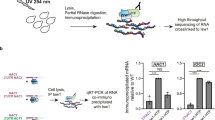
Abstract
Both p150 and p110 isoforms of ADAR1 convert adenosine to inosine in double-stranded RNA (dsRNA). ADAR1p150 suppresses the dsRNA-sensing mechanism that activates MDA5–MAVS–IFN signaling in the cytoplasm. In contrast, the biological function of the ADAR1p110 isoform, which is usually located in the nucleus, is largely unknown. Here, we show that stress-activated phosphorylation of ADAR1p110 by MKK6–p38–MSK MAP kinases promotes its binding to Exportin-5 and its export from the nucleus. After translocating to the cytoplasm, ADAR1p110 suppresses apoptosis in stressed cells by protecting many antiapoptotic gene transcripts that contain 3′-untranslated-region dsRNA structures primarily comprising inverted Alu repeats. ADAR1p110 competitively inhibits binding of Staufen1 to the 3′-untranslated-region dsRNAs and antagonizes Staufen1-mediated mRNA decay. Our study reveals a new stress-response mechanism in which human ADAR1p110 and Staufen1 regulate surveillance of a set of mRNAs required for survival of stressed cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Referenced accessions
NCBI Reference Sequence
References
Nishikura, K. A-to-I editing of coding and non-coding RNAs by ADARs. Nat. Rev. Mol. Cell Biol. 17, 83–96 (2016).
Mannion, N., Arieti, F., Gallo, A., Keegan, L.P. & O'Connell, M.A. New insights into the biological role of mammalian ADARs: the RNA editing proteins. Biomolecules 5, 2338–2362 (2015).
Hood, J.L. & Emeson, R.B. Editing of neurotransmitter receptor and ion channel RNAs in the nervous system. Curr. Top. Microbiol. Immunol. 353, 61–90 (2012).
Savva, Y.A., Rieder, L.E. & Reenan, R.A. The ADAR protein family. Genome Biol. 13, 252 (2012).
Kawahara, Y. et al. Redirection of silencing targets by adenosine-to-inosine editing of miRNAs. Science 315, 1137–1140 (2007).
Yang, W. et al. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nat. Struct. Mol. Biol. 13, 13–21 (2006).
Bazak, L., Levanon, E.Y. & Eisenberg, E. Genome-wide analysis of Alu editability. Nucleic Acids Res. 42, 6876–6884 (2014).
Ramaswami, G. et al. Accurate identification of human Alu and non-Alu RNA editing sites. Nat. Methods 9, 579–581 (2012).
Patterson, J.B. & Samuel, C.E. Expression and regulation by interferon of a double-stranded-RNA-specific adenosine deaminase from human cells: evidence for two forms of the deaminase. Mol. Cell. Biol. 15, 5376–5388 (1995).
Liu, Y., George, C.X., Patterson, J.B. & Samuel, C.E. Functionally distinct double-stranded RNA-binding domains associated with alternative splice site variants of the interferon-inducible double-stranded RNA-specific adenosine deaminase. J. Biol. Chem. 272, 4419–4428 (1997).
Hartner, J.C. et al. Liver disintegration in the mouse embryo caused by deficiency in the RNA-editing enzyme ADAR1. J. Biol. Chem. 279, 4894–4902 (2004).
Hartner, J.C., Walkley, C.R., Lu, J. & Orkin, S.H. ADAR1 is essential for the maintenance of hematopoiesis and suppression of interferon signaling. Nat. Immunol. 10, 109–115 (2009).
Wang, Q. et al. Stress-induced apoptosis associated with null mutation of ADAR1 RNA editing deaminase gene. J. Biol. Chem. 279, 4952–4961 (2004).
Mannion, N.M. et al. The RNA-editing enzyme ADAR1 controls innate immune responses to RNA. Cell Rep. 9, 1482–1494 (2014).
Pestal, K. et al. Isoforms of RNA-editing enzyme ADAR1 independently control nucleic acid sensor MDA5-driven autoimmunity and multi-organ development. Immunity 43, 933–944 (2015).
Liddicoat, B.J. et al. RNA editing by ADAR1 prevents MDA5 sensing of endogenous dsRNA as nonself. Science 349, 1115–1120 (2015).
Ward, S.V. et al. RNA editing enzyme adenosine deaminase is a restriction factor for controlling measles virus replication that also is required for embryogenesis. Proc. Natl. Acad. Sci. USA 108, 331–336 (2011).
Coulthard, L.R., White, D.E., Jones, D.L., McDermott, M.F. & Burchill, S.A. p38(MAPK): stress responses from molecular mechanisms to therapeutics. Trends Mol. Med. 15, 369–379 (2009).
Cuadrado, A. & Nebreda, A.R. Mechanisms and functions of p38 MAPK signalling. Biochem. J. 429, 403–417 (2010).
Gong, C. & Maquat, L.E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470, 284–288 (2011).
Gong, C., Tang, Y. & Maquat, L.E. mRNA–mRNA duplexes that autoelicit Staufen1-mediated mRNA decay. Nat. Struct. Mol. Biol. 20, 1214–1220 (2013).
Kim, Y.K., Furic, L., Desgroseillers, L. & Maquat, L.E. Mammalian Staufen1 recruits Upf1 to specific mRNA 3′UTRs so as to elicit mRNA decay. Cell 120, 195–208 (2005).
Anderson, P. Post-transcriptional regulons coordinate the initiation and resolution of inflammation. Nat. Rev. Immunol. 10, 24–35 (2010).
Schoenberg, D.R. & Maquat, L.E. Regulation of cytoplasmic mRNA decay. Nat. Rev. Genet. 13, 246–259 (2012).
Ng, S.K., Weissbach, R., Ronson, G.E. & Scadden, A.D. Proteins that contain a functional Z-DNA-binding domain localize to cytoplasmic stress granules. Nucleic Acids Res. 41, 9786–9799 (2013).
Seet, B.T., Dikic, I., Zhou, M.M. & Pawson, T. Reading protein modifications with interaction domains. Nat. Rev. Mol. Cell Biol. 7, 473–483 (2006).
Emrick, M.A., Hoofnagle, A.N., Miller, A.S., Ten Eyck, L.F. & Ahn, N.G. Constitutive activation of extracellular signal-regulated kinase 2 by synergistic point mutations. J. Biol. Chem. 276, 46469–46479 (2001).
Raingeaud, J., Whitmarsh, A.J., Barrett, T., Dérijard, B. & Davis, R.J. MKK3- and MKK6-regulated gene expression is mediated by the p38 mitogen-activated protein kinase signal transduction pathway. Mol. Cell. Biol. 16, 1247–1255 (1996).
Ahn, Y.H. et al. Map2k4 functions as a tumor suppressor in lung adenocarcinoma and inhibits tumor cell invasion by decreasing peroxisome proliferator-activated receptor γ2 expression. Mol. Cell. Biol. 31, 4270–4285 (2011).
Barancík, M. et al. SB203580, a specific inhibitor of p38-MAPK pathway, is a new reversal agent of P-glycoprotein-mediated multidrug resistance. Eur. J. Pharm. Sci. 14, 29–36 (2001).
Wood, C.D., Thornton, T.M., Sabio, G., Davis, R.A. & Rincon, M. Nuclear localization of p38 MAPK in response to DNA damage. Int. J. Biol. Sci. 5, 428–437 (2009).
Cho, D.S. et al. Requirement of dimerization for RNA editing activity of adenosine deaminases acting on RNA. J. Biol. Chem. 278, 17093–17102 (2003).
Nishikura, K., Sakurai, M., Ariyoshi, K. & Ota, H. Antagonistic and stimulative roles of ADAR1 in RNA silencing. RNA Biol. 10, 1240–1247 (2013).
Ota, H. et al. ADAR1 forms a complex with Dicer to promote microRNA processing and RNA-induced gene silencing. Cell 153, 575–589 (2013).
Hajdarpašić, A. & Ruggenthaler, P. Analysis of miRNA expression under stress in Arabidopsisthaliana . Bosn. J. Basic Med. Sci. 12, 169–176 (2012).
Jones-Rhoades, M.W. & Bartel, D.P. Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol. Cell 14, 787–799 (2004).
Fritz, J. et al. RNA-regulated interaction of transportin-1 and exportin-5 with the double-stranded RNA-binding domain regulates nucleocytoplasmic shuttling of ADAR1. Mol. Cell. Biol. 29, 1487–1497 (2009).
Gwizdek, C. et al. Minihelix-containing RNAs mediate exportin-5-dependent nuclear export of the double-stranded RNA-binding protein ILF3. J. Biol. Chem. 279, 884–891 (2004).
Wada, T. & Penninger, J.M. Mitogen-activated protein kinases in apoptosis regulation. Oncogene 23, 2838–2849 (2004).
Zarubin, T. & Han, J. Activation and signaling of the p38 MAP kinase pathway. Cell Res. 15, 11–18 (2005).
Vitali, P. & Scadden, A.D. Double-stranded RNAs containing multiple IU pairs are sufficient to suppress interferon induction and apoptosis. Nat. Struct. Mol. Biol. 17, 1043–1050 (2010).
de Lucas, S., Oliveros, J.C., Chagoyen, M. & Ortín, J. Functional signature for the recognition of specific target mRNAs by human Staufen1 protein. Nucleic Acids Res. 42, 4516–4526 (2014).
Elbarbary, R.A., Li, W., Tian, B. & Maquat, L.E. STAU1 binding 3′ UTR IRAlus complements nuclear retention to protect cells from PKR-mediated translational shutdown. Genes Dev. 27, 1495–1510 (2013).
Park, E. & Maquat, L.E. Staufen-mediated mRNA decay. Wiley Interdiscip. Rev. RNA 4, 423–435 (2013).
Balagopal, V., Fluch, L. & Nissan, T. Ways and means of eukaryotic mRNA decay. Biochim. Biophys. Acta 1819, 593–603 (2012).
Takao, N., Li, Y. & Yamamoto, K. Protective roles for ATM in cellular response to oxidative stress. FEBS Lett. 472, 133–136 (2000).
Raderschall, E. et al. Formation of higher-order nuclear Rad51 structures is functionally linked to p21 expression and protection from DNA damage-induced apoptosis. J. Cell Sci. 115, 153–164 (2002).
Wang, X., Vukovic, L., Koh, H.R., Schulten, K. & Myong, S. Dynamic profiling of double-stranded RNA binding proteins. Nucleic Acids Res. 43, 7566–7576 (2015).
Thornton, T.M. & Rincon, M. Non-classical p38 map kinase functions: cell cycle checkpoints and survival. Int. J. Biol. Sci. 5, 44–51 (2009).
Lai, F., Drakas, R. & Nishikura, K. Mutagenic analysis of double-stranded RNA adenosine deaminase, a candidate enzyme for RNA editing of glutamate-gated ion channel transcripts. J. Biol. Chem. 270, 17098–17105 (1995).
Gleghorn, M.L., Gong, C., Kielkopf, C.L. & Maquat, L.E. Staufen1 dimerizes through a conserved motif and a degenerate dsRNA-binding domain to promote mRNA decay. Nat. Struct. Mol. Biol. 20, 515–524 (2013).
Wagner, R.W., Smith, J.E., Cooperman, B.S. & Nishikura, K. A double-stranded RNA unwinding activity introduces structural alterations by means of adenosine to inosine conversions in mammalian cells and Xenopus eggs. Proc. Natl. Acad. Sci. USA 86, 2647–2651 (1989).
Brownawell, A.M. & Macara, I.G. Exportin-5, a novel karyopherin, mediates nuclear export of double-stranded RNA binding proteins. J. Cell Biol. 156, 53–64 (2002).
Nishi, K., Nishi, A., Nagasawa, T. & Ui-Tei, K. Human TNRC6A is an Argonaute-navigator protein for microRNA-mediated gene silencing in the nucleus. RNA 19, 17–35 (2013).
Pollard, V.W. et al. A novel receptor-mediated nuclear protein import pathway. Cell 86, 985–994 (1996).
Ricci, E.P. et al. Staufen1 senses overall transcript secondary structure to regulate translation. Nat. Struct. Mol. Biol. 21, 26–35 (2014).
Niranjanakumari, S., Lasda, E., Brazas, R. & Garcia-Blanco, M.A. Reversible cross-linking combined with immunoprecipitation to study RNA-protein interactions in vivo . Methods 26, 182–190 (2002).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Kelley, L.A., Mezulis, S., Yates, C.M., Wass, M.N. & Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Stefl, R. et al. The solution structure of the ADAR2 dsRBM-RNA complex reveals a sequence-specific readout of the minor groove. Cell 143, 225–237 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
The PyMOL Molecular Graphics System v. 1.8. (Schrödinger, LLC).
Li, B. & Dewey, C.N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Acknowledgements
We thank Z. Paroo (University of Illinois), M. Cobb (University of Texas), S. Janiki (Wistar Institute), G. Dreyfuss (University of Pennsylvania), and L. Maquat (University of Rochester) for various recombinant-protein-expression constructs. We also thank J. Hayden for assistance with fluorescence microscopy, Y. Sakurai for technical assistance, and M. Murphy and J.M. Murray for critical reading of the manuscript. Research reported in this publication was supported by grants to K.N. from the National Institutes of Health (R01GM040536 and R01CA175058), the Macula Vision Research Foundation, and the Japan Society for the Promotion of Science (S13204). C.S. is supported as a member of the Roy and Dianna Vagelos Scholars Program in Molecular Life Sciences at the University of Pennsylvania. M.S. is supported in part by a fellowship from the Japan Society for the Promotion of Science (JSPS 2010-22). We also acknowledge services provided by the Animal, Bioinformatics, Genomics, Imaging, Protein Expression, and Proteomics Shared Facilities of The Wistar Cancer Center, which are supported by the National Cancer Institute (P30CA010815). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
M.S., Y.S., and K.N. designed the study. M.S. conducted fluorescence microscopy experiments, apoptosis analysis, and RIP experiments. M.S. and H.O. conducted ADAR1-Xpo5 interaction studies. M.S. and Y.S. performed deep-sequencing sample preparation, bioinformatics data analysis, and gene expression analysis. C.S. prepared mutant ADAR1 constructs. A.V.K., J.W., and L.C.S. performed bioinformatics analysis of sequencing data. E.S. conducted molecular modeling studies. H.-Y.T. and D.W.S. performed LC-MS/MS analysis and data interpretation. M.S., Y.S., and K.N. supervised research, reviewed data, and wrote and edited the paper. All authors agree to the contents of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cellular localization of wild-type and phosphorylation mutants of mCherry-ADAR1p150 in A172 cells.
(a) UV irradiation did not affect overall cytoplasmic localization of ADAR1p150 in transiently transfected A172 human glioblastoma cells. (b) Fractional move to stress-granules was noted in A172 cells as reported previously (Weissbach and Scadden RNA 18, 462-71, 2012), as confirmed by co-localization of ADAR1p150 with a stress-granule marker, TIA1. TIA1-EGFP was a kind gift of John L. Goodier. (c) Cytoplasmic localization of ADAR1p150 was unaffected by phosphorylation site mutations. Scale bars, 20 μm.
Supplementary Figure 2 Cellular distribution of endogenous ADAR1p150 and p110 proteins in unirradiated and UV-irradiated A172 cells.
Nuclear and cytoplasmic fractions of control and UV-irradiated A172 cells were examined by western blotting analysis. Histone H3 and β-Actin were used as nuclear and cytoplasmic markers, respectively. UV irradiation increased significantly the cytoplasmic ADAR1p110, which was reversed by SB203580 p38 inhibitor.
Supplementary Figure 3 Phosphorylation of ADAR1p110 does not affect its A-to-I RNA-editing activity or its Dicer-promoting activity.
(a) In vitro A-to-I editing assay was carried out using HAT-ADAR1p110-WT, -T/S-to-D, or –T/S-to-A recombinant proteins and 32P-ATP labeled c-myc dsRNA substrate as described previously (Wagner et al. Proc. Natl. Acad. Sci. 86, 2647-51, 1989). A-to-I conversion was quantitated using TLC assay. (b) In vitro Dicing assay was carried out using 5’ 32P-labeled pre-let-7a substrate RNA, and results were quantitated as described previously (Ota et al. Cell 153, 575-89, 2013). (a, b) Data are mean ± S.D. (n = 3, technical replicates); significant differences were identified by two-tailed Student’s t-tests: ***P < 0.001, n.s., not significant. Source data are available online in Supplementary Data Set 1.
Supplementary Figure 4 UV-irradiation-induced cytoplasmic localization of ADAR1p110 depends on Xpo5.
(a) UV irradiation induced cytoplasmic localization of ADAR1p110-WT is inhibited by knockdown of Xpo5 in A172 cells. (b) Cytoplasmic localization of T/S-to-D phosphomimetic mutant of ADAR1p110 is also inhibited in un-irradiated A172 cells by knockdown of Xpo5. (c) Nuclear localization of T/S-to-A phosphorylation inhibitory mutant of ADAR1p110 is inhibited in un-irradiated A172 cells by knockdown of TRN1. Scale bars, 20 μm.
Supplementary Figure 5 Apoptosis induced in ADAR1-knockdown and UV-irradiated cells.
(a) Fluorescence microscopic detection of apoptosis induced in ADAR1 knocked down and UV-irradiated A172 cells in comparison to the cells treated with an inducer of apoptosis, Camptothecin (20 μM, 48 hrs). Apoptosis was evaluated by staining with AnnexinV-Alexa594 and also by CellEventTM Caspase-3/7 Green system (Themo Fischer), which utilizes a fluorogenic substrate for activated Caspase-3 and -7. Scale bars, 200 μm. (b) Apoptosis induced in UV-irradiated U2OS cells was evaluated by ApoTox-Glo Triplex apoptosis assay (Promega). ADAR1p150 specific knockdown did not cause UV-induced apoptosis. Knockdown of Staufen1 or UPF1 suppressed apoptosis induced by ADAR1 knockdown. Positive apoptosis control: un-irradiated U2OS cells treated with Camptothecin (20 μM) for 48 hrs. (c) The relative expression levels of FLAG-ADAR1p110 and endogenous ADAR1 (p150 and p110) in UV-irradiated A172 cells transiently transfected with FLAG-ADAR1p110 expression constructs were examined by western blotting analysis. Expression levels of the exogenous FLAG-ADAR1p110 proteins are about equal to those of endogenous ADAR1p110 proteins (compare lane 1 to lane 4 and 5). (d) The expression levels of ADAR1p150 and p110 as well as two ISGs, IFIT1 and MDA5, in A172 cells were examined by western blotting analysis of cell extracts. Note that UV-irradiation did not affect the expression of ADAR1 (both p150 and p110). Furthermore, ADAR1 knockdown alone did not induce the expression of IFIT1 or MDA5 in both un-irradiated and UV-irradiated cells. However, transfection of poly(I:C) dsRNA (Invivogen) for 12 hrs increased the expression of ADAR1p150 as well as IFIT1 and MDA5 in a dose dependent manner, indicating that the dsRNA sensing mechanism mediated by the MDA5-MAVS-IFN pathway remains intact in this cell line. (e) Apoptosis induced in UV-irradiated A172 cells was evaluated by ApoTox-Glo Triplex apoptosis assay (Promega). Knockdown of Staufen1 and/or UPF1 did not induce but suppressed apoptosis in UV irradiated and ADAR1 knocked down cells. (b, e) Data are mean ± S.D. (n = 4, cell cultures); significant differences were identified by two-tailed Student’s t-tests:***P < 0.001. Source data are available online in Supplementary Data Set 1.
Supplementary Figure 6 Secondary structures of 3′-UTR Alu dsRNAs.
The secondary structures of CASC5 (a) and ATM (b) 3’UTR Alu dsRNA predicted by mfold are shown. A-to-I editing sites detected by RNA-seq data analysis are indicated by red arrows with editing frequency. Primers used for RIP analysis are indicated as black arrows.
Supplementary Figure 7 Colocalization of Staufen1-EGFP and mCherry-ADAR1p110 in the cytoplasm of unirradiated and UV-irradiated A172 cells.
Staufen1 localizes in the cytoplasm of A172 cells regardless of UV irradiation. Scale bars, 20 μm.
Supplementary Figure 8 Quantitative RT–PCR analysis of pre-mRNAs.
Pre-mRNA levels of four SMD target genes containing 3’UTR Alu dsRNAs were analyzed by qRT-PCR. The expression of CCNG1 pre-mRNAs was slightly increased by ADAR1 knockdown. The pre-mRNA levels of the remaining SMD target genes were unchanged. Data are mean ± S.D. (n = 3 for HPRT1, 4 for other genes, technical replicates); significant differences were identified by two-tailed Student’s t-tests: *P < 0.05, **P < 0.01, n.s., not significant. Source data are available online in Supplementary Data Set 1.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1699 kb)
Supplementary Table 1
RNA-seq analysis for gene transcripts reduced by ADAR1 knockdown & rescued by Staufen1 knockdown. (XLSX 2338 kb)
Supplementary Table 2
RNA-seq analysis for ISG Transcripts (XLSX 1139 kb)
Supplementary Table 3
qRT-PCR analysis for 52 gene transcripts decreased by ADAR1 knockdown (XLSX 29 kb)
Supplementary Table 4
GO enrichment analysis (XLSX 21 kb)
Supplementary Table 5
Genes analyzed for RIP and qRT-PCR (XLSX 491 kb)
Supplementary Table 6
DNA and RNA oligos used in this study (XLSX 15 kb)
Supplementary Table 7
Antibodies used in this study (XLSX 10 kb)
Supplementary Data Set 1
Excel data source for all bar graphs (XLSX 364 kb)
Supplementary Data Set 2
PDF files for all western blotting original gel images (PDF 39102 kb)
Rights and permissions
About this article
Cite this article
Sakurai, M., Shiromoto, Y., Ota, H. et al. ADAR1 controls apoptosis of stressed cells by inhibiting Staufen1-mediated mRNA decay. Nat Struct Mol Biol 24, 534–543 (2017). https://doi.org/10.1038/nsmb.3403
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3403
This article is cited by
-
The role of ADAR1 through and beyond its editing activity in cancer
Cell Communication and Signaling (2024)
-
Inverted Alu repeats: friends or foes in the human transcriptome
Experimental & Molecular Medicine (2024)
-
Emerging role of the RNA-editing enzyme ADAR1 in stem cell fate and function
Biomarker Research (2023)
-
RNA modifications in cardiovascular health and disease
Nature Reviews Cardiology (2023)
-
The cellular and KSHV A-to-I RNA editome in primary effusion lymphoma and its role in the viral lifecycle
Nature Communications (2023)