Abstract
Nitrate transporters are an important component of plant growth and development. Chrysanthemum morifolium is an important ornamental species, for which a sufficient supply of nitrogenous fertilizer is required to maintain economic yields. In this study, the full-length cDNA of the nitrate transporter genes CmNRT2 and CmNAR2 were isolated. CmNRT2 transcript accumulation was inducible by both nitrate and ammonium, but the latter ion down-regulated the transcript accumulation of CmNAR2. CmNRT2 might be a plasma membrane localized protein, while CmNAR2 was distributed throughout the cell. CmNAR2 was shown to interact with CmNRT2 by in vitro and in vivo assays. Arabidopsis thaliana plants heterologously expressing CmNRT2 showed an increased rate of nitrate influx, while this trait was unaltered in plants expressing CmNAR2. Double transformants (CmNRT2 plus CmNAR2) exhibited an enhanced rate of nitrate influx into the root. Our data indicated that the interaction of CmNAR2 with CmNRT2 contributed to the uptake of nitrate.
Similar content being viewed by others
Introduction
In soil, inorganic nitrogen (N) is available for plants as nitrate and ammonium1 and plants have evolved various means of coping with nitrogen deficiency2. Genes belonging to the nitrate transporter families (NRT1, NRT2) and the nitrate assimilation related family (NAR2) ensure the capacity of root cells to regulate the uptake of nitrate, through two high-affinity transport systems (HATS) and one low-affinity transport system (LATS) depending on nitrate concentrations3,4. The Arabidopsis thaliana (A. thaliana) gene AtNPF6.3 has been shown to encode a dual-affinity nitrate transporter5. Certain elements of HATS are inducible (iHATS) while others are constitutively expressed6,7. Full activity of iHATS in the A. thaliana root requires the presence of both AtNRT2.1 and AtNRT2.28. Nevertheless, it was shown that the expression of high affinity nitrate transport in Chlamydomonas sp. requires the presence of NRT2.1, NRT2.2 and NAR29, an observation confirmed in experiments based on a Xenopus oocyte expression system10. In T-DNA insertional mutant atnar2.1-1 and atnar2.1-2, constitutive HATS (cHATS) was reduced by 34% and 89%, while iHATS was reduced by 92% and 96% respectively11. In the atnrt2.1-1 mutant, the HATS under either high or low nitrate growth concentrations was affected12. AtNAR2.1 and AtNRT2.1 interact with one another during the HATS process13, while the product of AtNAR2 is involved in the constitutively induced high affinity influx14. Similarly in rice, OsNAR2.1 interacts with OsNRT2.1, 2.2 and 2.3a15. In A. thaliana, the AtNRT2 transporters interact strongly with AtNAR2.1 except for AtNRT2.78.
Chrysanthemum (Chrysanthemum morifolium) is a commercially important ornamental species, for which a sufficient supply of nitrogenous fertilizer is required to promote vegetative growth16,17. However, excessive fertilizer application, especially when coupled with a low nitrate utilization rate, risks a significant wastage of nutrients, which can lead to problems of environmental pollution. Thus studying the HATS and LATS in chrysanthemum is necessary for cultivation. As yet, no genes involved in nitrate uptake have been isolated in chrysanthemum, nor have any interactions between nitrate transporters and nitrate assimilation related proteins been defined. Here, the isolation of CmNRT2 and CmNAR2 cDNA was described and their inducibility by nitrate and ammonium was described. A series of assays was used to show that their gene products interact with one another. The behavior of transgenic A. thaliana confirmed that CmNAR2 probably serves as a partner to CmNRT2 in controlling nitrate uptake.
Results
The CmNRT2 and CmNAR2 coding and promoter sequences
The degenerate PCR amplified a 959 bp fragment of CmNRT2 and a 300 bp fragment of CmNAR2. Extension from these amplicons to the full length cDNAs via RACE PCR identified that the two genes had an ORF of length, respectively, 1590 bp (DDBJ accession AB921547) and 597 bp (DDBJ accession AB921548), implying polypeptide products composed of 530 (molecular weight 57.6 kDa) and 199 (22.3 kDa) residues. The alignment of the CmNRT2 sequence with those of homologous NRT2 proteins showed levels of similarity ranging between 39.78% and 79.81% (Table S1), while that of CmNAR2 was between 39.13% and 54.50% (Table S2, Fig. S3). Only three conserved motifs LCYAP, WRK and YGQ were included in CmNAR2 with its related NAR2 sequences (Fig. S4). On the other hand, a comparison with a set of full-length NRT2 and NAR2 proteins from other species clustered CmNRT2 with NpNRT2.1 and CmNAR2 with CsNAR2 in the evolutionary, respectively (Fig. 1). The 1413 bp (CmNRT2) and 1683 bp (CmNAR2) sequences upstream of the translation initiation site were obtained by TAIL-PCR and anchored PCR. By reference to the PLACE database, the CmNRT2 promoter included a number of nitrate-responsive cis elements including 5 A(C/G)TCA and 22 GAGABOX motifs (Fig. S5), while the CmNAR2.1 promoter harbored 5 A(C/G)TCA motifs and 18 GAGABOX motifs, along with a single nitrogen-responsive EMHVCHORD motif (Fig. S6).
Phylogeny of NRT2 and NAR2 polypeptides.
Bootstrap percentages are given for each branch and the scale bar represents 0.05 substitutions per site. (A) The relationship of CmNRT2 with related NRT2s. The genes encoding these products and their GenBank accession numbers are: A. thalianaAtNRT2.1 (AAC64170), AtNRT2.2 (AAC35884), AtNRT2.3 (BAB10099), AtNRT2.4 (BAB10098), AtNRT2.5 (AAF78499), AtNRT2.6 (CAB89321) and AtNRT2.7 (CAB87624), Nicotiana. plumbaginifoliaNpNRT2.1 (CAA69387), Lotus japonicusLjNRT2 (CAC35729), soybean GmNRT2 (AAC09320), rice OsNRT2 (BAA33382), barley HvNRT2.1 (AAC49531), wheat TaNRT2.1 (AAK19519), Chlamydomonas reinhardtiiCrNRT2.1 (CAA80925), Medicago truncatulaMtNRT2 (XP_003596775). chrysanthemum CmNRT2 (DDBJ accession AB921547) is shown boxed. (B) The relationship of CmNAR2 with its related NAR2 sequences. The genes encoding these products and their GenBank accession numbers are: A. thaliana AtNAR2.1 (NP_199831), AtNAR2.2 (CAC36942), Cucumis sativus CsNAR2.1 (ACV33078), barley HvNAR2.1, HvNAR2.2 and HvNAR2.3 (AAP31850, AAP31851, AAP31852), L. japonicusLjNAR2.1 (BAI63584), rice OsNAR2.1 (AP004023.2), wheat TaNAR2.1 (AAV35210), maize ZmNAR2.1 (NP_001105929). chrysanthemum CmNAR2 (DDBJ accession AB921548) is shown boxed.
Transcript accumulation of CmNRT2 and CmNAR2 is inducible
To gain the expression pattern of CmNRT2 and CmNAR2 under various external NO3− and NH4+ concentrations, qRT-PCR was carried out to analyze samples. In the presence of 0.5 mM nitrate following a period of nitrogen starvation, CmNRT2 transcript abundance increased gradually over the first 4 h, but decreased marginally thereafter (Fig. 2A). Its behavior was similar at the two higher nitrate concentrations (Fig. 2C, E). CmNAR2 transcript accumulation was also induced, peaking 6 h after the initial provision of each of the three different concentrations of nitrate, then declining gradually (Fig. 2B, D, F). CmNRT2 transcript abundance was substantially raised after a 4 h exposure to 5 mM ammonium, but that of CmNAR2 was decreased (Fig. 2H).
The induced gene expression level of (A, C, E, G) CmNRT2 and (B, D, F, H) CmNAR2 in roots exposed, following a period of N starvation, to (A, B) 0.5 mM nitrate, (C, D) 1 mM nitrate, (E, F) 5 mM nitrate, (G, H) 5 mM ammonium, as assayed by qRT-PCR.
Mean values and standard deviation calculated from triplicated assays.
Subcellular localization of CmNRT2 and CmNAR2
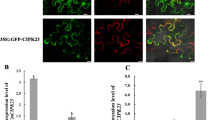
To explore subcellular localization of CmNRT2 and CmNAR2, the transient assays of GFP fusion constructs transformed into onion epidermal cells by particle bombardment were used. As shown in Fig. 3, the transient expression of CmNRT2::GFP and CmNAR2::GFP fusions in onion epidermal cells suggested that CmNAR2 localized throughout the entire cell, whereas CmNRT2 might localize to the plasma membrane, however a conclusion can't be drawn at present because of lack of co-localization with PM makers. In contrast, 35S-GFP alone resulted in diffused distribution of green fluorescence throughout the cell (Fig. 3).
Subcellular localization of CmNRT2 and CmNAR2.
Inserted plasmid carry GFP, CmNAR2::GFP; CmNRT2::GFP. Images DIC taken under bright light to display the morphology of the cell and images GFP under dark field to show GFP fluorescence; images Merge are formed by merging GFP and DIC, respectively. Bars: 50 μm.
Interaction between CmNRT2 and CmNAR2
To test whether CmNRT2 interacted with CmNAR2, the yeast two hybrid technique was used in our paper. CmNRT2 protein was fused to the mutated N-terminal half of ubiquitin (NubG) and the CmNAR2 protein was fused to the C-terminal half of ubiquitin (Cub) and the transcription factor LexA-VP1615. When NubG and Cub are brought together as fusions by CmNRT2 and CmNAR2 interacted, split-ubiquitin is restored. Then two types of reporter genes are activated by VP16 which allows growth tests on selective media (HIS3 and ADE2) and β-galactosidase activity assays (LacZ). Our datas showed that CmNAR2 interacted with CmNRT2 (Fig. 4A, B). Confirmation was obtained from X-α-Gal detection (Fig. 4C). No activity was detectable when either CmNAR2/pPR3-N or CmNRT2/pBT3-C were co-transformed. In addition, the BiFC technique was also used to characterize protein interactions invivo. The pSAT4A-CmNAR2 vector contains a fusion of CmNAR2 at the C terminus of YFP and the pSAT4A-CmNRT2 vector contains a fusion of CmNRT2 at the N terminus of YFP. The two recombinant DNA constructs were introduced into onion epidermal cells by particle bombardment. Signals were detected in the onion epidermal cell membrane when the cells were transformed with CmNRT2-YFPN/CmNAR2-YFPC, but were not observed when coexpressed with CmNRT2 -YFPN/YFPC, or YFPN/CmNAR2-YFPC, (Fig. 5).
The interaction between CmNRT2 and CmNAR2 based on the DUAL membrane pairwise interaction assay.
Cells grown on media lacking (A) Ade, His, Leu and Trp, (B) Leu and Trp. (C) β-galactosidase assay of the interaction between CmNRT2 and CmNAR2. Yeast strain NMY51 carried each pairs of bait and prey plasmid (pBT3-C and pPR3-N are the control vectors with no cloned cDNA). The construct pair of CmNRT2 and CmNAR2 fusions was used as positive control. The construct pairs of CmNRT2 and CmNAR2 with empty vector, respectively, were used as negative controls.
BiFC-based analysis of the CmNRT2 and CmNAR2 interaction in transiently transformed onion epidermal cells.
YFP fluorescence, DIC and merged images are shown for each transformation combination. The construct pair of CmNRT2-nYFP and CmNAR2-cYFP fusions was used as positive control. The construct pairs of CmNRT2-nYFP and CmNAR2-cYFP with empty vector, respectively, were used as negative controls. Bars: 50 μm.
Nitrate uptake in Xenopus oocytes
To assay the amount of nitrate accumulated inside the oocytes, a colorimetric method was used in the paper18. CmNRT2 and CmNAR2 were subcloned in to expression vector pT7Ts and then used as template to synthesize mRNA. After incubation in MBS solution including 0.5 mM NaNO3 for 16 h, there was no significant difference in nitrate uptake between the single injection (CmNRT2 or CmNAR2) and water injected oocytes. However the co-injection of CmNRT2 and CmNAR2 had a substantial positive increase on nitrate uptake (Fig. 6).
Nitrate uptake in Xenopus oocytes injected with the single gene assays or CmNRT2 and CmNAR2.
Water injected oocytes were as a control. Oocytes were incubated in MBS with 0.5 mM NaNO3 for 16 h and washed four times with NO3− free MBS solution. Four oocytes were pooled for each sample. Values were average of n = 8 ± SD. Asterisks signify that co-injection of NRT2 and NAR2 produced significantly higher nitrate uptake (by the t-test at P < 0.01) than others.
Root 15N influx in transgenic A. thaliana lines
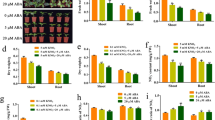
To confirm the function by in planta, we used different transgenic construction expressed in A. thaliana to measure root 15N influx. Through Agrobacterium-mediated transformation, four single-gene transformed lines (RT-3 and RT-20 carrying p1301-220-CmNRT2 and AR-1 and AR-2 carrying p1301-220-CmNAR2) and two dual-gene transformed lines (D1 and D2, carrying p1301-220-CmNAR2-CmNRT2) were arbitrarily chosen for study. Based on qRT-PCR assay, target genes CmNRT2 and CmNAR2 were expressed in transgenic A. thaliana lines but not wild-type (WT) and CaMV 35S::GUS (vector) A. thaliana plants (Fig. 7 A, B). After 5 min soaked in nutrient solution containing 0.2 mM 15N-NO3−, lines RT-3, RT-20, D1 and D2 showed a significantly enhanced rate of nitrate influx than wild-type and empty vector control transgenic lines (Fig. 8), indicating that the presence of CmNRT2 was responsible for heightened nitrate uptake. The 15N influx in the double transgenic gene lines was much greater than in the single gene ones indicatinig that CmNAR2 can interact with CmNRT2 to promote root 15N influx. There was no significant difference in nitrate uptake performance between the CmNAR2 transformants and the non-transformed control (Fig. 8). After 5 min culture in nutrient solution containing 6 mM 15N-NO3−, the data showed that no significant difference in root nitrate influx was found in transgenic and non-transgenic A. thaliana lines (Fig. S7).
Detection of CmNRT2 and CmNAR2 in the T3 generation of transgenic A. thaliana lines.
(A): The CmNRT2 expression based on qRT-PCR. (B): The CmNAR2 expression based on qRT-PCR. WT: wild type A. thaliana; Vector: empty vector control transgenic line; RT-3 and RT-20: CmNRT2 transgenics, AR-1 and AR-2: CmNAR2 transgenics; D1, D2: CmNRT2/CmNAR2 transgenics; AtUBQ: A. thaliana. reference gene. Transcript abundance was normalized against the expression of the constitutively expressed AtUBQ. The data efficiency of each reaction was 2−ΔΔCT (Livak and Schmittgen, 2001). Error bars represent SD of three biological replicates.
Influx of 15N influx into the root of transgenic and non-transgenic A. thaliana.
A. thaliana plants were grown for 42 d in hydroponics on 0.2 mM NO3−. Root 15NO3− was measured after 5 min labelling with complete nutrient solution containing 0.2 mM 15NO3−. WT: wild type A. thaliana; Vector: empty vector control transgenic line; RT-3 and RT-20: CmNRT2 transgenics, AR-1 and AR-2: CmNAR2 transgenics; D1, D2: CmNRT2/CmNAR2 transgenics. Values represent the mean ± SE (n = 5). Asterisks indicate means which differ significantly (by the t-test at P < 0.01) from the wild type value.
Discussion
Genes encoding NRT2 or NAR2 proteins are present in many plant species2,10,19,20,21. According to degenerated primers, seven chrysanthemum NRT2 homolog CmNRT2 and one NAR2 homolog CmNAR2 were isolated by RT-PCR and RACE methods (data not shown). Here, we focus on the molecular characterization of one CmNRT2 gene and CmNAR2. The CmNRT2 sequence proved to be highly homologous with those of other NRT2 proteins, but conservation of the CmNAR2 sequence was less marked, although the LCYAP, WRK and YGQ motifs were still retained. The regulation of these genes is believed to occur at transcriptional levels22,23 and the motif content of the promoter sequences of the two genes was consistent with their both being involved in nitrate uptake and/or signaling. Like other nitrate-inducible genes such as OsNRT2.1–2.4 in rice24, TaNRT2.1–2.3 in wheat25, AtNRT2.1 in A. thaliana26 and NpNRT2.1 in Nicotiana plumbaginifolia27, the expression level of CmNRT2 in the root was strongly affected by the nitrogen status of the growing medium (Fig. 2). The members of the AtNRT2 gene family are variously nitrate-inducible (AtNRT2.1, 2.2 and 2.4), nitrate-repressible (2.5) and constitutively expressed (2.3, 2.6 and 2.7)26. The transcript accumulation of CmNRT2 mirrored that of AtNRT2.4 in showing a nitrate-inducible peak, which decayed with time. CmNAR2 transcript accumulation followed a similar temporal profile, suggesting that these two chrysanthemum genes act in concert during nitrate uptake (Fig. 2). The transcript accumulation of CmNRT2, but not that of CmNAR2, was inducible by ammonium. In rice, it has been shown that the transcript accumulation in the root of OsNAR2.1 is suppressed by the presence of 5 mM ammonium, while that of various OsNRT2 genes (2.1, 2.2, 2.3a and 2.3b) is inhibited by much lower levels23. Ammonium also has an inhibitory effect on the transcript accumulation of the Dunaliella salina gene DsNRT2.128. Thus, the induction/inhibition of NRT2 transcript accumulation by nitrate or ammonium is quite gene and species dependent.
A number of techniques have been developed to reveal gene interactions, some involving in vitro assays and others carried out in vivo11,13,15,29. Here, use of the in vitro based yeast two-hybrid assay was able to establish that CmNAR2 interacts with CmNRT2 (Fig. 4) and this result was confirmed using the in vivo BiFC test (Fig. 5). The latter showed that the interaction took place in the plasma membrane, which again suggested a possible plasma membrane localization of CmNRT2 (Fig. 3). AtNRT2.1 localizes mostly to the plasma membrane of root cortical and epidermal cells and the BiFC assay has shown that an intact AtNRT2.1/AtNAR2.1 complex is formed within the A. thaliana plasma membrane29. CmNAR2 localized throughout the cell (Fig. 3). The natural hypothesis is therefore that CmNRT2 requires the presence of CmNAR2 to form a transporter unit. The effect on the rate of nitrate influx of constitutively expressing both chrysanthemum sequences as transgenes in A. thaliana simultaneously was much more positive than when either CmNAR2 or CmNRT2 on their own were expressed, underlining the importance of the CmNAR2/CmNRT2 interaction. In A. thaliana, AtNAR2.1 has been identified as chaperonin proteins and AtNRT2.1–2.6 all require the presence of AtNAR2.1 to be effective (this dos not apply to AtNRT2.7)8. In rice, the alteration in the influx of nitrate observed in an osnar2.1 knockdown mutant was taken to show that both HATS and LATS were impaired15. However, LATS was not affected using the different transgenic constructions in our results, which may be that CmNRT2 or CmNAR2 was responsible in HATS and CmNRT2 interacted with CmNAR2 at the protein level in HATS. Our work has further helped to confirm the proposed complementary roles of NRT2 and NAR2 proteins.
Methods
Plant materials and growing conditions
The chrysanthemum variety ‘Nannongxuefeng’ was obtained from the Chrysanthemum Germplasm Resource Preserving Centre, Nanjing Agricultural University, China. Cuttings were induced to form roots by a two week exposure to a liquid culture 1/4 MS. Phenotypically uniform seedlings at the 8 leaf stage were transferred into 8 L of 5 mM NH4NO3, 2.5 mM K2SO4, 1.5 mM MgSO4.7H2O, 1.33 mM NaH2PO4.2H2O, 2.0 mM CaCl2, 20 μM H3BO3, 9 μM MnCl2.4H2O, 0.77 μM ZnSO4·7H2O, 0.32 μM CuSO4·5H2O, 0.39 μM Na2MoO4.2H2O, 20 μM FeNaEDTA, pH 6.5. To inhibit nitrification, 7 μM dicyandiamide was added to the solution, which was refreshed each two days.
Isolation and sequencing of full-length cDNA
Batches of cuttings were grown in the above nutrient solution for four weeks and then were transferred into a nitrogen-free solution for one week. Total RNA was isolated from the roots 1 h after the addition of 5 mM KNO3, using the RNAiso reagent (TaKaRa, Japan) following the manufacturer's instructions and treating with RNase-free DNaseI (TaKaRa, Japan) to remove any DNA contamination. The concentration and the integrity of the resulting RNA were assessed following Gu et al30. The first cDNA strand was synthesized using Reverse Transcriptase M-MLV (RNase H−) (TaKaRa, Japan) according to the manufacturer's instructions. Two pairs of degenerate primers CmNRT2FD/RH and CmNAR2FD/RH (sequences given in Table 1) were designed based on conserved regions of, respectively, NRT2 (Fig. S1) and NAR2 (Fig. S2) homologs from a range of plant species. The remainder of the sequence was acquired using RACE-PCR following Shan et al.31. The open reading frame (ORF) within the amplicons was identified using ORF finder software (www.ncbi.nlm.nih.gov) and its deduced peptide sequence was subjected to a BLASTp search to identify homologs. The phylogeny of the sequences was derived using the DNAman software package.
Isolation and sequence analysis of promoters
Genomic DNA was isolated from leaves using the CTAB method32 and diluted to 50 ng/μL. The CmNAR2 and CmNRT2 promoter sequences were isolated from this genomic DNA, via three rounds of PCR (primer sequences given in Table 1), where each round used the product of the previous one as template, applying both TAIL-PCR and anchored PCR33,34. Fragments of CmNAR2 and CmNRT2 upstream of the translation initiation codon were isolated using the latter procedure, while the former used these sequences as the template. The TAIL-PCR primer pairs were P1, P2 and P3 in combination with AC1 and those for the anchored PCR were R1, R2 and R3 in combination with AAP and AUAP. The resulting amplicons were sequenced. Analyses of the sequences upstream of each gene relied on RSTA (rsat.ulb.ac.be/rsat/) software and the databases PLACE (dna.affrc.go.jp/PLACE/signalscan. html) and softberry (www.softberry.com/berry.phtml)23.
Real-time RT-PCR
Cuttings were grown in nutrient solution as above for four weeks and then starved of nitrogen for one week. Total RNA was isolated from the root tissue of plants supplied with either 0.5 mM, 1 mM or 5 mM KNO3 after either 0, 0.5, 1, 2, 4, 6, 8 or 12 h, or of those supplied with 5 mM NH4Cl after either 0, 4 or 6 h. The expression profiles of CmNAR2 and CmNRT2 were obtained by real time RT-PCR based on SYBR® Premix Ex Taq™ II (Perfect Real Time) (TAKARA, Japan). The PCR comprised an initial denaturation step of 95°C/60 s, followed by 40 cycles of 95°C/15 s, 55°C/30 s and 72°C/30 s. At the end of the PCR, the temperature was raised from 55 to 95°C at a rate of 0.5°C s−1 to generate the required denaturation curve. The primer pair qCmNRT2-F/qCmNRT2-R and qCmNAR2-F/qCmNAR2-R were applied to amplify CmNRT2 or CmNAR2 fragment, respectively (Table 1). The reference gene used was a fragment of CmpsaA (AB548817)30 with primer pair psaA-F/psaA-R (Table 1). Each PCR was repeated as three biological replicates and relative changes in transcript accumulation were analysed using the 2−ΔΔCT approach35.
Subcellular localization of CmNAR2 and CmNRT2
Plasmids for transient expression were generated using the Invitrogen Gateway system. The GUS sequence (without the stop codon) from pBI121, CmNAR2 and CmNRT2 ORFs (lacking their stop codon) were amplified employing primer pairs GUS-BamH I-103/GUS-Xho I-103, NAR2-BamH I-103/NAR2-Not I-103 and NRT2-Sal I-103/NRT2-Not I-103 (Table 1), then they were sub-cloned into pMD19-T (TAKARA, Japan) and confirmed by DNA sequencing. The pMD19-T constructs and the pENTRTM1A dual selection vector (Invitrogen) were cut by BamH I/Xho I (GUS), BamH I/Not I (CmNAR2) and Sal I/Not I (CmNRT2), ligated using T4 DNA ligase (TAKARA, Japan) and finally introduced into the pEarleyGate 103 plasmid using an LR Clonase™ II enzyme mix (Invitrogen) to produce pEarleyGate 103-GFP, pEarleyGate 103-CmNAR2/CmNRT2 vector respectively36. The transgenes were transiently expressed in onion epidermal cells using a He-driven PDS-1000 particle accelerator (Bio-Rad). pEarleyGate 103-GFP was used as a control. After bombardment, the onion peels were kept for 24 h in the dark on Murashige and Skoog (1962) medium plates. Confocal laser microscopy was used to monitor the expression of GFP.
Protein–protein interaction assays
The interactions between CmNAR2 and CmNRT2 were tested using a DUAL membrane pairwise interaction kit (Dualsystems Biotech, Schlieren, Switzerland). Full-length of CmNAR2 and CmNRT2 cDNAs were amplified using PrimeSTAR® HS DNA polymerase (TAKARA, Japan) with the primer pairs pBT3-C-NAR2-JS/pBT3-C-NAR2-JX and pPR3-N-2-JS/pPR3-N-2-JX (Table 1), then cloned into pBT3-C (LEU2, KanR) and pPR3-N (TRP1, AmpR) expression vectors, respectively and co-transformed into yeast strain NMY51 (MATa his3 trp1 leu2 ade2 LYS2::HIS3 ura3::lacZ ade2::ADE2 GAL4) using a DS yeast transformation kit (Dualsystems Biotech). Transformed colonies were selected in SD-LW medium to which had been added 5 mM 3-amino-1,2,4-triazole (3-AT) and allowed to grow out for the selection of positive transformants15. Several independent positive transformants were selected and grown in SD-LW liquid medium at 30°C overnight. When the OD546 of the cultures reached 1.0 they were serially diluted (10×, 100× and 1,000×). A 5 μL aliquot of each dilution was spotted on to SD-LW and SD-AHLW solid media, respectively and incubated at 30°C for 3 d. Positive clones were selected and assayed for β-galactosidase activity.
BiFC assay
The BiFC assay37 was used to identify the interaction between CmNAR2 and CmNRT2. The CmNAR2 ORF, excluding its termination codon, was amplified with the primer pair CmNAR2-Bi-S/X (Table 1) and introduced into the pSAT4A vector to give the construct pSAT4A-CmNAR2-cEYFP-N1. Similarly, the CmNRT2 ORF was amplified using the primer pair CmNRT2-Bi-S/X (Table 1) to construct pSAT4A-CmNRT2-nEYFP-N1. The two constructs were mixed with 1:1 gold particles (Bio-Rad) and transformed as described above for the transient expression study. Confocal laser microscopy was used to monitor the expression of YFP.
Functional characterization in Xenopus oocytes
Oocyte preparation and cRNA injection were carried out as described18,38. Oocytes injected with cRNA or water were incubated in NO3− MBS solution consisting of 88 mM NaCl, 1 mM KCl, 2.4 mM NaHCO3, 0.71 mM CaCl2, 0.82 mM MgSO4 and 15 mM HEPES of pH 7.4 for 48 h. To minitor NO3−, injected oocytes were incubated in MBS solution supplied with 0.5 mM NaNO3 at 18°C for 16 h18. Then, they were washed with NO3− free MBS solution. The supernatants were collected using methods as described by Feng et al.18. A 25 ng aliquot of cRNA was injected for each of the single gene assays (CmNRT2) and 25 ng of each of CmNRT2 and CmNAR2 for the two gene assay8. Nitrate uptake assays were performed as set out in the nitrate/nitrite colorimetric assay kit (Cayman Chemical Company).
Transgene construction and A. thaliana transformation
The CmNAR2 and CmNRT2 ORFs were amplified using the primer pair CmNAR (CmNRT2)F/R (Table 1) and inserted into the BamHI and SacI sites of pCAMBIA1301 following digestion with these two enzymes to generate 35S::CmNRT2 and 35S::CmNAR2 constructs. The constructs were named p1301-220-CmNAR2 and p1301-220-CmNRT2. CmNRT2 was amplified using the primer pair CmNRT2 BS/BX (Table 1) and the amplicon cloned into the BglII/BstEII site of p1301-220-CmNAR2 replacing the GUS sequence to form p1301-220-CmNAR2-CmNRT2. This construct was transformed into Agrobacterium tumefaciens strain EHA105 using the freeze-thaw method39. A. thaliana Col-0 was transformed via the floral dip method40. T1 seedlings were raised on agar containing 20 mg/L hygromycin and 25 mg/L ampicillin. Positive transformants were validated by PCR. Specific primers of quantitative RT-PCR for CmNRT2 and CmNAR2 were CmNRT2-qF/CmNRT2-qR for CmNRT2 and NAR2-qF/NAR2-qR for CmNAR2 (Table 1). A portion of the A. thalianaAtUBQ gene (NM_116771.5), amplified by primer pair AtUBQs/AtUBQx, was used as the reference (Table 1).
Root 15N influx
The influx of 15N into seedlings of transgenic and non-transgenic lines was assayed as described previously12. All lines were in the similar developmental stage for root 15N influx testing. Plants were supplied with nutrient media as follows: 0.1 mM KNO3, 0.05 mM Ca(NO3)2, 2.45 mM K2SO4, 2.15 mM CaCl2 and 0.7 mM CaCl2. Nutrient solutions were used at half-strength during the first 2 weeks and changed every 2 days12. After 42 days culture, plants were first exposed to 0.1 mM CaSO4 for 1 min and then to a complete nutrient solution containing 0.2 mM 15N-NO3− for 5 min and finally to 0.1 mM CaSO4 for 1 min. Homogenized root tissue was dried by baking overnight at 80°C and analyzed using the ANCA-MS system (PDZ Europa). The influx of 15N was calculated from the 15N concentrations of the roots.
References
Tang, Z. et al. Knockdown of a rice stelar nitrate transporter alters long-distance translocation but not root influx. Plant Physiol. 160, 2052–2063 (2012).
Kiba, T. et al. The Arabidopsis Nitrate Transporter NRT2.4 Plays a Double Role in Roots and Shoots of Nitrogen-Starved Plants. Plant Cell 24, 245–258 (2012).
Tsay, Y.-F., Ho, C.-H., Chen, H.-Y. & Lin, S.-H. Integration of nitrogen and potassium signaling. Annu. Rev. Plant Biol. 62, 207–226 (2011).
Dechorgnat, J. et al. From the soil to the seeds: the long journey of nitrate in plants. J. Exp. Bot. 62, 1349–1359 (2011).
Guo, F. Q., Wang, R., Chen, M. & Crawford, N. M. The Arabidopsis dual-affinity nitrate transporter gene AtNRT1.1 (CHL1) is activated and functions in nascent organ development during vegetative and reproductive growth. Plant Cell 13, 1761–1777 (2001).
Krouk, G., Crawford, N. M., Coruzzi, G. M. & Tsay, Y. F. Nitrate signaling: adaptation to fluctuating environments. Curr. Opin. Plant Biol. 13, 265–272 (2010).
Forde, B. G. Nitrate transporters in plants: structure, function and regulation. BBA-Biomembranes 1465, 219–235 (2000).
Kotur, Z. et al. Nitrate transport capacity of the Arabidopsis thaliana NRT2 family members and their interactions with AtNAR2. 1. New Phytol. 194, 724–731 (2012).
Quesada, A., Galvan, A. & Fernandez, E. Identification of nitrate transporter genes in Chlamydomonas reinhardtii. Plant J. 5, 407–419 (1994).
Zhou, J. J., Fernández, E., Galván, A. & Miller, A. J. A high affinity nitrate transport system from Chlamydomonas requires two gene products. FEBS lett. 466, 225–227 (2000).
Okamoto, M. et al. High-affinity nitrate transport in roots of Arabidopsis depends on expression of the NAR2-like gene AtNRT3.1. Plant Physiol. 140, 1036–1046 (2006).
Orsel, M., Eulenburg, K., Krapp, A. & Daniel-Vedele, F. Disruption of the nitrate transporter genes AtNRT2.1 and AtNRT2.2 restricts growth at low external nitrate concentration. Planta 219, 714–721 (2004).
Orsel, M. et al. Characterization of a two-component high-affinity nitrate uptake system in Arabidopsis. Physiology and protein-protein interaction. Plant Physiol. 142, 1304–1317 (2006).
Glass, A. D. et al. Nitrogen transport in plants, with an emphasis on the regulations of fluxes to match plant demand. J. Plant Nutr. Soil Sc. 164, 199–207 (2001).
Yan, M. et al. Rice OsNAR2. 1 interacts with OsNRT2.1, OsNRT2.2 and OsNRT2.3a nitrate transporters to provide uptake over high and low concentration ranges. Plant Cell Environ. 34, 1360–1372 (2011).
Huang, C. B., Fang, W. M., Yang, Y., Chen, F. & Xu, Y. Effects of different nitrogen levels and forms on the appearance quality and photosynthesis characters of potted chrysanthemum with small inflorescences. Acta Agr. Zhejiangensis 1, 45–50 (2010).
Jiang, B. B., Fang, W. M., Chen, F. D. & Gu, J. J. Effects of N, P and K ratio on the growth and development of cut chrysanthemum ‘Jinba’. J. Zhejiang Forestry College 6, 692–627 (2008).
Feng, H., Xia, X., Fan, X., Xu, G. & Miller, A. J. Optimizing plant transporter expression in Xenopus oocytes. Plant Methods 9, 48 (2013).
Unkles, S. et al. crnA encodes a nitrate transporter in Aspergillus nidulans. P. Natl. Acad. Sci. USA 88, 204–208 (1991).
Unkles, S. E., Zhou, D., Siddiqi, M. Y., Kinghorn, J. R. & Glass, A. D. Apparent genetic redundancy facilitates ecological plasticity for nitrate transport. EMBO J. 20, 6246–6255 (2001).
Vidmar, J. J., Zhuo, D., Siddiqi, M. Y. & Glass, A. D. M. Isolation and Characterization of HvNRT2.3 and HvNRT2.4, cDNAs Encoding High-Affinity Nitrate Transporters from Roots of Barley. Plant Physiol. 122, 783–792 (2000).
Wirth, J. et al. Regulation of root nitrate uptake at the NRT2.1 protein level in Arabidopsis thaliana. J. Biol. Chem. 282, 23541–23552 (2007).
Feng, H. et al. Spatial expression and regulation of rice high-affinity nitrate transporters by nitrogen and carbon status. J. Exp. Bot. 62, 2319–2332 (2011).
Cai, C. et al. Gene structure and expression of the high-affinity nitrate transport system in rice roots. J. Integr. Plant Biol. 50, 443–451 (2008).
Cai, C. et al. Regulation of the high-affinity nitrate transport system in wheat roots by exogenous abscisic acid and glutamine. J. Integr. Plant Biol. 49, 1719–1725 (2007).
Okamoto, M., Vidmar, J. J. & Glass, A. D. M. Regulation of NRT1 and NRT2 gene families of Arabidopsis thaliana: responses to nitrate provision. Plant Cell Physiol. 44, 304–317 (2003).
Krapp, A. et al. Expression studies of Nrt2:1Np, a putative high-affinity nitrate transporter: evidence for its role in nitrate uptake. Plant J. 14, 723–731 (1998).
He, Q. et al. Cloning and expression study of a putative high-affinity nitrate transporter gene from Dunaliella salina. J. Appl. Phycol. 16, 395–400 (2004).
Yong, Z., Kotur, Z. & Glass, A. D. M. Characterization of an intact two-component high-affinity nitrate transporter from Arabidopsis roots. Plant J. 63, 739–748 (2010).
Gu, C. et al. Reference gene selection for quantitative real-time PCR in Chrysanthemum subjected to biotic and abiotic stress. Mol. Biotechnol. 49, 192–197 (2011).
Shan, H. et al. Heterologous Expression of the Chrysanthemum R2R3-MYB Transcription Factor CmMYB2 Enhances Drought and Salinity Tolerance, Increases Hypersensitivity to ABA and Delays Flowering in Arabidopsis thaliana. Mol. Biotechnol. 51, 160–173 (2011).
Murray, M. & Thompson, W. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4326 (1980).
Liu, Y. G., Mitsukawa, N., Oosumi, T. & Whittier, R. F. Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J. 8, 457–463 (1995).
Chen, B., Sun, C., Wang, Y., Hu, Y. & Lin, Z. Anchored PCR (A-PCR): a new method for chromosome walking. Chinese Sci. Bull. 49, 1772–1774 (2004).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Earley, K. W. et al. Gateway-compatible vectors for plant functional genomics and proteomics. Plant J. 45, 616–629 (2006).
Han, Y. et al. Rice root architecture associated1 binds the proteasome subunit RPT4 and is degraded in a D-box and proteasome-dependent manner. Plant Physiol. 148, 843–855 (2008).
Li, J. et al. The Arabidopsis nitrate transporter NRT1.8 functions in nitrate removal from the xylem sap and mediates cadmium tolerance. Plant Cell 22, 1633–1646 (2010).
Liu, Z. et al. Heterologous expression of a Nelumbo nucifera phytochelatin synthase gene enhances cadmium tolerance in Arabidopsis thaliana. Appl. Biochem. Biotech. 166, 722–734 (2012).
Liu, Z. et al. Identification and expression of an APETALA2-Like gene from Nelumbo nucifera. Appl. Biochem. Biotech. 168, 383–391 (2012).
Acknowledgements
We thank Professor Guohua Xu and Dr. Xiudong Xia (College of Resources and Environmental Sciences, Nanjing Agricultural University) for offer the yeast strains. The study was supported by the National Natural Science Foundation of China (31071820, 31272196, 31272202), the Program for New Century Excellent Talents in University of the Chinese Ministry of Education (NCET-10-0492, NCET-12-0890), the Program for Sci-Tech Support Research, Jiangsu, China (BE2012350, BE2011325), the Fundamental Research Funds for the Central Universities (KYZ201147).
Author information
Authors and Affiliations
Contributions
C.G., S.C. and F.C. conceived the study and designed the experiments. C.G., X.Z., J.J., Z.G. and S.Z. performed the experiments and C.G., W.F. and Y.L. analysed the data with suggestions by S.C. and F.C. and C.G. wrote the manuscript. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
supplementary file 1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Gu, C., Zhang, X., Jiang, J. et al. Chrysanthemum CmNAR2 interacts with CmNRT2 in the control of nitrate uptake. Sci Rep 4, 5833 (2014). https://doi.org/10.1038/srep05833
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05833
This article is cited by
-
An R2R3-MYB Transcription Factor CmMYB42 Improves Low-Nitrogen Stress Tolerance in Chrysanthemum
Journal of Plant Growth Regulation (2023)
-
New insights into the role of chrysanthemum calcineurin B–like interacting protein kinase CmCIPK23 in nitrate signaling in Arabidopsis roots
Scientific Reports (2022)
-
Effects of the Fractionation of the Nitrogen Fertilization on Root Nitrate Uptake and Vine Agronomic Performance in Pinot Gris Grapevines in a Temperate Climate
Journal of Soil Science and Plant Nutrition (2022)
-
Cloning and Molecular Characterization of CcNRT2.1/CcNAR2, a Putative Inducible High Affinity Nitrate Transport System in Capsicum chinense Jacq. Roots
Tropical Plant Biology (2020)
-
CmFTL2 is involved in the photoperiod- and sucrose-mediated control of flowering time in chrysanthemum
Horticulture Research (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.