Featured
-
-
Article
| Open Access SDE2 integrates into the TIMELESS-TIPIN complex to protect stalled replication forks
SDE2 integrates into the TIMELESS-TIPIN complex to protect stalled replication forks
The fork protection complex (FPC), including the proteins TIMELESS and TIPIN, stabilizes the replisome to ensure unperturbed fork progression during DNA replication. Here the authors reveal that that SDE2, a PCNA-associated protein, plays an important role in maintaining active replication and protecting stalled forks by regulating the replication fork protection complex (FPC).
- Julie Rageul
- , Jennifer J. Park
- & Hyungjin Kim
-
Article
| Open Access Polymerization and editing modes of a high-fidelity DNA polymerase are linked by a well-defined path
Polymerization and editing modes of a high-fidelity DNA polymerase are linked by a well-defined path
In high fidelity DNA polymerases the exonuclease site is distal from the polymerization site and it is unknown how the primer strand travels between the two sites when mis-incorporated nucleotides must be removed. Here, the authors perform MD simulations and identify an optimal path for DNA primer strand translocation in the E. coli replicative DNA polymerase III and characterise the kinetics and dynamics of the Pol III pol-to-exo mode transition, which is validated with mutagenesis experiments.
- Thomas Dodd
- , Margherita Botto
- & Ivaylo Ivanov
-
Article
| Open Access Mechanisms of telomerase inhibition by oxidized and therapeutic dNTPs
Mechanisms of telomerase inhibition by oxidized and therapeutic dNTPs
Telomerase enzymes add telomeric repeats to the end of linear chromosomes. Here the authors reveal mechanisms by which oxidized dNTPs and therapeutic dNTPs inhibit telomerase-mediated telomere elongation.
- Samantha L. Sanford
- , Griffin A. Welfer
- & Patricia L. Opresko
-
Article
| Open Access Evolution of DNA replication origin specification and gene silencing mechanisms
Evolution of DNA replication origin specification and gene silencing mechanisms
Contrary to most eukaryotes that lack sequence-specific origins of replication, S. cerevisiae origins are defined by specific DNA sequence motifs. Here the authors reveal that multiple subunits of ORC, including Orc2 and Orc4, contribute to the sequence-specificity of origins in S. cerevisiae.
- Y. Hu
- , A. Tareen
- & B. Stillman
-
Matters Arising
| Open Access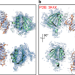 Reply to: “Does PCNA diffusion on DNA follow a rotation-coupled translation mechanism?”
Reply to: “Does PCNA diffusion on DNA follow a rotation-coupled translation mechanism?”
- Matteo De March
- , Silvia Onesti
- & Alfredo De Biasio
-
Article
| Open Access Cross-regulation of viral kinases with cyclin A secures shutoff of host DNA synthesis
Cross-regulation of viral kinases with cyclin A secures shutoff of host DNA synthesis
Herpesviruses code for conserved protein kinases (CHPKs) that exert several regulatory functions by interacting with cellular factors. Here, the authors use affinity purification mass spectrometry (AP–MS) to identify differential interaction partners of CHPKs from seven different human herpesviruses, finding Cyclin A and associated factors as a specific signature of β-herpesvirus kinases.
- Boris Bogdanow
- , Max Schmidt
- & Lüder Wiebusch
-
Article
| Open Access A predictable conserved DNA base composition signature defines human core DNA replication origins
A predictable conserved DNA base composition signature defines human core DNA replication origins
In metazoan the DNA sequence elements characterizing origin specification are unknown. By generating and analysing 19 SNS-seq datasets from different human cell types, the authors reveal a class and features of Core origins of replication which can be predicted by an algorithm.
- Ildem Akerman
- , Bahar Kasaai
- & Marcel Méchali
-
Article
| Open Access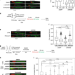 Resolution of R-loops by INO80 promotes DNA replication and maintains cancer cell proliferation and viability
Resolution of R-loops by INO80 promotes DNA replication and maintains cancer cell proliferation and viability
In mammalian cells, during transcription and replication, RNA:DNA hybrid structures known as R-loops can arise, posing as obstacles to replication fork progression. Here the authors reveal that the ATP-dependent chromatin remodelling INO80 complex promotes resolution of R-loops to prevent replication associated DNA damage in cancer cells.
- Lisa Prendergast
- , Urszula L. McClurg
- & Manolis Papamichos-Chronakis
-
Article
| Open Access Antibiotic susceptibility signatures identify potential antimicrobial targets in the Acinetobacter baumannii cell envelope
Antibiotic susceptibility signatures identify potential antimicrobial targets in the Acinetobacter baumannii cell envelope
A unique cell envelope contributes to the antibiotic resistance of the pathogen Acinetobacter baumannii. Here, Geisinger et al. identify A. baumannii mutants with altered antibiotic susceptibility, infer the function of uncharacterized proteins involved in envelope synthesis, and predict antibiotic synergies.
- Edward Geisinger
- , Nadav J. Mortman
- & Ralph R. Isberg
-
Article
| Open Access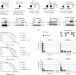 Warsaw Breakage Syndrome associated DDX11 helicase resolves G-quadruplex structures to support sister chromatid cohesion
Warsaw Breakage Syndrome associated DDX11 helicase resolves G-quadruplex structures to support sister chromatid cohesion
WABS patient derived cells display loss of sister chromatid cohesion. Here the authors by analyzing WABS patient derived cells, reveal a role of the DDX11 helicase in resolving G-Quadruplex structures to support sister chromatid cohesion.
- Janne J. M. van Schie
- , Atiq Faramarz
- & Job de Lange
-
Article
| Open Access Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6
Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6
The origin recognition complex (ORC) is essential for loading the Mcm2–7 replicative helicase onto DNA during DNA replication initiation. Here, the authors describe several cryo-electron microscopy structures of Drosophila ORC bound to DNA and its cofactor Cdc6 and also report an in vitro reconstitution system for Drosophila Mcm2–7 loading, revealing unexpected features of ORC’s DNA binding and remodeling mechanism during Mcm2–7 loading.
- Jan Marten Schmidt
- & Franziska Bleichert
-
Article
| Open Access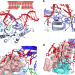 Molecular basis for DNA repair synthesis on short gaps by mycobacterial Primase-Polymerase C
Molecular basis for DNA repair synthesis on short gaps by mycobacterial Primase-Polymerase C
Mycobacteria Prim-PolC performs short gap synthesis following removal of lesions during excision repair. Here the authors resolve crystal structures of pre- and post-catalytic Prim-PolC complexes bound to gapped DNA substrates to define its mechanism.
- Nigel C. Brissett
- , Katerina Zabrady
- & Aidan J. Doherty
-
Article
| Open Access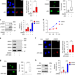 BRD4 prevents the accumulation of R-loops and protects against transcription–replication collision events and DNA damage
BRD4 prevents the accumulation of R-loops and protects against transcription–replication collision events and DNA damage
In order to avoid transcription-replication conflicts (TRCs) on shared DNA templates, cell must maintain strict spatiotemporal co-ordination of transcription with replication. Here the authors uncover a role for BRD4 in preventing TRCs and DNA damage checkpoint signaling in oncogenic cells.
- Fred C. Lam
- , Yi Wen Kong
- & Michael B. Yaffe
-
Article
| Open Access DONSON and FANCM associate with different replisomes distinguished by replication timing and chromatin domain
DONSON and FANCM associate with different replisomes distinguished by replication timing and chromatin domain
Eukaryotic replisomes are multiprotein complexes. Here the authors reveal two distinct stressed replisomes, associated with DONSON and FANCM, displaying a bias in replication timing and chromatin domain.
- Jing Zhang
- , Marina A. Bellani
- & Michael M. Seidman
-
Article
| Open Access Ethanol exposure increases mutation rate through error-prone polymerases
Ethanol exposure increases mutation rate through error-prone polymerases
Whereas the toxic effects of ethanol are well-documented, the underlying mechanism is obscure. This study uses the eukaryotic model S. cerevisiae to reveal how exposure to sublethal ethanol concentrations causes DNA replication stress and an increased mutation rate.
- Karin Voordeckers
- , Camilla Colding
- & Kevin J. Verstrepen
-
Article
| Open Access 3D genome organization contributes to genome instability at fragile sites
3D genome organization contributes to genome instability at fragile sites
Common fragile sites are regions susceptible to replication stress and are prone to chromosomal instability. Here, the authors, by analyzing the contribution of 3D chromatin organization, identify and characterize a fragility signature and precisely map these fragility regions.
- Dan Sarni
- , Takayo Sasaki
- & Batsheva Kerem
-
Article
| Open Access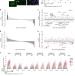 Sequential role of RAD51 paralog complexes in replication fork remodeling and restart
Sequential role of RAD51 paralog complexes in replication fork remodeling and restart
Replication stress has been associated with transient remodelling of replication intermediates into reversed forks, followed by efficient fork restart. Here the authors systematically analyse the role of RAD51 paralogs in these transactions, providing insights on the mechanistic role of different complexes of these proteins.
- Matteo Berti
- , Federico Teloni
- & Massimo Lopes
-
Article
| Open Access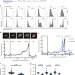 E2F-dependent transcription determines replication capacity and S phase length
E2F-dependent transcription determines replication capacity and S phase length
DNA replication is tightly regulated during S phase of the cell cycle to ensure timely and accurate genome duplication. Here, the authors reveal that E2F-dependent transcription determines the maximal amount of DNA a cell is able to synthesise per unit time throughout S phase.
- Betheney R. Pennycook
- , Eva Vesela
- & Robertus A. M. de Bruin
-
Article
| Open Access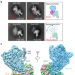 Structure of the polymerase ε holoenzyme and atomic model of the leading strand replisome
Structure of the polymerase ε holoenzyme and atomic model of the leading strand replisome
DNA polymerase epsilon (Pol ε) is responsible for leading strand synthesis during DNA replication. Here the authors use Cryo-EM to describe the architecture of the Pol ε holoenzyme and to provide an atomic model for the leading strand replisome.
- Zuanning Yuan
- , Roxana Georgescu
- & Huilin Li
-
Article
| Open Access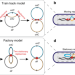 Direct observation of independently moving replisomes in Escherichia coli
Direct observation of independently moving replisomes in Escherichia coli
How chromosome replication and segregation is organised in E. coli is a matter of debate. Here the authors visualise the bacterial chromosome and the replisomes during DNA replication, providing support for a previously suggested train track model.
- Aleksandre Japaridze
- , Christos Gogou
- & Cees Dekker
-
Article
| Open Access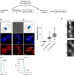 Budding yeast complete DNA synthesis after chromosome segregation begins
Budding yeast complete DNA synthesis after chromosome segregation begins
In the S phase of the cell cycle, the full genome needs to be replicated before cell division occurs. Here, authors show that in budding yeast DNA synthesis is completed after chromosome segregation begins.
- Tsvetomira Ivanova
- , Michael Maier
- & Manuel Mendoza
-
Article
| Open Access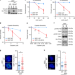 Ubiquitinated-PCNA protects replication forks from DNA2-mediated degradation by regulating Okazaki fragment maturation and chromatin assembly
Ubiquitinated-PCNA protects replication forks from DNA2-mediated degradation by regulating Okazaki fragment maturation and chromatin assembly
PCNA is essential for DNA replication and cellular proliferation. Here, the authors reveal that PCNA ubiquitination protects stalled replication forks from DNA2-mediated degradation via regulation of Okazaki fragment maturation and chromatin assembly.
- Tanay Thakar
- , Wendy Leung
- & George-Lucian Moldovan
-
Article
| Open Access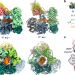 Structural basis for the increased processivity of D-family DNA polymerases in complex with PCNA
Structural basis for the increased processivity of D-family DNA polymerases in complex with PCNA
Replicative DNA polymerases (DNAPs) have evolved the ability to copy the genome with high processivity and fidelity. Here, the authors present a cryo-EM structure of the DNA-bound PolD–PCNA complex from Pyrococcus abyssi to reveal the molecular basis for the interaction and cooperativity between a replicative DNAP and PCNA.
- Clément Madru
- , Ghislaine Henneke
- & Ludovic Sauguet
-
Article
| Open Access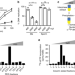 SSRP1-mediated histone H1 eviction promotes replication origin assembly and accelerated development
SSRP1-mediated histone H1 eviction promotes replication origin assembly and accelerated development
During embryonic development, it is vital to maintain rapid genome duplication. Here, the authors shed light on the mechanism by revealing that SSRP1 stimulates replication origin assembly on somatic nuclei in Xenopus laevis egg extract by promoting histone H1 eviction from somatic chromatin.
- Lucia Falbo
- , Erica Raspelli
- & Vincenzo Costanzo
-
Article
| Open Access In vitro self-replication and multicistronic expression of large synthetic genomes
In vitro self-replication and multicistronic expression of large synthetic genomes
A main objective of synthetic biology is the creation of chemical systems capable of replication and evolution. Here, the authors demonstrate combined self-replication and expression of multipartite genomes in vitro.
- K. Libicher
- , R. Hornberger
- & H. Mutschler
-
Article
| Open Access H3K4 methylation at active genes mitigates transcription-replication conflicts during replication stress
H3K4 methylation at active genes mitigates transcription-replication conflicts during replication stress
Transcription-replication conflicts (TRC) can contribute to genome instability. Here the authors reveal that under replication stress H3K4 methylation can play a role in TRC prevention.
- Shin Yen Chong
- , Sam Cutler
- & Cheng-Fu Kao
-
Article
| Open Access The nuclear pore complex prevents sister chromatid recombination during replicative senescence
The nuclear pore complex prevents sister chromatid recombination during replicative senescence
The Nuclear Pore Complex has been linked to DNA damage processing. Here the authors reveal that the Nup1 C-terminus is critical for the relocalization of eroded telomeres to nuclear pores and that modification of Nup1 promotes sister chromatid recombination and unleashes a new telomere maintenance mechanism.
- Paula Aguilera
- , Jenna Whalen
- & Vincent Géli
-
Article
| Open Access ATAD5 promotes replication restart by regulating RAD51 and PCNA in response to replication stress
ATAD5 promotes replication restart by regulating RAD51 and PCNA in response to replication stress
How the replisome machinery contributes to fork stability under replication stress is currently not clear. Here the authors reveal a role for ATAD5 in maintaining genome integrity during replication stress by promoting replication restart through RAD51/PCNA regulation.
- Su Hyung Park
- , Nalae Kang
- & Kyungjae Myung
-
Article
| Open Access Transcription-mediated organization of the replication initiation program across large genes sets common fragile sites genome-wide
Transcription-mediated organization of the replication initiation program across large genes sets common fragile sites genome-wide
Common Fragile Sites (CFSs) are chromosome regions prone to breakage upon replication stress known to drive chromosome rearrangements during oncogenesis. Here the authors use genome-wide and single cell techniques to assess how replication timing and transcriptional activity correlate with genome stability.
- Olivier Brison
- , Sami El-Hilali
- & Chun-Long Chen
-
Comment
| Open Access A model of DNA damage response activation at stalled replication forks by SPRTN
A model of DNA damage response activation at stalled replication forks by SPRTN
The process of DNA replication is threatened by many factors, including DNA lesions, and machineries acting as obstacles. Here we discuss and speculate on a recently proposed mechanism of DNA damage response activation in response to lesions that challenge the progression of DNA replication forks.
- Christopher Bruhn
- & Marco Foiani
-
Article
| Open Access Two-tiered enforcement of high-fidelity DNA ligation
Two-tiered enforcement of high-fidelity DNA ligation
DNA ligases catalyze the joining of DNA strands to complete DNA replication, recombination and repair transactions. Here the authors present X-ray structures and kinetic analyses of LIG1 complexes with undamaged and oxidatively damaged DNA that unveil determinants of LIG1 substrate recognition and enzymatic fidelity.
- Percy P. Tumbale
- , Thomas J. Jurkiw
- & R. Scott Williams
-
Article
| Open Access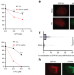 ZFP161 regulates replication fork stability and maintenance of genomic stability by recruiting the ATR/ATRIP complex
ZFP161 regulates replication fork stability and maintenance of genomic stability by recruiting the ATR/ATRIP complex
The ATR pathway is active during DNA replication stress to maintain genome stability. Here the authors reveal the role of the zinc finger containing protein 161 (ZFP161) to facilitate replication fork stability by acting as a scaffold to facilitate the interaction between RPA and ATR/ATRIP.
- Wootae Kim
- , Fei Zhao
- & Jian Yuan
-
Article
| Open Access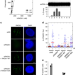 Non-enzymatic roles of human RAD51 at stalled replication forks
Non-enzymatic roles of human RAD51 at stalled replication forks
RAD51 has been implicated in replication fork processing and restart in response to replication stress. Here, authors reveal mechanistic aspects of non-enzymatic roles of RAD51 for fork reversal and cooperation with FBH1.
- Jennifer M. Mason
- , Yuen-Ling Chan
- & Douglas K. Bishop
-
Article
| Open Access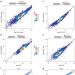 Distinct transcriptional roles for Histone H3-K56 acetylation during the cell cycle in Yeast
Distinct transcriptional roles for Histone H3-K56 acetylation during the cell cycle in Yeast
H3K56Ac promotes nucleosome turnover at promoter-proximal locations and assembly of new nucleosomes during S phase. Here the authors find that H3K56Ac is both a genome-wide activator of transcription in yeast while also repressing promiscuous transcription that occurs after replication fork passage.
- Salih Topal
- , Pauline Vasseur
- & Craig L. Peterson
-
Article
| Open Access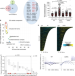 MRE11-RAD50-NBS1 promotes Fanconi Anemia R-loop suppression at transcription–replication conflicts
MRE11-RAD50-NBS1 promotes Fanconi Anemia R-loop suppression at transcription–replication conflicts
Accumulations of R-loops can lead to genome instability. Here the authors reveal a role for the MRN complex in suppressing R-loops and associated DNA damage at transcription–replication conflicts.
- Emily Yun-Chia Chang
- , Shuhe Tsai
- & Peter C. Stirling
-
Article
| Open Access Replication stress induces mitotic death through parallel pathways regulated by WAPL and telomere deprotection
Replication stress induces mitotic death through parallel pathways regulated by WAPL and telomere deprotection
Mitotic catastrophe is a regulated mechanism that responds to aberrant mitoses leading to removal of damaged cells. Here the authors reveal how replication stress induces mitotic death through pathways regulated by WAPL and telomere deprotection.
- V. Pragathi Masamsetti
- , Ronnie Ren Jie Low
- & Anthony J. Cesare
-
Article
| Open Access Roles for DNA polymerase δ in initiating and terminating leading strand DNA replication
Roles for DNA polymerase δ in initiating and terminating leading strand DNA replication
DNA polymerases epsilon and delta, respectively, perform the majority of leading and lagging strand replication of the eukaryotic nuclear genome. Here the authors map the ribonucleotide fingerprints of the polymerases to show the special roles of polymerase delta on both strands during replication initiation and termination.
- Zhi-Xiong Zhou
- , Scott A. Lujan
- & Thomas A. Kunkel
-
Article
| Open Access Mild replication stress causes chromosome mis-segregation via premature centriole disengagement
Mild replication stress causes chromosome mis-segregation via premature centriole disengagement
Chromosome instability can be caused by replication stress, although the mechanism is unclear. Here, the authors show that inducing mild replication stress in cancerous and non-cancerous cell lines leads to centriole disengagement and the subsequent formation of lagging chromosomes and micronuclei.
- Therese Wilhelm
- , Anna-Maria Olziersky
- & Patrick Meraldi
-
Article
| Open Access Involvement of G-quadruplex regions in mammalian replication origin activity
Involvement of G-quadruplex regions in mammalian replication origin activity
Origins of replications are associated with potential G quadruplexes forming structures (G4s). Here the authors reveal the functional role of G4 elements in DNA replication initiation.
- Paulina Prorok
- , Marie Artufel
- & Marcel Méchali
-
Article
| Open Access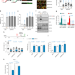 SPRTN protease and checkpoint kinase 1 cross-activation loop safeguards DNA replication
SPRTN protease and checkpoint kinase 1 cross-activation loop safeguards DNA replication
Cells deficient in SPRTN protease activity exhibit severe DNA-protein crosslink induced replication stress and genome instability. Here the author reveal a functional link between the SPRTN protease and the CHK1 kinase during physiological DNA replication.
- Swagata Halder
- , Ignacio Torrecilla
- & Kristijan Ramadan
-
Article
| Open Access DNA translocation mechanism of the MCM complex and implications for replication initiation
DNA translocation mechanism of the MCM complex and implications for replication initiation
Eukaryotes and archaea use a heximeric ring-shaped MCM helicase to unwind the DNA template during replication. Here the authors present a crystal structure of the MCM complex from archaeon S. solfataricus bound to single-stranded DNA, and to a combination of ADP, and ATP-mimic, ADP-BeF3.
- Martin Meagher
- , Leslie B. Epling
- & Eric J. Enemark
-
Article
| Open Access The ORC ubiquitin ligase OBI1 promotes DNA replication origin firing
The ORC ubiquitin ligase OBI1 promotes DNA replication origin firing
DNA replication is initiated at defined genomic sites called origins of replication following ORC pre-replicative complex assembly. Here the authors identify a protein ubiquitylating ORC that is involved in origin activation and may act as a selector of origins to be fired.
- Philippe Coulombe
- , Joelle Nassar
- & Marcel Méchali
-
Article
| Open Access qDSB-Seq is a general method for genome-wide quantification of DNA double-strand breaks using sequencing
qDSB-Seq is a general method for genome-wide quantification of DNA double-strand breaks using sequencing
Measuring relative frequencies of DNA double-strand breaks between loci does not provide the full physiological relevance of those breaks. Here Rowicka and colleagues present qDSB-Seq method which uses spike-in double-strand breaks induced by a restriction enzyme to accurately quantify DNA damage.
- Yingjie Zhu
- , Anna Biernacka
- & Maga Rowicka
-
Article
| Open Access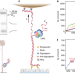 The mechanism of DNA unwinding by the eukaryotic replicative helicase
The mechanism of DNA unwinding by the eukaryotic replicative helicase
How the eukaryotic helicase unzips DNA during replication is not well understood. By measuring the real-time motion of purified CMG unwinding DNA with magnetic tweezers, the authors reveal the dynamics where isolated CMG unwinds via a biased random walk with proclivity to pause.
- Daniel R. Burnham
- , Hazal B. Kose
- & Hasan Yardimci
-
Article
| Open Access A requirement for STAG2 in replication fork progression creates a targetable synthetic lethality in cohesin-mutant cancers
A requirement for STAG2 in replication fork progression creates a targetable synthetic lethality in cohesin-mutant cancers
Cohesin complex mediates cohesion of sister chromatids and DNA loop formation. Here the authors show another role of cohesin in replication fork progression, suggesting a potential therapeutic strategy for cohesin-mutant cancers.
- Gourish Mondal
- , Meredith Stevers
- & David A. Solomon
-
Article
| Open Access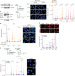 Rad52 prevents excessive replication fork reversal and protects from nascent strand degradation
Rad52 prevents excessive replication fork reversal and protects from nascent strand degradation
Stabilisation of stalled replication forks prevents excessive fork reversal and genome instability. Here authors reveal a RAD52-dependent replication fork protection mechanism.
- Eva Malacaria
- , Giusj Monia Pugliese
- & Pietro Pichierri
-
Article
| Open Access Overexpression of Claspin and Timeless protects cancer cells from replication stress in a checkpoint-independent manner
Overexpression of Claspin and Timeless protects cancer cells from replication stress in a checkpoint-independent manner
Oncogene-induced replication stress (RS) promotes cancer development. Here, the authors report that cancer cells adapt to oncogene-induced RS by overexpressing downstream components of ATR-CHK1 pathway, Claspin and Timeless, which have protective role at the replication forks independent of their checkpoint function.
- Julien N. Bianco
- , Valérie Bergoglio
- & Philippe Pasero
-
Article
| Open Access Maintenance of epigenetic landscape requires CIZ1 and is corrupted in differentiated fibroblasts in long-term culture
Maintenance of epigenetic landscape requires CIZ1 and is corrupted in differentiated fibroblasts in long-term culture
The inactive X chromosome (Xi) is a model for establishment and maintenance of repressed chromatin and the function of polycomb repressive complexes. Here the authors show that Xi transiently relocates from the nuclear periphery during replication in a CIZ1-dependent manner, which plays a role in maintaining PRC-mediated repressed chromatin.
- Emma R. Stewart
- , Robert M. L. Turner
- & Dawn Coverley
-
Article
| Open Access Replication timing and epigenome remodelling are associated with the nature of chromosomal rearrangements in cancer
Replication timing and epigenome remodelling are associated with the nature of chromosomal rearrangements in cancer
The connection between DNA replication timing and changes that occur to the epigenome in cancer are still poorly understood. Here, the authors perform Repli-Seq and integrated epigenome analyses and find that genomic regions that undergo long-range epigenetic deregulation in prostate cancer also show concordant differences in replication timing.
- Qian Du
- , Saul A. Bert
- & Susan J. Clark

 Mus81-Mms4 endonuclease is an Esc2-STUbL-Cullin8 mitotic substrate impacting on genome integrity
Mus81-Mms4 endonuclease is an Esc2-STUbL-Cullin8 mitotic substrate impacting on genome integrity