Abstract
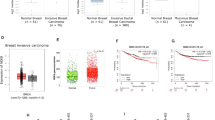
Breast cancer is a major cause of cancer-related deaths in American women; therefore, the identification of novel breast cancer-related molecules for the discovery of new markers and drug targets remains essential. The human DEK gene, which encodes a chromatin-binding protein and DNA topology regulator, is upregulated in many types of cancer. DEK has been implicated as an oncogene in breast cancer based on mRNA expression studies, but its functional significance in breast cancer growth and progression has not yet been tested directly. We demonstrate that DEK is highly expressed in breast cancer cells compared with normal tissue, and functionally important for cellular growth, invasion and mammosphere formation. DEK overexpression in non-tumorigenic MCF10A cells resulted in increased growth and motility, with a concomitant downregulation of E-cadherin. Conversely, DEK knockdown in MCF7 and MDA-MB-468 breast cancer cells resulted in decreased growth and motility with upregulation of E-cadherin. The use of DEK-proficient and -deficient breast cancer cells in orthotopic xenografts provided further in vivo evidence that DEK contributes to tumor growth. Activation of the β-catenin signaling pathway is important for normal and cancer stem cell character, growth and metastasis. We show that DEK expression stimulated, and DEK knockdown repressed β-catenin nuclear translocation and activity. Importantly, the expression of constitutively active β-catenin rescued breast cancer invasion defects of DEK knockdown cells. Together, our data indicate that DEK expression stimulates the growth, stem cell character and motility of breast cancer cells, and that DEK-dependent cellular invasion occurs at least in part via β-catenin activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Abba MC, Sun H, Hawkins KA, Drake JA, Hu Y, Nunez MI et al. (2007). Breast cancer molecular signatures as determined by SAGE: correlation with lymph node status. Mol Cancer Res 5: 881–890.
ACS, Breast Cancer Facts & Figures 2009-2010. American Cancer Society, Inc.: Atlanta, 2009.
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF . (2003). Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 100: 3983–3988.
Alexiadis V, Waldmann T, Andersen J, Mann M, Knippers R, Gruss C . (2000). The protein encoded by the proto-oncogene DEK changes the topology of chromatin and reduces the efficiency of DNA replication in a chromatin-specific manner. Genes Dev 14: 1308–1312.
Allan AL, George R, Vantyghem SA, Lee MW, Hodgson NC, Engel CJ et al. (2006). Role of the integrin-binding protein osteopontin in lymphatic metastasis of breast cancer. Am J Pathol 169: 233–246.
Andreassen PR, Margolis RL . (1994). Microtubule dependency of p34cdc2 inactivation and mitotic exit in mammalian cells. J Cell Biol 127: 789–802.
Bosco EE, Knudsen ES . (2007). RB in breast cancer: at the crossroads of tumorigenesis and treatment. Cell Cycle 6: 667–671.
Bowles E, Corson TW, Bayani J, Squire JA, Wong N, Lai PB et al. (2007). Profiling genomic copy number changes in retinoblastoma beyond loss of RB1. Genes Chromosomes Cancer 46: 118–129.
Campillos M, Garcia MA, Valdivieso F, Vazquez J . (2003). Transcriptional activation by AP-2alpha is modulated by the oncogene DEK. Nucleic Acids Res 31: 1571–1575.
Carro MS, Spiga FM, Quarto M, Di Ninni V, Volorio S, Alcalay M et al. (2006). DEK Expression is controlled by E2F and deregulated in diverse tumor types. Cell Cycle 5: 1202–1207.
Dignam JD, Lebovitz RM, Roeder RG . (1983). Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res 11: 1475–1489.
Dontu G, Abdallah WM, Foley JM, Jackson KW, Clarke MF, Kawamura MJ et al. (2003). In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev 17: 1253–1270.
Dontu G, Liu S, Wicha MS . (2005). Stem cells in mammary development and carcinogenesis: implications for prevention and treatment. Stem Cell Rev 1: 207–213.
Euhus DM, Hudd C, LaRegina MC, Johnson FE . (1986). Tumor measurement in the nude mouse. J Surg Oncol 31: 229–234.
Evans AJ, Gallie BL, Jewett MA, Pond GR, Vandezande K, Underwood J et al. (2004). Defining a 0.5-mb region of genomic gain on chromosome 6p22 in bladder cancer by quantitative-multiplex polymerase chain reaction. Am J Pathol 164: 285–293.
Goodell MA, Brose K, Paradis G, Conner AS, Mulligan RC . (1996). Isolation and functional properties of murine hematopoietic stem cells that are replicating in vivo. J Exp Med 183: 1797–1806.
Guarino M, Rubino B, Ballabio G . (2007). The role of epithelial-mesenchymal transition in cancer pathology. Pathology 39: 305–318.
Heuberger J, Birchmeier W . (2010). Interplay of cadherin-mediated cell adhesion and canonical Wnt signaling. Cold Spring Harb Perspect Biol 2: a002915.
Johung K, Goodwin EC, DiMaio D . (2007). Human papillomavirus E7 repression in cervical carcinoma cells initiates a transcriptional cascade driven by the retinoblastoma family, resulting in senescence. J Virol 81: 2102–2116.
Kappes F, Burger K, Baack M, Fackelmayer FO, Gruss C . (2001). Subcellular localization of the human proto-oncogene protein DEK. J Biol Chem 276: 26317–26323.
Kappes F, Fahrer J, Khodadoust MS, Tabbert A, Strasser C, Mor-Vaknin N et al. (2008). DEK is a poly(ADP-ribose) acceptor in apoptosis and mediates resistance to genotoxic stress. Mol Cell Biol 28: 3245–3257.
Khodadoust MS, Verhaegen M, Kappes F, Riveiro-Falkenbach E, Cigudosa JC, Kim DS et al. (2009). Melanoma proliferation and chemoresistance controlled by the DEK oncogene. Cancer Res 69: 6405–6413.
Kondoh N, Wakatsuki T, Ryo A, Hada A, Aihara T, Horiuchi S et al. (1999). Identification and characterization of genes associated with human hepatocellular carcinogenesis. Cancer Res 59: 4990–4996.
Le Hir H, Gatfield D, Izaurralde E, Moore MJ . (2001). The exon-exon junction complex provides a binding platform for factors involved in mRNA export and nonsense-mediated mRNA decay. Embo J 20: 4987–4997.
Li Y, Welm B, Podsypanina K, Huang S, Chamorro M, Zhang X et al. (2003). Evidence that transgenes encoding components of the Wnt signaling pathway preferentially induce mammary cancers from progenitor cells. Proc Natl Acad Sci USA 100: 15853–15858.
Lu ZL, Luo DZ, Wen JM . (2005). Expression and significance of tumor-related genes in HCC. World J Gastroenterol 11: 3850–3854.
McGarvey T, Rosonina E, McCracken S, Li Q, Arnaout R, Mientjes E et al. (2000). The acute myeloid leukemia-associated protein, DEK, forms a splicing-dependent interaction with exon-product complexes. J Cell Biol 150: 309–320.
Miller LD, Smeds J, George J, Vega VB, Vergara L, Ploner A et al. (2005). An expression signature for p53 status in human breast cancer predicts mutation status, transcriptional effects, and patient survival. Proc Natl Acad Sci USA 102: 13550–13555.
Prahalad P, Calvo I, Waechter H, Matthews JB, Zuk A, Matlin KS . (2004). Regulation of MDCK cell-substratum adhesion by RhoA and myosin light chain kinase after ATP depletion. Am J Physiol Cell Physiol 286: C693–C707.
Privette LM, Gonzalez ME, Ding L, Kleer CG, Petty EM . (2007). Altered expression of the early mitotic checkpoint protein, CHFR, in breast cancers: implications for tumor suppression. Cancer Res 67: 6064–6074.
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D et al. (2004). ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6: 1–6.
Ribeiro-Silva A, Zambelli Ramalho LN, Britto Garcia S, Zucoloto S . (2003). The relationship between p63 and p53 expression in normal and neoplastic breast tissue. Arch Pathol Lab Med 127: 336–340.
Richardson AL, Wang ZC, De Nicolo A, Lu X, Brown M, Miron A et al. (2006). X chromosomal abnormalities in basal-like human breast cancer. Cancer Cell 9: 121–132.
Sammons M, Wan SS, Vogel NL, Mientjes EJ, Grosveld G, Ashburner BP . (2006). Negative regulation of the RelA/p65 transactivation function by the product of the DEK proto-oncogene. J Biol Chem 281: 26802–26812.
Schmittgen TD, Livak KJ . (2008). Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3: 1101–1108.
Shibata T, Kokubu A, Miyamoto M, Hosoda F, Gotoh M, Tsuta K et al. (2010). DEK oncoprotein regulates transcriptional modifiers and sustains tumor initiation activity in high-grade neuroendocrine carcinoma of the lung. Oncogene 29: 4671–4681.
Soares LM, Zanier K, Mackereth C, Sattler M, Valcarcel J . (2006). Intron removal requires proofreading of U2AF/3′ splice site recognition by DEK. Science 312: 1961–1965.
Soule HD, Maloney TM, Wolman SR, Peterson Jr WD, Brenz R, McGrath CM et al. (1990). Isolation and characterization of a spontaneously immortalized human breast epithelial cell line, MCF-10. Cancer Res 50: 6075–6086.
Tomayko MM, Reynolds CP . (1989). Determination of subcutaneous tumor size in athymic (nude) mice. Cancer Chemother Pharmacol 24: 148–154.
Tubiana M, Koscielny S . (1999). The rationale for early diagnosis of cancer-the example of breast cancer. Acta Oncol 38: 295–303.
van't Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M et al. (2002). Gene expression profiling predicts clinical outcome of breast cancer. Nature 415: 530–536.
van de Vijver MJ, He YD, van't Veer LJ, Dai H, Hart AA, Voskuil DW et al. (2002). A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347: 1999–2009.
Waldmann T, Baack M, Richter N, Gruss C . (2003). Structure-specific binding of the proto-oncogene protein DEK to DNA. Nucleic Acids Res 31: 7003–7010.
Waldmann T, Eckerich C, Baack M, Gruss C . (2002). The ubiquitous chromatin protein DEK alters the structure of DNA by introducing positive supercoils. J Biol Chem 277: 24988–24994.
Waldmann T, Scholten I, Kappes F, Hu HG, Knippers R . (2004). The DEK protein-an abundant and ubiquitous constituent of mammalian chromatin. Gene 343: 1–9.
Wang Y, Klijn JG, Zhang Y, Sieuwerts AM, Look MP, Yang F et al. (2005). Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365: 671–679.
Wise-Draper TM, Allen HV, Jones EE, Habash KB, Matsuo H, Wells SI . (2006). Apoptosis inhibition by the human DEK oncoprotein involves interference with p53 functions. Mol Cell Biol 26: 7506–7519.
Wise-Draper TM, Allen HV, Thobe MN, Jones EE, Habash KB, Munger K et al. (2005). The human DEK proto-oncogene is a senescence inhibitor and an upregulated target of high-risk human papillomavirus E7. J Virol 79: 14309–14317.
Wise-Draper TM, Mintz-Cole RA, Morris TA, Simpson DS, Wikenheiser-Brokamp KA, Currier MA et al. (2009a). Overexpression of the cellular DEK protein promotes epithelial transformation in vitro and in vivo. Cancer Res 69: 1792–1799.
Wise-Draper TM, Morreale RJ, Morris TA, Mintz-Cole RA, Hoskins EE, Balsitis SJ et al. (2009b). DEK proto-oncogene expression interferes with the normal epithelial differentiation program. Am J Pathol 174: 71–81.
Woodward WA, Chen MS, Behbod F, Rosen JM . (2005). On mammary stem cells. J Cell Sci 118: 3585–3594.
Zhang M, Behbod F, Atkinson RL, Landis MD, Kittrell F, Edwards D et al. (2008). Identification of tumor-initiating cells in a p53-null mouse model of breast cancer. Cancer Res 68: 4674–4682.
Acknowledgements
We thank Yi Zheng for critical evaluation of this manuscript. We also thank James Lessard, Aaron Zorn, Jose Cancelas, James Mulloy and James Wells for reagents and discussion, Gerard Grosveld for Dek knockout mice, Gina Kavanaugh for the immortalized Dek wild-type and knockout mouse embryonic fibroblasts and the Viral Vector Core at CCHMC. We would like to acknowledge the assistance of the Research Flow Cytometry Core in the Division of Rheumatology at Cincinnati Children's Hospital Medical Center, supported in part by NIH AR-47363. This research was supported by Kirschstein National Research Service Awards (NRSA) F32CA139931 and T32HL091805 from the National Cancer Institute (L.M.P.V.), and Public Health Service grants CA116316 (S.I.W.), CA100002 (S.E.W.), and HL079193 (K. W-B.)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Privette Vinnedge, L., McClaine, R., Wagh, P. et al. The human DEK oncogene stimulates β-catenin signaling, invasion and mammosphere formation in breast cancer. Oncogene 30, 2741–2752 (2011). https://doi.org/10.1038/onc.2011.2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.2
Keywords
This article is cited by
-
A three layered histone epigenetics in breast cancer metastasis
Cell & Bioscience (2020)
-
Role of the DEK oncogene in the development of squamous cell carcinoma
International Journal of Clinical Oncology (2020)
-
Optical Redox Imaging Detects the Effects of DEK Oncogene Knockdown on the Redox State of MDA-MB-231 Breast Cancer Cells
Molecular Imaging and Biology (2019)
-
The potential predictive value of DEK expression for neoadjuvant chemoradiotherapy response in locally advanced rectal cancer
BMC Cancer (2018)
-
DEK is required for homologous recombination repair of DNA breaks
Scientific Reports (2017)