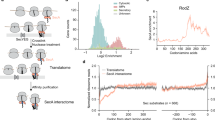
Abstract
Ribosomes synthesizing inner membrane proteins in Escherichia coli are targeted to the membrane by the signal recognition particle (SRP) pathway. By rapid kinetic analysis we show that after initial binding to the ribosome, SRP undergoes dynamic fluctuations in search of additional interactions. Non-translating ribosomes, or ribosomes synthesizing non-membrane proteins, do not provide these contacts, allowing SRPs to dissociate rapidly. A nascent peptide in the exit tunnel stabilizes SRPs in a standby state. Binding to the emerging signal-anchor sequence (SAS) of a nascent membrane protein halts the fluctuations of SRP, resulting in complex stabilization and recruitment of the SRP receptor. We propose a kinetic model where SRP rapidly scans all ribosomes until it encounters a ribosome exposing an SAS. Binding to the SAS switches SRP into the targeting mode, in which dissociation is slow and docking of the SRP receptor is accelerated.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bibi, E. Early targeting events during membrane protein biogenesis in Escherichia coli. Biochim. Biophys. Acta 1808, 841–850 (2011).
Grudnik, P., Bange, G. & Sinning, I. Protein targeting by the signal recognition particle. Biol. Chem. 390, 775–782 (2009).
Luirink, J., von Heijne, G., Houben, E. & de Gier, J.W. Biogenesis of inner membrane proteins in Escherichia coli. Annu. Rev. Microbiol. 59, 329–355 (2005).
Bornemann, T., Jöckel, J., Rodnina, M.V. & Wintermeyer, W. Signal sequence-independent membrane targeting of ribosomes containing short nascent peptides within the exit tunnel. Nat. Struct. Mol. Biol. 15, 494–499 (2008).
Zhang, X., Rashid, R., Wang, K. & Shan, S.O. Sequential checkpoints govern substrate selection during cotranslational protein targeting. Science 328, 757–760 (2010).
Jensen, C.G. & Pedersen, S. Concentrations of 4.5S RNA and Ffh protein in Escherichia coli: the stability of Ffh protein is dependent on the concentration of 4.5S RNA. J. Bacteriol. 176, 7148–7154 (1994).
Gu, S.Q., Peske, F., Wieden, H.J., Rodnina, M.V. & Wintermeyer, W. The signal recognition particle binds to protein L23 at the peptide exit of the Escherichia coli ribosome. RNA 9, 566–573 (2003).
Buskiewicz, I.A., Jockel, J., Rodnina, M.V. & Wintermeyer, W. Conformation of the signal recognition particle in ribosomal targeting complexes. RNA 15, 44–54 (2009).
Halic, M. et al. Signal recognition particle receptor exposes the ribosomal translocon binding site. Science 312, 745–747 (2006).
Zheng, N. & Gierasch, L.M. Domain interactions in E. coli SRP: stabilization of M domain by RNA is required for effective signal sequence modulation of NG domain. Mol. Cell 1, 79–87 (1997).
Bradshaw, N., Neher, S.B., Booth, D.S. & Walter, P. Signal sequences activate the catalytic switch of SRP RNA. Science 323, 127–130 (2009).
Jagath, J.R., Rodnina, M.V., Lentzen, G. & Wintermeyer, W. Interaction of guanine nucleotides with the signal recognition particle from Escherichia coli. Biochemistry 37, 15408–15413 (1998).
Peluso, P. et al. Role of 4.5S RNA in assembly of the bacterial signal recognition particle with its receptor. Science 288, 1640–1643 (2000).
Schaffitzel, C. & Ban, N. Generation of ribosome nascent chain complexes for structural and functional studies. J. Struct. Biol. 158, 463–471 (2007).
Bhushan, S. et al. SecM-stalled ribosomes adopt an altered geometry at the peptidyl transferase center. PLoS Biol. 9, e1000581 (2011).
Saraogi, I., Zhang, D., Chandrasekaran, S. & Shan, S.O. Site-specific fluorescent labeling of nascent proteins on the translating ribosome. J. Am. Chem. Soc. 133, 14936–14939 (2011).
Janda, C.Y. et al. Recognition of a signal peptide by the signal recognition particle. Nature 465, 507–510 (2010).
Hainzl, T., Huang, S., Merilainen, G., Brannstrom, K. & Sauer-Eriksson, A.E. Structural basis of signal-sequence recognition by the signal recognition particle. Nat. Struct. Mol. Biol. 18, 389–391 (2011).
Egea, P.F. et al. Substrate twinning activates the signal recognition particle and its receptor. Nature 427, 215–221 (2004).
Focia, P.J., Shepotinovskaya, I.V., Seidler, J.A. & Freymann, D.M. Heterodimeric GTPase core of the SRP targeting complex. Science 303, 373–377 (2004).
Estrozi, L.F., Boehringer, D., Shan, S.O., Ban, N. & Schaffitzel, C. Cryo-EM structure of the E. coli translating ribosome in complex with SRP and its receptor. Nat. Struct. Mol. Biol. 18, 88–90 (2011).
Mircheva, M. et al. Predominant membrane localization is an essential feature of the bacterial signal recognition particle receptor. BMC Biol. 7, 76 (2009).
Braig, D. et al. Signal sequence-independent SRP-SR complex formation at the membrane suggests an alternative targeting pathway within the SRP cycle. Mol. Biol. Cell 22, 2309–2323 (2011).
Stjepanovic, G. et al. Lipids trigger a conformational switch that regulates signal recognition particle (SRP)-mediated protein targeting. J. Biol. Chem. 286, 23489–23497 (2011).
Zhang, X., Schaffitzel, C., Ban, N. & Shan, S.O. Multiple conformational switches in a GTPase complex control co-translational protein targeting. Proc. Natl. Acad. Sci. USA 106, 1754–1759 (2009).
Shen, K., Zhang, X. & Shan, S.O. Synergistic actions between the SRP RNA and translating ribosome allow efficient delivery of the correct cargos during cotranslational protein targeting. RNA 17, 892–902 (2011).
Zhang, X., Kung, S. & Shan, S.O. Demonstration of a multistep mechanism for assembly of the SRP x SRP receptor complex: implications for the catalytic role of SRP RNA. J. Mol. Biol. 381, 581–593 (2008).
Buskiewicz, I., Kubarenko, A., Peske, F., Rodnina, M.V. & Wintermeyer, W. Domain rearrangement of SRP protein Ffh upon binding 4.5S RNA and the SRP receptor FtsY. RNA 11, 947–957 (2005).
Buskiewicz, I. et al. Conformations of the signal recognition particle protein Ffh from Escherichia coli as determined by FRET. J. Mol. Biol. 351, 417–430 (2005).
Kramer, G. et al. L23 protein functions as a chaperone docking site on the ribosome. Nature 419, 171–174 (2002).
Rodnina, M.V. & Wintermeyer, W. GTP consumption of elongation factor Tu during translation of heteropolymeric mRNAs. Proc. Natl. Acad. Sci. USA 92, 1945–1949 (1995).
Milon, P. et al. Transient kinetics, fluorescence, and FRET in studies of initiation of translation in bacteria. Methods Enzymol. 430, 1–30 (2007).
Acknowledgements
We thank E. Deuerling (University of Konstanz, Germany) for the E. coli strain lacking ribosomal protein L23 and A. Bursy, F. Hummel, T. Wiles, S. Kappler and O. Geintzer for expert technical assistance. The work was supported by the Deutsche Forschungsgemeinschaft (grant WI 626/18-1 to W.W.).
Author information
Authors and Affiliations
Contributions
W.H., T.B., M.V.R. and W.W. conceived the research and designed experiments. W.H. and S.L. prepared materials and conducted experiments. W.H., S.L., T.S. and M.V.R. analyzed the data. W.W. and M.V.R wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Note (PDF 1414 kb)
Rights and permissions
About this article
Cite this article
Holtkamp, W., Lee, S., Bornemann, T. et al. Dynamic switch of the signal recognition particle from scanning to targeting. Nat Struct Mol Biol 19, 1332–1337 (2012). https://doi.org/10.1038/nsmb.2421
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2421
This article is cited by
-
Ribosome-bound Get4/5 facilitates the capture of tail-anchored proteins by Sgt2 in yeast
Nature Communications (2021)
-
Kinetic control of nascent protein biogenesis by peptide deformylase
Scientific Reports (2021)
-
Cotranslational protein targeting to the membrane: Nascent-chain transfer in a quaternary complex formed at the translocon
Scientific Reports (2018)
-
The signal recognition particle contacts uL23 and scans substrate translation inside the ribosomal tunnel
Nature Microbiology (2017)
-
Signal recognition particle prevents N-terminal processing of bacterial membrane proteins
Nature Communications (2017)