Abstract
In this study, we have investigated genome-wide occurrence of Histone Acetyltransferases (HATs) in genomes of Mus musculus and Danio rerio on the basis of presence of HAT domain. Our study identified a group of proteins that lacks characteristic features of known HAT families, relatively smaller in size and has no other associated domains. Most of the proteins in this unclassified group are Camello proteins, which are not yet known and classified as functional HATs. Our in vitro and in vivo analysis revealed that Camello family proteins are active HATs and exhibit specificity towards histone H4. Interestingly, Camello proteins are among the first identified HATs showing perinuclear localization. Moreover, Camello proteins are evolutionarily conserved in all chordates and are observed for the first time in cnidarians in phylogeny. Furthermore, knockdown of Camello protein (CMLO3) in zebrafish embryos exhibited defects in axis elongation and head formation. Thus, our study identified a novel family of active HATs that is specific for histone H4 acetylation, exhibits perinuclear localization and is essential for zebrafish development.
Similar content being viewed by others
Introduction
Gene expression in eukaryotes is a tightly controlled process involving a complex interplay between chromatin proteins and transcription factors. The functional availability of these factors and accessibility of DNA sequence define the state of gene activation or repression. DNA in chromatin is wrapped around histone octamers comprising of two copies each of the four core histone proteins (H2A, H2B, H3 and H4) to form discrete nucleosome units. The N-terminal tails of these core-histones protrude from the nucleosome particles and are subjected to various post-translational modifications such as acetylation, methylation, phosphorylation and ubiqutination1,2.
Histone acetylation by histone acetyltransferases (HATs) is one of the most extensively studied covalent histone modifications. HATs modify physico–chemical properties of core histones through acetylation, influence the nucleosome structure and participate in transcription regulation. However, many HATs can act on non-histone proteins (cytoplasmic as well as nuclear) and are now renamed as lysine acetyltransferases (KATs)3. Acetylation of core-histone and non-histone proteins is correlated with various cellular processes such as transcription regulation, chromatin assembly, DNA repair and cell cycle progression4.
Characterization of HATs on the basis of protein sequence and domain organization reveals five distinct families of HATs5. (i) Largest of these families is the GNAT (GCN5-related N-acetyltransferase) family whose members share a highly conserved acetylation-related structural motif. GCN5, one of the members of the GNAT family is the best-characterized HAT protein and serves as a prototype for histone acetyltransferase studies. One of the characteristic features of the GNAT family is a carboxy-terminal bromo-domain, which helps in targeting proteins to the substrate6. GNAT family proteins are also known to acetylate non-histone proteins as well as small molecules7. (ii) Another family is the MYST (MOZ, Ybf2/Sas3, Sas2 and Tip60) family, which also has an acetylation-related structural motif. Many of the MYST family proteins contain zinc fingers as well as chromo-domain5. Presence of chromo-domain in the MYST family suggests that they might interact with the heterochromatin-associated proteins8. GNAT and MYST families contain dozens of lysine acetyltransferase enzymes and are mostly part of multi-subunit transcriptional co-activator complexes. (iii) The P300/CBP (CREB-binding protein) family consists of two paralogous proteins, P300 and CBP. These two proteins have interchangeable functions. Members of the P300/CBP family contain many functional domains including acetylation-related structural motif which is involved in acetyl-CoA binding, three zinc finger regions and a bromo-domain. P300/CBP act as co-activators and harbor domains for interaction with many transcription factors9. (iv) The fourth group of HATs is the basal transcription factor family, which is related to mammalian TAFII250, the largest subunit of the transcription factor complex TFIID2,10. Basal transcription factor family proteins also act as HATs but do not harbor acetylation related structural motif. (v) Last of the HAT families is the nuclear receptor cofactors family, which is largely specific to mammals6. Members of this family include nuclear receptor co-activators such as steroid receptor co-activators (SRC1) and clock circadian regulator (CLOCK). This family of HATs is also functionally known to act as HAT but they do not have any acetylation related structural motif11,12,13.
Here, we performed genome-wide survey of lysine acetyltransferase proteins in mouse and zebrafish genomes. Our genome-wide bioinformatics analysis identified a novel family of HATs, namely Camello proteins, which harbors the HAT domain. We demonstrated that Camello-family of proteins are active HATs and have specificity towards histone H4 acetylation. We also show that Camello proteins have perinuclear localization and their overexpression leads to increased acetylation of histone H4. Finally, we demonstrated in vivo role of camello histone acetyltransferases by knockdown of CMLO3 in zebrafish embryos. Morpholino-mediated knockdown of CMLO3 exhibited defects in axis elongation and head formation, suggesting its critical role in zebrafish development.
Results
Genome-wide identification of HATs in mouse and zebrafish genomes
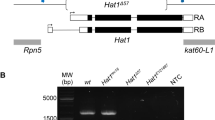
The mouse genome sequence was searched for homologs of known histone acetyltransferases. Briefly, we employed a query set of HATs from all kingdoms of life as proteins harboring known HAT domains and previously classified e.g. GCN5. A total of 293 HAT domain-containing proteins were identified from all kingdoms of life and their homologs were surveyed in the mouse proteome database. After removing redundant sequences and false positives, we obtained 33 putative HAT-like proteins in the mouse proteome. These 33 putative HAT-like proteins are encoded by 21 mouse genes indicating presence of isoforms for few of these proteins. Phylogenetic analysis of these 33 HATs revealed that there are groups of proteins from MYST family, GNAT family (GCN5/PCAF, ATAC2, ARD), P300/CBP family as well as a group of proteins which are not yet classified as HATs (Figure 1A). Most of the proteins in unclassified group are Camello proteins and sequence alignment of their acetyl-CoA binding domain with the GCN5 suggests that acetyl-CoA binding motif is conserved between them (Figure 1B). Further, to confirm that the Camello group of proteins also occur in other chordates, we performed similar analysis in zebrafish and identified 19 putative HAT proteins encoded by 18 genes suggesting their conservation in zebrafish genome (Additional file 1: Figure S1).
Complement of HAT homologs in mouse genome.
(A) The dendrogram was constructed using the UPGMA method of MEGA version 5.1 after 100 bootstrap resampling. Dendrogram of HATs in mouse classified into known and new family containing Camello proteins. Homologs of different HAT families are represented in different colors. The new family of HAT “Camello family” identified in this study is highlighted in grey. (B) Multiple sequence alignment of seven protein sequences belonging to Camello family with GCN5 shows presence of acetyl-coA-binding motif in all the members of Camello family, (C) General domain organization of GCN5 and Camello protein, CMLO3. HAT domain of GCN5 is tethered with additional domains whereas CMLO3 has just the HAT domain.
Next, we analyzed sequence features of the Camello proteins, which are smaller in size as compared to other HAT families. CMLO3, a representative example of one of the Camello proteins is 226 amino acids long and the HAT domain extends from 78 to 204 amino acids (Figure 1C). The subcellular localization of CMLO3 is predicted to be nuclear (with low confidence) and likely to have two transmembrane helices between 37–56th and 60–77th positions of sequence by PSORT and Phobius web-servers, respectively. Presence of two transmembrane helices suggests that CMLO3 is likely to be a nuclear membrane protein. This in turn suggests that C-terminal region of the CMLO3 containing HAT domain is nuclear and can still access nuclear histones for acetylation. Interestingly, the amino acid sequence between 61 to 84 (partially overlaps with the transmembrane helix) positions also shows similarity to the homeobox signature motif suggesting putative DNA-binding activity of the protein. Thus, our genome-wide bioinformatics analysis identified a novel family of HATs, which harbors the HAT domain but has no other associated domains.
Recombinant Camello proteins exhibit histone acetyltransferase activity specific for Histone H4
Next, to test the enzymatic activity of the Camello proteins identified in our bioinformatics analysis, we expressed recombinant full-length mouse Camello proteins (CMLO3, CMLO2 and E0CRY6) in bacterial cells and purified them to homogeneity (Additional file 2: Figure S2). HAT assays were performed using purified recombinant Camello proteins and histones H3 and H4 as substrates to study the functional status of the proteins. We found that Camello proteins (E0CYR6 and CMLO3) are active histone acetyltransferases with substrate preference for histone H4 in HAT assay (Figure 2A, lanes 2 and 3). However, CMLO2 did not exhibit any HAT activity under the conditions employed (Figure 2A, lane 4). There are two possibilities for the lack of acetylation of histone H4 by CMLO2, (i) it might act on other histones or non-histone proteins as a substrate, or (ii) it might require other partners or modifications of the substrate for its activity. Additionally, specific antibodies (H3ac, H4ac and H4K5ac, H4K8ac and H4K12ac) were used to cross-examine the acetylation levels on histone H3 and H4 by CMLO3 and E0CYR6 (Figure 2B). Further, to identify the residues modified by CMLO3 and E0CYR6, mass spectroscopic analysis was performed using histones H3 and H4 that were modified by CMLO3 and E0CYR6 in vitro. Both CMLO3 and E0CYR6 catalyzed acetylation of multiple lysines (H4K5, H4K8, H4K12 and H4K16) on Histone H4 (Figure 2C and 2D). No acetylated residues on histone H3 could be identified using similar strategy. Thus, these in vitro assays demonstrated that both CMLO3 and E0CYR6 are indeed active histone acetyltransferases with substrate preference for histone H4.
Camello family proteins CMLO3 and E0CYR6 are active histone acetyltransferases.
(A) HAT assay using baculo produced recombinant histones, H3 and H4. E0CYR6 and CMLO3 are showing histone H4 acetylation (lanes 2 and 3, respectively). (Full-length blot is provided as Additional file8: Figure S7) (B) Western blots using site-specific antibodies (H3ac, H4ac and H4K5, 8, 12ac). E0CYR6 and CMLO3 are specific towards histone H4 acetylation and have no activity on histone H3. All the samples were run on 15% SDS-polyacrylamide gels under similar conditions. (C) Mass spectrometric analysis of gel-cut histones, H3 and H4. Both CMLO3 and E0CYR6 were found to acetylate multiple lysines residues (H4K5, H4K8, H4K12 and H4K16) on Histone H4. First column indicates the mass spectral peptide. Second column indicates the position of the lysine residues on histone H4. Xcorr is the cross-correlation value, Xcorr above 2.0 are considered as good correlation. Charge is state of peptide. X in CMLO3 and E0CYR6 columns indicate the positive acetylation by these two proteins. (D) Complete mass spectrum of intensity vs. mass-to-charge ratio of the modified peptides.
Camello proteins CMLO3 and E0CYR6 exhibit perinuclear localization
Camello group of proteins in the pfam database are described as probable N-acetyltransferases (NAT) based on their sequence similarity to human Nat8 and Nat8l, which are reported to be aspartate acetyltransferases (AAT) specific to kidney and liver in human14,15. Furthermore, Nat8 and Nat8l are shown to be localized to the secretary pathways primarily in the endoplasmic reticulum and display lysine acetyltransferase activity16. To investigate the probable subcellular localization of Camello proteins we generated CMLO3-GFP and E0CYR6-GFP fusion constructs and expressed them in HeLa cells. Our results suggest that like human NAT8, CMLO3 and E0CYR6 are not localized in the endoplasmic reticulum (data shown for CMLO3, Figure 3A and Additional file 3: Figure S3). Interestingly, we found that CMLO3 and E0CYR6 primarily exhibit perinuclear localization (data shown for CMLO3, Figure 3B and 3C and Additional file 4: Figure S4). CMLO3-GFP fusion protein was also found in cytoplasm in cells exhibiting strong overexpression of CMLO3-GFP fusion protein. Moreover, localization of CMLO3 into nuclear compartment was also shown by Western blots after cytoplasmic and nuclear fractionation (Figure 3D). Furthermore, overexpression of CMLO3 and E0CYR6 in HeLa cells resulted in higher acetylation of endogenous histone H4 (Figure 3E, right panel lanes 2 and 3) suggesting the involvement of Camello proteins in acetylation of histone H4. To test whether the observed subcellular localization of CMLO3-GFP fusion protein is driven by bulky C-terminal GFP tag, we introduced his-myc tag separately at the C-terminal and N-terminal of CMLO3. We found that his-myc tagging of CMLO3 at C-terminal recapitulated the localization profile of the CMLO3-GFP fusion protein and its overlap with lamin B1, marker of the inner nuclear membrane and further strengthened our observation that CMLO3 is indeed localized in the nucleus (Additional file 5, Figure S5). However, N-terminal tagging of CMLO3 with his-myc tag affected the localization of CMLO3 and it was found mostly as punctuate structures (Additional file 5, Figure S5). Such localization pattern of CMLO3 could be presumably due to interference or hindrance of its N-terminal transmembrane helix by the his-myc tag. This in turn indicates that Camello proteins are indeed different than NAT8 protein as they have different functions and subcellular localization. Collectively, our analysis suggests that CMLO3 and E0CYR6 have perinuclear localization and their overexpression leads to higher acetylation of histone H4 in vivo.
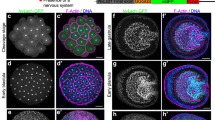
CMLO3 has perinuclear localization.
(A) CMLO3 is expressed as GFP fusion protein in HeLa cells. Co-localization of CMLO3-GFP fusion protein was compared with endoplasmic reticulum marker, calnexin. CMLO3 is not co-localized with endoplasmic reticulum marker. (B, C) CMLO3 has perinuclear localization and it co-localizes with the perinuclear membrane marker, Lamin B1. The line in the merged image denotes a linear fluorescence profile of co-localization. CMLO3-GFP fusion protein was also present in cytoplasm in cells having strong overexpression of CMLO3-GFP fusion protein. (D) Western blot analysis of CMLO3-GFP overexpressing HeLa cells after cytoplasmic and nuclear fractionation. Lamin B1 and tubulin were used as markers of nuclear and cytoplasmic fractions respectively. CMLO3-GFP was mainly observed in nuclear fraction. C - Cytoplasmic fraction; N- Nuclear fraction. (Full-length blot is provided as Additional file9: Figure S8) (E) Overexpression of CMLO3 and E0CYR6 leads to higher acetylation of endogenous histone H4. Tubulin was used as a loading control. Bar graph on the right represents the quantification of the relative protein intensity of the gel image by normalizing with tubulin. Overexpression of CMLO3 and E0CYR6 leads to 1.5- and 2.5-fold enrichment of endogenous hitsone H4 acetylation, respectively.
Camello family of HATs is conserved across chordates but originated in cnidaria
To study the origin, conservation and evolution of Camello–family of proteins we performed sequence homology search for CMLO3 along the length in 23 representative eukaryotic genomes (Additional file 6: Table S1). Out of the 450 homologous proteins detected by jackhammer in 23 genomes, only 46 proteins aligned along the length of CMLO3 protein (Additional file 7: Figure S6). Representative species were selected to construct multiple sequence alignment (MSA) for CMLO3 proteins. Figure 4A shows the overall conservation of several positions and the location of predicted transmembrane helices in MSA. The green boxes depict location of transmembrane helices predicted in the CMLO3 protein sequence (Fig. 4A). The transmembrane helices show less conservation except at a few positions. The location of acetyltransferase domain is shown in pink colored box. Acetyl co-A-binding motif is shown with black solid line suggesting possibility of catalytically active protein sequences. CMLO3 homologs are present in all chordates analyzed in this study including earliest chordate Branchiostomafloridae. The only non-chordate genome that is an exception to this trend is the cnidarian Nematostella, which encodes full length CMLO3 homolog. These results suggest the early origin of CMLO3 proteins in metazoan phylogeny. Furthermore, CMLO3 protein is absent in Drosophila melanogaster, Caenorhabditis elegans and Strongylocentrotus purpuratus. However, comparative genomic analysis suggests that Cnidarians are closely related to the chordates than that of Drosophila melanogaster and Caenorhabditis elegans. Previous studies also suggest heavy gene loss in these two organisms17,18. This in turn would suggest that CMLO3 might have emerged in Nematostella but subsequently lost in other invertebrates19,20.
Multiple sequence alignment and Neighbor-joining tree of 25 CMLO3 homologs.
(A) Proteins that showed significant similarity with CMLO3 across entire length were selected for MSA construction. Green and pink colored boxes depict location of predicted transmembrane helix and acetyltransferase domain in CMLO3 protein sequence, respectively. Color shading of MSA is based on clustalx. Positions are colored if they have more than 30% amino acids conserved. (B) Phylogenetic tree was constructed using neighbor-joining method for 221 positions. Statistical significance was assessed using 100 bootstraps. Two groups of CMLO3-like proteins clearly demarcated on the basis of Phylogenetic tree are shown in brown and navy blue color. Tree is rooted with ATS042_NEMVE sequence. Sequence identifiers can be used as identifiers for species as follows, HUMAN: Homo sapiens, MOUSE: Mus musculus, CHICK: Gallus gallus, XENTR: Xenopus tropicalis, DANRE: Danio rerio, BRAFL: Branchistoma floridae and NEMVE: Nematostella vectensis.
Phylogenetic tree of these 25 proteins clearly demarcate two groups of CMLO3 proteins which are likely to have diverged in gnathostomes from a single ancestral gene (Figure 4B). Bootstrap support for this split is 92. This suggests probable duplication of CMLO3 protein in the last common ancestor of Danio rerio and extant gnathostomes. The split between these two groups is evident using maximum-likelihood method with bootstrap support of 89. However, statistical support for other clades varies. Thus, CMLO3-like proteins seem to have first appeared in cnidarians with subsequent loss in few invertebrates and subsequently acquired by all chordates.
Morpholino mediated knockdown of CMLO3 in Zebrafish results in defective axis elongation and head formation
Our study suggests that Camello family of HATs is evolutionary conserved in all chordates and appeared for the first time in Cnidarians. The acquisition of Camello proteins coincides with the emergence of Cnidarians, suggesting that Camello proteins might have evolved for performing functions related to pattern formation during gastrulation. We therefore used zebrafish as a model system to study the in vivo functional relevance of CMLO3 protein in early embryonic development. To downregulate CMLO3 two antisense morpholinos were used which can target its mRNA sequence spanning the first methionine codon. Knockdown upon injection of 4 ng of control and two separate CMLO3 morpholinos led to defects in axis formation (Figure 5), which is typically seen in embryos with abnormalities in convergent extension20. CMLO3 morpholino injected embryos showed shortened body axis, deformed notochord and short trunk and tail at 18 hours post-fertilization (hpf). These observations suggest potential role of CMLO3 protein in convergent extension process in zebrafish. Further studies would help to understand the mechanisms by which CMLO3 is involved in this process.
Morpholino mediated knockdown of CMLO3 in zebrafish embryos.
(A) Lateral views of 18 hpf embryos injected with control MO, CMLO3 MO1 and CMLO3 MO2. (B) Stacked bar graph representing the percentage of embryos which are normal, dead or deformed after injecting control and CMLO3 morpholinos. Control (n = 220), CMLO3 MO1 (n = 180) and CMLO3 MO2 (n = 160) injected embryos were analyzed in this experiment, where n represents numbers of embryos injected.
Discussion
In this study, we performed classification of HATs into various families on the basis of protein sequences and domain organizations5,6,21. We could identify all the known families of HATs based on the presence of HAT domain in mouse and zebrafish genomes. Our genome-wide bioinformatics analysis also identified a group of HAT proteins namely Camello proteins and we wondered if Camello proteins represent a distinct novel family of HATs. Properties including absence of characteristic feature of the GNAT family and the p300/CBP family (bromo domain) and the MYST family (chromo domain), relatively small size of Camello proteins, absence of any other associated domain and their association with the nuclear membrane all indicate that Camello family might represent a novel family of HATs.
To our knowledge, Camello proteins are the only HATs exhibiting perinuclear localization. Our analysis also indicates that CMLO3 has the homeobox signature motif suggesting its DNA-binding activity. The nuclear pore-complex (NPC) in perinuclear membrane is considered a key regulator of the molecular trafficking between cytoplasm and nucleus. However, various experimental evidences suggest that the NPC also acts as bridge between nuclear transport and gene regulation by a phenomenon known as ‘gene-gating’22. The cross talks between chromatin remodeling, transcription and messenger-RNA export at NPC can act as a checkpoint for precise gene expression23. The transcription and export factor, Sus1 is one such example, which is a stable component of SAGA, the histone acetyltransferase complex and TREX-2, the nuclear pore associated transcription and export complex24. Sus1 functions in conjugation with Nucleoporins, Nup1 and Nup60 to recruit mRNA export machinery at the NPC, which are also the sites for physical tethering of the actively transcribed gene25. Based on above observations, we speculate that Camello proteins might mediate a dynamic attachment between transcriptionally active chromatin and the nuclear matrix and assist in gene regulation. Further, systematic investigation of the function of Camello proteins should reveal novel functions in the regulatory repertoire of the multi-faceted HAT complexes.
Camello family of HATs is evolutionary conserved in all chordates and appeared for the first time in Cnidarians. The acquisition of Camello proteins coincides with the emergence of Cnidarians, the early branching multicellular organisms with two germ layers, body axis and pattern formation19. Till date, CMLO3 is only studied in Xenopus levis, where overexpression of CMLO3 leads to inhibition of gastrulation26, suggesting that Camello proteins might have evolved for performing functions related to pattern formation during gastrulation. Interestingly, CMLO3 protein is absent in Drosophila melanogaster, Caenorhabditis elegans and Strongylocentrotus purpuratus presumably due to heavy gene loss in these organisms17,18. Thus, CMLO3 might have emerged in Nematostella but subsequently lost in other invertebrates and have functions associated with pattern formation.
The correlation between the histone acetylation and gene expression is crucial to determine the fate of embryo during embryonic development. Knockout of lysine actyltransferases p300, GCN5 and CBP is shown to exhibit embryonic lethality in mice5. p300 null mice exhibit defect in neurulation and heart development. CBP is required for neural tube closure and hematopoietic differentiation and GCN5 null mice exhibits defects in neural tube closure and development of mesodermal lineages5. This in turn suggests that independent HATs are required for specific gene expression program during the embryonic development. Similarly, knockdown of Camello protein (CMLO3) in zebrafish is embryonic lethal and exhibited defects in axis elongation and head formation, suggesting its critical role in pattern formation during early embryogenesis. This effect might be primarily because of abnormalities in convergent extension process in zebrafish embryos. Essential role of CMLO3 in zebrafish development could be attributed to the fact that many of the HATs are shown to exert a high degree of functional specificity during the early embryonic development, where each HAT plays independent and distinct role in stage-specific manner27.
Conclusion
In this study we have identified a novel family of HATs and demonstrated that it is specific for histone H4 acetylation. Our study is the first demonstration of any HAT in the perinuclear membrane. Camello family proteins are evolutionarily conserved in all the chrodates and originated from cnidarians. We also found that Camello proteins are essential for zebrafish development. Knockdown of CMLO3 in zebrafish leads to defects in axis elongation and head formation, suggesting its critical role in pattern formation during early embryogenesis. Focus of our future research would be to dissect the molecular mechanism associated with CMLO3 mediated acetylation and its effect on transcription, mRNA export and nuclear organization.
Methods
Protein sequence retrieval and bioinformatics analyses
The complete set of protein sequences from the ORFs of Mus musculus and Danio rerio have been obtained from UNIPROT database. The two genomes were then surveyed for putative histone acetyltransferases (HAT) against known HAT domains present in pfam database using sensitive sequence homology search algorithms such as BLAST28 and PSIBLAST29 with stringent e-value 0.0001. The hits identified were further pruned using the CD-HITS program30 to eliminate the redundant sequences which are 100% identical. Truncated sequences were further removed from the analysis. Hits which were lacking significant sequence similarity with the query were further examined manually and submitted to fold prediction method such as Phyre31 for the compatibility of sequence with 3D structural fold of HATs. The final data set of 33 putative mouse HATs were further analyzed for the associated domains which are identified using the HMMer32 against the Pfam database33.
Homologs of mouse CMLO3 protein and phylogenetic analysis
Homologs of CMLO3 protein of mouse were fished by searching against the database of protein sequences from 23 completely sequenced eukaryotic organisms (Additional file 6: Table S1). Details of homology search and phylogenetics analysis are provided in Supplementary Information.
Cloning of CMLO genes and protein expression
Full-length CMLO genes were PCR amplified from mouse embryo cDNA library (E8.5) using gene-specific primers. Amplified PCR products were cloned in pET28b (Novagen) in NdeI and XhoI restriction sites. The clones thus obtained were confirmed by DNA sequencing. CMLO3 proteins were expressed in E. coli (BL21) as 6× histidine tagged fusion proteins by induction with 0.4 mM isopropyl–B-D thiogalactopyrranoside (IPTG) for 6 h at 25°C. Proteins were affinity purified using Ni-NTA beads (Qiagen). CMLO3-GFP and E0CYR6-GFP fusion constructs were prepared by cloning CMLO3 and E0CYR6 in pEGFP-N1 plasmid (Clontech).
C-and N-terminal His-Myc tagging of CMLO3
C-terminal tagging of CMLO3 was performed by digesting pET28b-CMLO3 plasmid with XbaI and XhoI restriction enzymes. Resultant insert was ligated with NheI and XhoI digested pcDNA3.1(−)/myc-His A (Addgene) plasmid. N-terminal Myc-His tag CMLO3 was produced by cloning of His-CMLO3 between SalI and AflII restriction sites of pcDNA3.1(−)/myc-His A.
Histone acetyltransferase assays
Camello protein (100 ng) was incubated for 30 min at 30°C in HAT buffer (20 mM Tris-HCl, pH 8.8, 1.5 mM MgCl2, 10 mM NaCl and acetyl-CoA with 400 ng recombinant histones H3 and H4. Reactions were run on 15% SDS-PAGE and Western blot was performed using Pan-lysine acetyl (06-933, Upstate), H3ac (Ab47915), H4ac (06-866, Millipore) antibodies. Mass spectroscopic analysis of the proteins after HAT assays was performed at the W.M. Keck Biomedical Mass Spectrometry Laboratory, University of Virginia Health System, USA.
Cell culture, transfection, immunostaining and microscopy
HeLa cells were grown in DMEM medium supplemented with 10% fetal bovine serum at 37°C with 5% CO2. Details of transfection, immunostaining and microscopy are provided in Supplementary Information.
Morpholino (MO) injections in Zebrafish
Tubingen (Tu) strain of zebrafish was used for morpholino injections. Control morpholino and CMLO3 morpholino were designed and obtained from Gene tools LLC. Four ng of 5 base mismatch control MO (5′-TCCCCTCCATGCACATAACACGAGA-3′) and CMLO3 MO1 (5′ –GACGAATCTGCACCTCATCCATGAC-3′) and CMLO3 MO2 (5′-TCGCCTCGATCCAGATAAGACGAGA-3′) were injected at one cell stage in WT zebrafish embryos and phenotypes were scored at 18 hpf. For zebrafish maintenance and experimentation, the guidelines recommended by the Committee for the Purpose of Control and Supervision of Experiments on Animals (CPCSEA), Government of India, were followed. Zebrafish experiments were also approved by the Tata Institute of Fundamental Research (TIFR), India.
Ethics statements
Zebrafish experiments were performed in accordance with the guidelines of Committee for the Purpose of Control and Supervision of Experiments on Animals (CPCSEA), Government of India. All animal procedures carried out in this study were reviewed, approved and supervised by the Institutional Ethics Committee of Tata Institute of Fundamental Research (TIFR), India.
References
Kouzarides, T. Chromatin modifications and their function. Cell. 128, 693–705 (2007).
Zentner, G. E. & Henikoff, S. Regulation of nucleosome dynamics by histone modifications. Nat Struct Mol Biol. 20, 259–266 (2013).
Talbert, P. B. et al. A unified phylogeny-based nomenclature for histone variants. Epigenetics Chromatin. 21, 7 (2012).
Sterner, D. E. & Berger, S. L. Acetylation of Histones and Transcription-Related Factors. Microbiol Mol Biol Rev. 64, 435 (2000).
Roth, S. Y., Denu, J. M. & Allis, C. D. Histone acetyltransferases. Annu Rev Biochem. 70, 81–120 (2000).
Yang, X. J. The diverse superfamily of lysine acetyltransferases and their roles in leukemia and other diseases. Nucleic Acids Res. 32, 959–76 (2004).
Pandey, R. et al. Analysis of histone acetyltransferase and histone deacetylase families of Arabidopsis thaliana suggests functional diversification of chromatin modification among multicellular eukaryotes. Nucleic Acids Res. 30, 5036–5055 (2002).
Roth, S. Y. Chromatin-mediated transcriptional repression in yeast. Curr Opin Genet Dev. 5, 168–73 (1995).
Giles, R. H., Peters, D. J. & Breuning, M. H. Conjunction dysfunction: CBP/p300 in human disease. Trends Genet. 14, 178–183 (1998).
Mizzen, C. A. et al. The TAF(II)250 subunit of TFIID has histone acetyltransferase activity. Cell. 87, 1261–1270 (1996).
Chen, H. et al. Nuclear receptor coactivator ACTR is a novel histone acetyltransferase and forms a multimeric activation complex with P/CAF and CBP/p300. Cell 90, 569–580 (1997).
Doi, M., Hirayama, J. & Sassone-Corsi, P. Circadian regulator CLOCK is a histone acetyltransferase. Cell. 125, 497–508 (2006).
Leo, C. & Chen, J. D. The SRC family of nuclear receptor coactivators. Gene. 245, 1–11 (2000).
Ariyannur, P. S. Methamphetamine-induced neuronal protein NAT8L is the NAA biosynthetic enzyme: implications for specialized acetyl coenzyme A metabolism in the CNS. Brain Res. 1335, 1–13 (2010).
Veiga-da-Cunha, M. et al. Molecular identification of NAT8 as the enzyme that acetylates cysteine S-conjugates to mercapturic acids. J Biol Chem. 285, 18888–98 (2010).
Ko, M. H. & Puglielli, L. Two endoplasmic reticulum (ER)/ER Golgi intermediate compartment-based lysine acetyltransferases post-translationally regulate BACE1 levels. J Biol Chem. 284, 2482–2492 (2009).
Cutter, A. D., Dey, A. & Murray, R. L. Evolution of the Caenorhabditis elegans genome. Mol Biol Evol. 26, 1199–234 (2009).
Ranz, J. M. et al. Principles of genome evolution in the Drosophila melanogaster species group. PLoS Biol. 5, e152 (2007).
Technau, U. & Steele, R. E. Evolutionary crossroads in developmental biology: Cnidaria. Development. 138, 1447–58 (2011).
Jessen, J. R. et al. Zebrafish trilobite identifies new roles for Strabismus in gastrulation and neuronal movements. Nat Cell Biol. 4, 610–615 (2002).
Carrozza, M. J., Utley, R. T., Workman, J. L. & Côté, J. The diverse functions of histone acetyltransferase complexes. Trends Genet. 9, 321–29 (2003).
Strambio-De-Castillia, C., Niepel, M. & Routm, M. P. The nuclear pore complex: bridging nuclear transport and gene regulation. Nat Rev Mol Cell Biol. 11, 490–501 (2010).
Rodríguez-Navarro, S. & Hurt, E. Linking gene regulation to mRNA production and export. Curr Opin Cell Biol. 23, 302–09 (2011).
Pascual-García, P. et al. Sus1 is recruited to coding regions and functions during transcription elongation in association with SAGA and TREX2. Genes Dev. 22, 2811–22 (2008).
Rodríguez-Navarro, S. et al. Sus1, a functional component of the SAGA histone acetylase complex and the nuclear pore-associated mRNA export machinery. Cell. 116, 75–86 (2004).
Popsueva, A. E. et al. Overexpression of camello, a member of a novel protein family, reduces blastomere adhesion and inhibits gastrulation in Xenopus laevis. Dev Biol. 234, 483–96 (2001).
Anamika, K. et al. Lessons from genome-wide studies: an integrated definition of the coactivator function of histone acetyl transferases. Epigenetics Chromatin. 3, 3–18 (2010).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J Mol Biol. 215, 403–410 (2010).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–02 (1997).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics. 22, 1658–59 (2006).
Kelley, L. A. & Sternberg, M. J. E. Protein structure prediction on the web: a case study using the Phyre server. Nature Protocols. 4, 363–71 (2009).
Eddy, S. R. Profile hidden Markov models. Bioinformatics. 14, 755–63 (1998).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 30, 276–280 (2002).
Acknowledgements
Work was supported by grants from the Centre of Excellence in Epigenetics program of the Department of Biotechnology, Government of India and IISER Pune. We thank Prof. Laszlo Tora for valuable suggestions, R. L. Praveena and Vijay Vithal for help with confocal microscopy and Ashwin K. and Gayathri P. for critical reading of the manuscript. We also thank Dr. Mahendra Sonawane for providing access to zebrafish facility.
Author information
Authors and Affiliations
Contributions
K.A. conceived and performed classification of HATs and prepared Figure 1. Evolutionary analysis is performed by V.Y.M. and prepared Figure 4. Zebrafish experiments were performed by S.J.P. and Y.B. and prepared Figure 5. All other experiments were performed by K.K. and prepared Figures 2–3. K.K. and S.G. analyzed data. K.K. and S.G. wrote the manuscript, planned, coordinated and supervised the project.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Karmodiya, K., Anamika, K., Muley, V. et al. Camello, a novel family of Histone Acetyltransferases that acetylate histone H4 and is essential for zebrafish development. Sci Rep 4, 6076 (2014). https://doi.org/10.1038/srep06076
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06076
This article is cited by
-
Spatiotemporal expression of Prdm genes during Xenopus development
Cytotechnology (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.