang Zhang
- , Sai Ma
- & Chang Lu
-
Article
| Open Access Integrative analysis of transcriptome dynamics during human craniofacial development identifies candidate disease genes
Integrative analysis of transcriptome dynamics during human craniofacial development identifies candidate disease genes
Craniofacial disorders are among the most common congenital defects. Here, the authors examined the genetic causes of non-syndromic craniofacial disorders during human development through analysis of gene expression and epigenomics.
- Tara N. Yankee
- , Sungryong Oh
- & Justin Cotney
-
Article
| Open Access Genomic insight into domestication of rubber tree
Genomic insight into domestication of rubber tree
Understanding the genetic basis of rubber tree domestication is critical for improving natural rubber production. Here, the authors assemble the genome of the rubber tree clone CATAS8-79 and conduct population and genetic association analyses to reveal the function of phytosulfokine in regulating number of laticifer rings.
- Jinquan Chao
- , Shaohua Wu
- & Wei-Min Tian
-
Article
| Open Access Rapid gene content turnover on the germline-restricted chromosome in songbirds
Rapid gene content turnover on the germline-restricted chromosome in songbirds
Songbirds have an extra chromosome with unknown function found only in their germline. This study assembles and compares this chromosome in two closely related nightingale species, finding large differences in genetic content and only one conserved gene with probable essential function.
- Stephen A. Schlebusch
- , Jakub Rídl
- & Radka Reifová
-
Article
| Open Access Genetic variation in the immunoglobulin heavy chain locus shapes the human antibody repertoire
Genetic variation in the immunoglobulin heavy chain locus shapes the human antibody repertoire
Haplotype diversity in the human immunoglobulin heavy chain (IGH) locus is poorly characterized. Here, the authors use long-read sequencing to discover extensive IGH diversity and link germline variants to variation in the antibody repertoire.
- Oscar L. Rodriguez
- , Yana Safonova
- & Corey T. Watson
-
Article
| Open Access Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
In single-cell RNA-seq analyses, it would be critical to measure the relationships between genes. Here, the authors develop a framework for single-cell dimensionality reduction that incorporates gene-specific relationships - GeneVector -, and use it for tasks such as annotating cell types and analysing pathway variation after treatment.
- Nicholas Ceglia
- , Zachary Sethna
- & Andrew McPherson
-
Article
| Open Access Insertion sequence transposition inactivates CRISPR-Cas immunity
Insertion sequence transposition inactivates CRISPR-Cas immunity
CRISPR-Cas immunity systems safeguard prokaryotic genomes by inhibiting the invasion of mobile genetic elements. Here, the authors show that insertion sequences can efficiently insert into cas genes, thus inactivating CRISPR defenses and increasing bacterial susceptibility to foreign DNA invasion.
- Yong Sheng
- , Hengyu Wang
- & Qianjin Kang
-
Article
| Open Access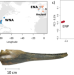 Ancient dolphin genomes reveal rapid repeated adaptation to coastal waters
Ancient dolphin genomes reveal rapid repeated adaptation to coastal waters
The chronology and mode of parallel evolution remain unclear. Here, the authors compare mid-Holocene and contemporary bottlenose dolphin adaptations between pelagic and coastal ecosystems with paleogenomics, finding rapid adaptation to newly emerged habitat from standing genetic variation.
- Marie Louis
- , Petra Korlević
- & Andrew D. Foote
-
Article
| Open Access High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
There is a need for methods that allow the analysis of single-cell long-read sequencing data without depending on known barcode lists or short-read sequencing. Here, the authors develop scNanoGPS, a tool that can independently deconvolute long reads into single cells and single molecules, and apply it on tumour and cell line data.
- Cheng-Kai Shiau
- , Lina Lu
- & Ruli Gao
-
Article
| Open Access Optimal strategies for learning multi-ancestry polygenic scores vary across traits
Optimal strategies for learning multi-ancestry polygenic scores vary across traits
There are challenges with transferring genetic risk scores from ancestry in which they were generated to another. Here, the authors investigate the use of multi-ancestry versus single-ancestry training sets to construct polygenic scores and find that the optimal strategy varies across traits.
- Brieuc Lehmann
- , Maxine Mackintosh
- & Chris Holmes
-
Article
| Open Access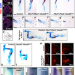 A spatio-temporally constrained gene regulatory network directed by PBX1/2 acquires limb patterning specificity via HAND2
A spatio-temporally constrained gene regulatory network directed by PBX1/2 acquires limb patterning specificity via HAND2
Many key developmental transcriptional regulators are broadly expressed but perform distinct functions in specific tissues. Here they show that ubiquitously expressed PBX factors gain limb bud functionality by interaction with HAND2, uncovering fundamental principles of cooperation between promiscuous and tissue-specific regulators to instruct developmental programs.
- Marta Losa
- , Iros Barozzi
- & Licia Selleri
-
Article
| Open Access Single-cell profiling of lncRNA expression during Ebola virus infection in rhesus macaques
Single-cell profiling of lncRNA expression during Ebola virus infection in rhesus macaques
Long non-coding RNAs (lncRNAs) play key roles in the immune response but their properties at the single-cell level are less well understood. Here, the authors characterize differential features of lncRNAs and protein-coding genes upon Ebola infection in macaques at single-cell resolution.
- Luisa Santus
- , Maria Sopena-Rios
- & Marta Melé
-
Article
| Open Access Genomic screening of 16 UK native bat species through conservationist networks uncovers coronaviruses with zoonotic potential
Genomic screening of 16 UK native bat species through conservationist networks uncovers coronaviruses with zoonotic potential
Certain bats species have previously been identified as ancestral sources of coronaviruses that infect humans but there is limited data on the genomic diversity or zoonotic potential of viruses infecting bats in the UK. Here, the authors use deep sequencing and in vitro assays to characterise coronaviruses recovered from 48 bat faecal samples.
- Cedric C. S. Tan
- , Jahcub Trew
- & Vincent Savolainen
-
Article
| Open Access Epistatic interactions between the high pathogenicity island and other iron uptake systems shape Escherichia coli extra-intestinal virulence
Epistatic interactions between the high pathogenicity island and other iron uptake systems shape Escherichia coli extra-intestinal virulence
The virulence of extra-intestinal pathogenic Escherichia coli is associated with multiple different genes in different lineages. Here, Royer et al. show that the emergence of virulence is associated with acquisition of the siderophore-encoding high-pathogenicity island (HPI), and full virulence is associated with the additional presence of the aer or sit operons.
- Guilhem Royer
- , Olivier Clermont
- & Erick Denamur
-
Article
| Open Access Phylodynamic of SARS-CoV-2 during the second wave of COVID-19 in Peru
Phylodynamic of SARS-CoV-2 during the second wave of COVID-19 in Peru
The second SARS-CoV-2 wave in Peru had a high case fatality rate with Lambda and Gamma causing most cases. Using phylodynamics, the authors here show that Lambda most likely originated in Peru from where it spread to other South American countries and that the center of Peru played a key role in transmission to other regions.
- Santiago Justo Arevalo
- , Carmen Sofia Uribe Calampa
- & Joao Renato Rebello Pinho
-
Article
| Open Access Genomics of cold adaptations in the Antarctic notothenioid fish radiation
Genomics of cold adaptations in the Antarctic notothenioid fish radiation
The notothenioid radiation is a remarkable group of fish adapted to life in the icy waters of the Southern Ocean. This study investigates the evolutionary history of this group and the basis of their adaption to cold environments through genomic analysis of 24 new genome assemblies.
- Iliana Bista
- , Jonathan M. D. Wood
- & Richard Durbin
-
Article
| Open Access Origin of minicircular mitochondrial genomes in red algae
Origin of minicircular mitochondrial genomes in red algae
While the organelle genome is commonly considered to be a single circular DNA molecule, extensive variation exists. Here, the authors report multipartite minicircular genomes in red algae and indicate an origin driven by recombination due to loss of DNA replication, recombination, and repair genes.
- Yongsung Lee
- , Chung Hyun Cho
- & Hwan Su Yoon
-
Article
| Open Access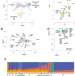 South Asian medical cohorts reveal strong founder effects and high rates of homozygosity
South Asian medical cohorts reveal strong founder effects and high rates of homozygosity
South Asia is home to almost 2 billion people but is extremely underrepresented in human genetics. This study uses genomes from ~5,000 South Asians to characterize genetic variation and help facilitate future South Asian genetic studies.
- Jeffrey D. Wall
- , J. Fah Sathirapongsasuti
- & Andrew S. Peterson
-
Article
| Open Access Transposon signatures of allopolyploid genome evolution
Transposon signatures of allopolyploid genome evolution
Assigning assembled chromosomes to subgenome in allopolypoid genome analysis is challenging. Here, the authors report a statistical formwork for identifying evolutionarily coherent subgneomes relying on transposable elements to group chromosomes into sets with shared ancestry and apply it in cyprinids, false flax and strawberry.
- Adam M. Session
- & Daniel S. Rokhsar
-
Article
| Open Access Linear time complexity de novo long read genome assembly with GoldRush
Linear time complexity de novo long read genome assembly with GoldRush
Current state-of-the-art de novo long read genome assemblers follow the Overlap-Layout-Consensus paradigm. GoldRush departs from this paradigm, generating highly contiguous assemblies with linear time complexity and using an order of magnitude less RAM than state-of-the-art methods.
- Johnathan Wong
- , Lauren Coombe
- & Inanç Birol
-
Article
| Open Access Distinct genomic routes underlie transitions to specialised symbiotic lifestyles in deep-sea annelid worms
Distinct genomic routes underlie transitions to specialised symbiotic lifestyles in deep-sea annelid worms
Annelid worms have colonised extreme ecological niches, such as hydrothermal vents and whale falls thanks to symbiotic bacteria. This study finds that Osedax worms and the related Vestimentifera have evolved different genomic adaptations to sustain their bacterial symbioses and exploit different resources, such as decaying bone.
- Giacomo Moggioli
- , Balig Panossian
- & José M. Martín-Durán
-
Article
| Open Access Base-resolution UV footprinting by sequencing reveals distinctive damage signatures for DNA-binding proteins
Base-resolution UV footprinting by sequencing reveals distinctive damage signatures for DNA-binding proteins
Proteins binding to DNA can locally alter DNA damage formation by UV light. Here, Elliott et al. generate high-resolution quantitative UV damage profiles for genomic regions of interest, revealing distinctive damage signatures for specific proteins and elevated UV damage at melanoma mutation hotspots.
- Kerryn Elliott
- , Vinod Kumar Singh
- & Erik Larsson
-
Article
| Open Access Single-nucleus RNA-sequencing of autosomal dominant Alzheimer disease and risk variant carriers
Single-nucleus RNA-sequencing of autosomal dominant Alzheimer disease and risk variant carriers
Mutations in amyloid precursor protein (APP) and presenilin 1 (PSEN1) cause autosomal dominant AD (ADAD). Here, the authors perform single-nucleus RNA-sequencing of ADAD and other disease risk modifying variant carriers and report altered expression states of specific brain cell types.
- Logan Brase
- , Shih-Feng You
- & Oscar Harari
-
Article
| Open Access Identifying high-impact variants and genes in exomes of Ashkenazi Jewish inflammatory bowel disease patients
Identifying high-impact variants and genes in exomes of Ashkenazi Jewish inflammatory bowel disease patients
Inflammatory bowel disease (IBD) is highly prevalent among the Ashkenazi Jewish population. Here, the authors identify novel IBD-associated variants and genes, validated by transcriptomic and phenome-wide associations.
- Yiming Wu
- , Kyle Gettler
- & Yuval Itan
-
Matters Arising
| Open AccessReply to “Subgenome-aware analyses suggest a reticulate allopolyploidization origin in three Papaver genomes”
- Xiaofei Yang
- , Shenghan Gao
- & Kai Ye
-
Article
| Open Access Multi-omic underpinnings of epigenetic aging and human longevity
Multi-omic underpinnings of epigenetic aging and human longevity
Here, the authors integrate genomic, bulk and single-cell transcriptomic, and metabolomic data sets to compare the biological underpinning of four epigenetic clocks and human longevity, offering novel insights into aging biology.
- Lucas A. Mavromatis
- , Daniel B. Rosoff
- & Falk W. Lohoff
-
Matters Arising
| Open Access Subgenome-aware analyses suggest a reticulate allopolyploidization origin in three Papaver genomes
Subgenome-aware analyses suggest a reticulate allopolyploidization origin in three Papaver genomes
- Ren-Gang Zhang
- , Chaoxia Lu
- & Wei Zhao
-
Article
| Open Access Comparative epidemic expansion of SARS-CoV-2 variants Delta and Omicron in the Brazilian State of Amazonas
Comparative epidemic expansion of SARS-CoV-2 variants Delta and Omicron in the Brazilian State of Amazonas
The Amazonas region has been the most heavily affected by COVID-19 in Brazil. In this study, the authors conduct phylodynamic analyses to assess SARS-CoV-2 lineage replacement dynamics in the region and infer the impact of population immunity on the spread and severity of the Delta and Omicron variants.
- Ighor Arantes
- , Gonzalo Bello
- & Felipe Gomes Naveca
-
Article
| Open Access Analyses of a chromosome-scale genome assembly reveal the origin and evolution of cultivated chrysanthemum
Analyses of a chromosome-scale genome assembly reveal the origin and evolution of cultivated chrysanthemum
Chrysanthemum is an important ornamental species with great economic value. Here, the authors assemble the haploid genome of C. morifolium, reveal its segmental allopolyploid genomic composition (AA’B), and identify candidate genes associated with flower development, petal shape, and flower colour.
- Aiping Song
- , Jiangshuo Su
- & Fadi Chen
-
Article
| Open Access The evolution and international spread of extensively drug resistant Shigella sonnei
The evolution and international spread of extensively drug resistant Shigella sonnei
An increase in shigellosis cases among men who have sex with men in the United Kingdom has been linked to an extensively drug-resistant strain of Shigella sonnei. In this genomic epidemiology study, the authors investigate the genetic basis, evolutionary history, and international dissemination of the outbreak strain.
- Lewis C. E. Mason
- , David R. Greig
- & Kate S. Baker
-
Article
| Open Access A comprehensive platform for analyzing longitudinal multi-omics data
A comprehensive platform for analyzing longitudinal multi-omics data
The analysis of longitudinal bulk and single-cell multi-omics data is a highly complex task. Here, the authors introduce PALMO, a software platform with five modules to analyse longitudinal bulk and single-cell multi-omics data, which is extensively tested in external datasets that include multiple omics modalities.
- Suhas V. Vasaikar
- , Adam K. Savage
- & Xiao-jun Li
-
Article
| Open Access Pan-cancer classification of single cells in the tumour microenvironment
Pan-cancer classification of single cells in the tumour microenvironment
The accuracy and granularity of classifying cell types in the tumour microenvironment (TME) from single-cell RNA-seq data is impacted by heterogeneity among cancer cells and similarities among functionally related immune cells. Here, the authors develop scATOMIC, a tumour and TME cell type classifier based on a hierarchical approach that can be applied to pan-cancer datasets.
- Ido Nofech-Mozes
- , David Soave
- & Sagi Abelson

 Deciphering complex breakage-fusion-bridge genome rearrangements with Ambigram
Deciphering complex breakage-fusion-bridge genome rearrangements with Ambigram