Abstract
The association of Zika virus (ZIKV) infections with microcephaly has resulted in an ongoing public-health emergency. Here we report the crystal structure of a C-terminal fragment of ZIKV nonstructural protein 1 (NS1), a major host-interaction molecule that functions in flaviviral replication, pathogenesis and immune evasion. Comparison with West Nile and dengue virus NS1 structures reveals conserved features but diverse electrostatic characteristics at host-interaction interfaces, thus possibly implying different modes of flavivirus pathogenesis.
Similar content being viewed by others
Main
ZIKV, a Flaviviridae family member transmitted to humans by mosquitoes of the genus Aedes, was first isolated in Africa in 1947 (ref. 1) and has been found to commonly circulate in tropical regions in Africa and Asia2. After a large Zika outbreak in French Polynesia from 2013 to 2014, the virus was able to emerge into new territories of the American continent3, and it has been found circulating in 26 countries and territories in South America and the Caribbean4. Before this outbreak, research on ZIKV pathogenesis was largely neglected because infected individuals are often asymptomatic or have mild symptoms. However there is now growing evidence that ZIKV infections may be linked to fetal and newborn microcephaly and serious neurological complications, such as Guillain–Barré syndrome4. ZIKV has been detected in the amniotic fluid of pregnant women whose fetuses had microcephaly syndrome and has also been detected in microcephalic fetal brain tissues5,6. Moreover, ZIKV infects human cortical neural progenitor cells and attenuates their growth7. Another matter of concern is that ZIKV may also be transmitted by sexual activity, and high viral loads have been detected in semen from infected patients8. To date, no clinically approved vaccines or therapeutics are available to prevent and control ZIKV infection. Given this urgent situation, great efforts are needed to develop new vaccines and antiviral therapeutics; to do so, a comprehensive understanding of the pathogenesis and molecular basis of ZIKV infection is required.
As a member of the Flaviviridae family, ZIKV has a single positive sense RNA genome, which is initially translated as a single polyprotein which is then cleaved post-translationally into three structural proteins (C, PrM or M, and E) and seven nonstructural (NS) proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B and NS5). Six of the NS proteins (NS2A to NS5) form a replication complex on the cytoplasmic side of the endoplasmic reticulum membrane. The glycosylated NS1, which associates with lipids, forms a homodimer inside the cells and is necessary for viral replication and late in infection9. NS1 is also secreted into the extracellular space as a hexameric lipoprotein particle9, which is involved in immune evasion and pathogenesis by interacting with components from both the innate and adaptive immune systems, as well as other host factors10,11. NS1 is the major antigenic marker for viral infection12, and it has been suggested, in combination with other markers, as a biomarker for early detection of dengue virus (DENV) infection13. The molecular mechanisms of NS1 are relatively well established for DENV and West Nile virus (WNV)14,15, and the NS1-encoding sequence is suspected to be a major genetic factor underlying the diverse clinical consequences of infections caused by flaviviruses (over 70 members)16,17. However, little is known about the NS1 of ZIKV, which displays different pathogenesis from that of typical flaviviruses.
To address this lack of information, we expressed the ZIKV NS1172–352 fragment of the BeH819015 strain isolated from Brazil in 2015 in Escherichia coli as inclusion bodies and obtained the soluble protein by in vitro refolding (Online Methods). We then solved its crystal structure by molecular replacement to a resolution of 2.2 Å (Rwork and Rfree values of 0.205 and 0.246, respectively) (Supplementary Table 1). The ZIKV NS1172–352 protein crystallized as a rod-like homodimer with a length of ~9 nm (Fig. 1a). Sedimentation velocity analytical ultracentrifugation analyses confirmed that the ZIKV NS1172–352 protein exists as a homodimer (~40 kDa) in solution (Supplementary Fig. 1).
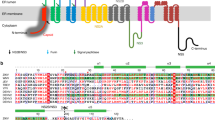
(a) ZIKV NS1172–352 forms a head-to-head dimer with one extended β-ladder platform and one loop surface on the opposite side. (b) Topology diagram for NS1172–352. Glycosylation sites are indicated with green hexagons, and disulfide bonds are indicated with yellow circles. η represents the 310 helix, and β represents the β-sheet.
The ZIKV NS1172–352 homodimer structure has a continuous β-sheet on one surface, with 20 β-strands arranged like the rungs of a ladder (Fig. 1a), in which each monomer contributes ten rungs to the antiparallel β-ladder. On the opposite side of the homodimer, an irregular surface is formed by a complex arrangement of loop structures (Fig. 1a). Most of those interstrand loops are short, except for a long 'spaghetti loop' between β4 and β5 that lacks secondary structure (Fig. 1a,b). A potential N-linked glycosylation site that is highly conserved in the Flaviviridae family is located in the β3–β4 loop (Fig. 1b).
We built a phylogenetic tree by using the ZIKV NS1 protein sequence and those of ten other flaviviruses—WNV, DENV types 1 to 4 (DENV1–4), Japanese encephalitis virus (JEV), St. Louis encephalitis virus (SLEV), yellow fever virus (YFV), Murray Valley encephalitis virus (MVEV) and tick-borne encephalitis virus (TBEV) (Supplementary Fig. 2). This analysis revealed that these flaviviruses can be classified into four groups on the basis of their NS1 sequences: the DENV group (DENV1–4), the WNV group (WNV, JEV, SLEV and MVEV), the YFV group (YFV and TBEV) and an independent ZIKV group. This result suggests that different flaviviruses may have group-specific NS1 characteristics and that the ZIKV NS1 may have properties that are unique among the four groups.
We compared the ZIKV NS1172–352 structure with the available NS1 structures of DENV1, DENV2 and WNV (Fig. 2). ZIKV NS1172–352 is structurally similar to the aforementioned NS1 structures (Fig. 2a), despite having protein sequence identities in the range of 53–56%. Interestingly, the electrostatic surface potential maps of these proteins show several group-specific features in the loop surface (Fig. 2b). The DENV1 and DENV2 NS1 structures both display a positively charged surface in the central regions of their loop surfaces, whereas the WNV NS1 structure has a negatively charged central region in its loop surface. For ZIKV, the loop surface exhibits a composite platform containing both a positively and negatively charged central region and a negative charge toward the two distal ends.
(a) Superimposed structures in ribbon representation of NS1 from ZIKV (orange), dengue virus type 1 (DENV1, blue), dengue virus type 2 (DENV2, green) and West Nile virus (WNV, magenta). (b) Electrostatic surface views of the same four NS1172–352 structures, showing common features on the β-ladder face but diverse characteristics on the loop surface.
The similarities between the NS1 structures of ZIKV and DENV2 allowed us to create a ZIKV NS1 hexamer model (Supplementary Data Set 1) based on the DENV2 NS1 hexamer structure (PDB 4O6B)14. The ZIKV NS1 model shows a hexameric arrangement of three dimers forming a symmetric barrel shape (Fig. 3a). In the center of the barrel, a hydrophobic hole could easily accommodate host lipids to form a complex lipoprotein. In this model, the loop surface of NS1172–352 is fully exposed, facing outward, while the β-ladder surface faces inward (Fig. 3b).
(a) The ZIKV NS1 hexamer, modeled by using the full-length DENV2 NS1 structure (PDB 4O6B)14. The actual NS1172–352 structure is colored in orange, and the predicted NS11–171 structure is colored in light gray. NS1172–352 is located in the outer layer of the hexamer barrel. (b) Electrostatic presentation of the NS1172–352 structure in the hexamer model.
The loop surface has been suggested to play a crucial role in interactions of secreted NS1 hexamers with host factors and antibodies15,18. Furthermore, antibodies to DENV NS1 have been implicated in immune pathogenesis16,19, and epitopes of antibodies with cross-reactivity to human proteins have been mapped to the conserved tip (residues 311–330) of NS1172–352 (Supplementary Fig. 3). Within this conserved tip, ZIKV NS1 has a negatively charged residue Glu315, which is not present in our sequence alignment with ten other flaviviruses (Supplementary Fig. 3).
In summary, our study describes the crystal structure of a C-terminal fragment of ZIKV NS1. Despite having high structural similarity with other flavivirus NS1 proteins, ZIKV NS1 displays a loop-surface interface with divergent electrostatic potential that may result in altered binding properties to host factors and to known protective antibodies to flavivirus NS1; this possibility should be further studied in the near future. Eliciting such antibodies is considered to be an attractive approach in vaccine development, because they would circumvent the enhancement of infection associated with anti-envelope antibodies in DENV13. The unique surface electrostatic potentials in the three known flavivirus NS1 structures (for DENV, WNV and ZIKV) provide a new direction for future studies of more NS1 structures of flavivirus members with diverse clinical outcomes. The unique surface characteristics of ZIKV NS1 may be exploited in the development of new diagnostic tools for ZIKV infection. These future studies may elucidate a clearer scenario of the role that NS1 plays in flavivirus pathogenesis.
Methods
Gene cloning, protein expression and purification.
Coding-sequence-optimized synthetic DNA encoding the ZIKV NS1 C-terminal region (amino acids 172–352, GenBank AMA12085) was cloned into the pET-21a vector (Novagen) with NdeI and XhoI restriction sites and was transformed into Escherichia coli strain BL21(DE3) for protein expression. The inclusion bodies of recombinant proteins were purified and then refolded as previously described, by gradual dilution of the inclusion bodies in a refolding buffer (100 mM Tris-HCl, 2 mM EDTA, 400 mM L-arginine, 0.5 mM oxidized glutathione and 5 mM reduced glutathione, pH 8.0) at 4 °C. The refolded protein was concentrated and purified by gel-filtration chromatography in a buffer containing 20 mM Tris-HCl and 50 mM NaCl, pH 8.0, on a HiLoad 16/60 Superdex 200 PG column (GE Healthcare) and then was further purified by ion-exchange chromatography with a 6-ml RESOURCE Q column (GE Healthcare). For crystallization, the proteins were further purified by gel-filtration chromatography with a Superdex 75 10/300 GL column (GE Healthcare) with a running buffer of 20 mM Tris-HCl and 50 mM NaCl, pH 8.0, and the collected protein fractions were concentrated to 10 mg/mL with a membrane concentrator with a molecular-weight cutoff of 10 kDa (Millipore).
Crystallization, data collection and structure determination.
The initial screening trials were set up with commercial crystallization kits (Molecular Dimensions) with the sitting-drop vapor-diffusion method. Typically, 1 μL protein (5 or 10 mg/mL in 20 mM Tris, pH 8.0, and 50 mM NaCl) was mixed with 1 μL reservoir solution. The resultant drop was then sealed and equilibrated against 100 μL reservoir solution at 4 or 18 °C. Diffractable crystals were obtained in a reservoir solution containing 0.03 M NaNO3, 0.03 M Na2HPO4, 0.03 M (NH4)2SO4, 0.1 M buffer mix, pH 6.5 (44.5 mM imidazole, pH 10.23, and 55.5 mM MES, pH 3.11), 12.5% MPD, 12.5% PEG 1000 and 12.5% PEG 3350. Crystals were flash-cooled in liquid nitrogen after briefly being soaked in reservoir solution with 17% (v/v) glycerol added. The X-ray diffraction data were collected under cryogenic conditions (100 K) at a wavelength of 0.97915 Å at the Shanghai Synchrotron Radiation Facility (SSRF) beamline BL17U and were indexed, integrated and scaled with HKL2000 (ref. 20).
The ZIKV NS1172–352 structure was solved by the molecular replacement method with Phaser21 from the CCP4 program suite22, with the structure of DENV2 NS1172–352 (PDB 4OIG) as the search model. Initial restrained rigid-body refinement and manual model building were performed with REFMAC5 (ref. 23) and COOT24, respectively. Further refinement was performed with Phenix25. In the final Ramachandran plot, 93.43% and 99.43% of residues were in the favored and allowed regions, respectively. The final statistics for data collection and structure refinement are shown in Supplementary Table 1.
Modeling of ZIKV NS1 hexamer.
The structural model of ZIKV NS11–171 was built with the SWISS-MODEL online server (http://www.swissmodel.expasy.org/). The crystal structure of DENV2 NS1 (PDB 4O6B) was used as a template to build the model. The hexamer structural model of ZIKV NS1 was constructed by superimposing the ZIKV NS11–171 model and the ZIKV NS1172–352 crystal structure with the DENV2 NS1 hexamer structure (PDB 4O6B). The coordinate file for this model is provided as Supplementary Data Set 1.
Biochemical characterization of the protein.
The purified protein was analyzed with an analytical gel-filtration assay with a calibrated Superdex 75 10/300 GL column (GE Healthcare). The samples were further analyzed with SDS–PAGE.
The analytical ultracentrifugation assay was performed according to a previously reported method26. The proteins were prepared in 20 mM Tris, pH 8.0, and 50 mM NaCl at a concentration of A280 = 0.8. The assay was performed on an optimal XL-I analytical ultracentrifuge (Beckman Coulter) at a speed of 60,000 r.p.m.
Accession codes.
Coordinates and structure factors have been deposited in the Protein Data Bank under accession code PDB 5IY3.
References
Dick, G.W., Kitchen, S.F. & Haddow, A.J. Trans. R. Soc. Trop. Med. Hyg. 46, 509–520 (1952).
Faye, O. et al. PLoS Negl. Trop. Dis. 8, e2636 (2014).
Cao-Lormeau, V.M. & Musso, D. Lancet 384, 1571–1572 (2014).
Petersen, E. et al. Int. J. Infect. Dis. 44, 11–15 (2016).
Calvet, G. et al. Lancet Infect. Dis. http://dx.doi.org/10.1016/S1473-3099(16)00095-5 (2016).
Mlakar, J. et al. N. Engl. J. Med. 374, 951–958 (2016).
Tang, H. et al. Cell Stem Cell http://dx.doi.org/10.1016/j.stem.2016.02.016 (2016).
Mansuy, J.M. et al. Lancet Infect. Dis. http://dx.doi.org/10.1016/S1473-3099(16)00138-9 (2016).
Suthar, M.S., Diamond, M.S. & Gale, M. Jr. Nat. Rev. Microbiol. 11, 115–128 (2013).
Avirutnan, P. et al. J. Exp. Med. 207, 793–806 (2010).
Chung, K.M. et al. Proc. Natl. Acad. Sci. USA 103, 19111–19116 (2006).
Young, P.R., Hilditch, P.A., Bletchly, C. & Halloran, W. J. Clin. Microbiol. 38, 1053–1057 (2000).
Muller, D.A. & Young, P.R. Antiviral Res. 98, 192–208 (2013).
Akey, D.L. et al. Science 343, 881–885 (2014).
Edeling, M.A., Diamond, M.S. & Fremont, D.H. Proc. Natl. Acad. Sci. USA 111, 4285–4290 (2014).
Cheng, H.J. et al. Exp. Biol. Med. (Maywood) 234, 63–73 (2009).
Kuno, G., Chang, G.J., Tsuchiya, K.R., Karabatsos, N. & Cropp, C.B. J. Virol. 72, 73–83 (1998).
Avirutnan, P. et al. PLoS Pathog. 3, e183 (2007).
Liu, I.J., Chiu, C.Y., Chen, Y.C. & Wu, H.C. J. Biol. Chem. 286, 9726–9736 (2011).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Read, R.J. Acta Crystallogr. D Biol. Crystallogr. 57, 1373–1382 (2001).
Collaborative Computational Project, Number 4. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P. & Cowtan, K. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Zhang, N. et al. Nat. Commun. 2, 577 (2011).
Acknowledgements
This work was supported by the Emergency Task-Force Project (grant no. 81641001) of the National Natural Science Foundation of China (NSFC), the China Ministry of Science and Technology National 973 Project (grant no. 2015CB910500) and the Task Force of Zika Virus Research from the Chinese Academy of Sciences (CAS) and the Zika Special Project of the National Infectious Disease Control S&T Grand Project and Strategic Priority Research Program of the CAS (XDB08020100). We thank the staff of the BL17U beamline at the Shanghai Synchrotron Radiation Facility for assistance during data collection. We thank X. Xu, H. Wang and Z. Su for assistance in protein preparation. Y.S. is supported by the Excellent Young Scientist Program of the CAS and the Youth Innovation Promotion Association, CAS (2015078). G.F.G. is supported partly as a leading principal investigator of the NSFC Innovative Research Group (grant no. 81321063).
Author information
Authors and Affiliations
Contributions
G.F.G. and Y.S. designed and supervised the study. H.S. conducted the experiments. J.Q. collected the data sets and solved the structures. Y.S., J.H., H.S. and G.F.G. analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Biophysical characterization of the ZIKV NS1172–352 protein
(a) Analytical gel filtration of ZIKV NS1172-352 protein. The 280-nm absorbance curve and the SDS-PAGE migration profile of the pooled sample are shown. (b) Ultracentrifugation sedimentation profile of ZIKV NS1172-352. The calculated molecular weight of the indicated protein species is shown.
Supplementary Figure 2 Phylogenetic analysis of NS1 proteins from different flaviviruses.
We applied a neighbor-joining method on NS1 protein sequences from ZIKV and ten representative flaviviruses. The eleven flaviviruses can be classified into four groups. ZIKV is an independent group, separated from the ten representative flaviviruses.
Accession codes: Zika virus (ZIKV) strain BeH819015, AMA12085; West Nile virus (WNV), Q9Q6P4; Dengue virus 1 Nauru/West Pac/1974(DENV1), P17763; Dengue virus 2 Puerto Rico/PR159-S1/1969 (DENV2), P12823; Dengue virus 3 Singapore/8120/1995 (DENV3), Q5UB51; Dengue virus 4 Thailand/0348/1991 (DENV4), Q2YHF0; Japanese encephalitis strain K94P05 (JEV), AAC02714; St. Louis encephalitis (SLEV), D8L537; Yellow fever 17D vaccine strain (YFV), CAA27332; Murray Valley encephalitis (MVEV), NP_051124; Tick-borne encephalitis (TBEV), NP_043135.
Supplementary Figure 3 Secondary structure and sequence alignment of flavivirus NS1 proteins.
The charged residues that contribute to the electrostatic characters in the loop surface of ZIKV NS1172-352 are indicated by red and blue squares. The region (residues 311 to 330) recognized by autoreactive antibodies is indicated by dot lines. The unique residue of ZIKV in the highly conserved region is indicated by a dark blue arrowhead.
Accession numbers for the NS1 protein sequence are described in the supplementary figure 2 legend.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Table 1 (PDF 827 kb)
Supplementary Data Set 1
Hexamer model of ZIKV NS1 (TXT 1207 kb)
Rights and permissions
About this article
Cite this article
Song, H., Qi, J., Haywood, J. et al. Zika virus NS1 structure reveals diversity of electrostatic surfaces among flaviviruses. Nat Struct Mol Biol 23, 456–458 (2016). https://doi.org/10.1038/nsmb.3213
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3213
This article is cited by
-
Structural characterization of SARS-CoV-2 dimeric ORF9b reveals potential fold-switching trigger mechanism
Science China Life Sciences (2023)
-
TRIM22 suppresses Zika virus replication by targeting NS1 and NS3 for proteasomal degradation
Cell & Bioscience (2022)
-
ZIKV viral proteins and their roles in virus-host interactions
Science China Life Sciences (2021)
-
Association of high pressure and alkaline condition for solubilization of inclusion bodies and refolding of the NS1 protein from zika virus
BMC Biotechnology (2018)
-
Purified Inactivated Zika Vaccine Candidates Afford Protection against Lethal Challenge in Mice
Scientific Reports (2018)