Abstract
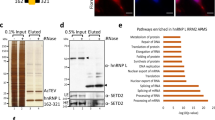
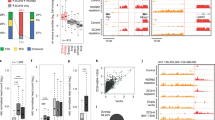
Argonaute proteins play a major part in transcriptional gene silencing in many organisms, but their role in the nucleus of somatic mammalian cells remains elusive. Here, we have immunopurified human Argonaute-1 and Argonaute-2 (AGO1 and AGO2) chromatin-embedded proteins and found them associated with chromatin modifiers and, notably, with splicing factors. Using the CD44 gene as a model, we show that AGO1 and AGO2 facilitate spliceosome recruitment and modulate RNA polymerase II elongation rate, thereby affecting alternative splicing. Proper AGO1 and AGO2 recruitment to CD44 transcribed regions required the endonuclease Dicer and the chromobox protein HP1γ, and resulted in increased histone H3 lysine 9 methylation on variant exons. Our data thus uncover a new model for the regulation of alternative splicing, in which Argonaute proteins couple RNA polymerase II elongation to chromatin modification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Filipowicz, W., Jaskiewicz, L., Kolb, F.A. & Pillai, R.S. Post-transcriptional gene silencing by siRNAs and miRNAs. Curr. Opin. Struct. Biol. 15, 331–341 (2005).
Fabian, M.R., Sonenberg, N. & Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 79, 351–379 (2010).
Grishok, A., Sinskey, J.L. & Sharp, P.A. Transcriptional silencing of a transgene by RNAi in the soma of C. elegans. Genes Dev. 19, 683–696 (2005).
Martienssen, R.A., Zaratiegui, M. & Goto, D.B. RNA interference and heterochromatin in the fission yeast Schizosaccharomyces pombe. Trends Genet. 21, 450–456 (2005).
Matzke, M.A. & Birchler, J.A. RNAi-mediated pathways in the nucleus. Nat. Rev. Genet. 6, 24–35 (2005).
Fagegaltier, D. et al. The endogenous siRNA pathway is involved in heterochromatin formation in Drosophila. Proc. Natl. Acad. Sci. USA 106, 21258–21263 (2009).
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004).
Zaratiegui, M. et al. RNAi promotes heterochromatic silencing through replication-coupled release of RNA Pol II. Nature 479, 135–138 (2011).
Reuter, M. et al. Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature 480, 264–267 (2011).
Weinmann, L. et al. Importin 8 is a gene-silencing factor that targets Argonaute proteins to distinct mRNAs. Cell 136, 496–507 (2009).
Benhamed, M., Herbig, U., Ye, T., Dejean, A. & Bischof, O. Senescence is an endogenous trigger for microRNA-directed transcriptional gene silencing in human cells. Nat. Cell Biol. 14, 266–275 (2012).
Kim, D.H., Villeneuve, L.M., Morris, K.V. & Rossi, J.J. Argonaute-1 directs siRNA-mediated transcriptional gene silencing in human cells. Nat. Struct. Mol. Biol. 13, 793–797 (2006).
Morris, K.V., Chan, S.W., Jacobsen, S.E. & Looney, D.J. Small interfering RNA–induced transcriptional gene silencing in human cells. Science 305, 1289–1292 (2004).
Wang, E.T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Cooper, T.A., Wan, L. & Dreyfuss, G. RNA and disease. Cell 136, 777–793 (2009).
Barash, Y. et al. Deciphering the splicing code. Nature 465, 53–59 (2010).
Allemand, E., Batsche, E. & Muchardt, C. Splicing, transcription, and chromatin: a ménage à trois. Curr. Opin. Genet. Dev. 18, 145–151 (2008).
Luco, R.F., Allo, M., Schor, I.E., Kornblihtt, A.R. & Misteli, T. Epigenetics in alternative pre-mRNA splicing. Cell 144, 16–26 (2011).
Brodsky, A.S. et al. Genomic mapping of RNA polymerase II reveals sites of cotranscriptional regulation in human cells. Genome Biol. 6, R64 (2005).
de la Mata, M., Lafaille, C. & Kornblihtt, A.R. First come, first served revisited: factors affecting the same alternative-splicing event have different effects on the relative rates of intron removal. RNA 16, 904–912 (2010).
Sims, R.J. III et al. Recognition of trimethylated histone H3 lysine 4 facilitates the recruitment of transcription postinitiation factors and pre-mRNA splicing. Mol. Cell 28, 665–676 (2007).
Luco, R.F. et al. Regulation of alternative splicing by histone modifications. Science 327, 996–1000 (2010).
Batsché, E., Yaniv, M. & Muchardt, C. The human SWI/SNF subunit Brm is a regulator of alternative splicing. Nat. Struct. Mol. Biol. 13, 22–29 (2006).
Saint-André, V., Batsche, E., Rachez, C. & Muchardt, C. Histone H3 lysine 9 trimethylation and HP1γ favor inclusion of alternative exons. Nat. Struct. Mol. Biol. 18, 337–344 (2011).
Alló, M. et al. Control of alternative splicing through siRNA-mediated transcriptional gene silencing. Nat. Struct. Mol. Biol. 16, 717–724 (2009).
Chu, C.Y. & Rana, T.M. Translation repression in human cells by microRNA-induced gene silencing requires RCK/p54. PLoS Biol. 4, e210 (2006).
Meister, G. et al. Identification of novel Argonaute-associated proteins. Curr. Biol. 15, 2149–2155 (2005).
Landthaler, M. et al. Molecular characterization of human Argonaute-containing ribonucleoprotein complexes and their bound target mRNAs. RNA 14, 2580–2596 (2008).
Höck, J. et al. Proteomic and functional analysis of Argonaute-containing mRNA-protein complexes in human cells. EMBO Rep. 8, 1052–1060 (2007).
Nielsen, S.J. et al. Rb targets histone H3 methylation and HP1 to promoters. Nature 412, 561–565 (2001).
Cammas, F., Herzog, M., Lerouge, T., Chambon, P. & Losson, R. Association of the transcriptional co-repressor TIF1-β with heterochromatin protein 1 (HP1): an essential role for progression through differentiation. Genes Dev. 18, 2147–2160 (2004).
Will, C.L. & Luhrmann, R. Spliceosomal UsnRNP biogenesis, structure and function. Curr. Opin. Cell Biol. 13, 290–301 (2001).
Long, J.C. & Caceres, J.F. The SR protein family of splicing factors: master regulators of gene expression. Biochem. J. 417, 15–27 (2009).
Taft, R.J. et al. Nuclear-localized tiny RNAs are associated with transcription initiation and splice sites in metazoans. Nat. Struct. Mol. Biol. 17, 1030–1034 (2010).
Valen, E. et al. Biogenic mechanisms and utilization of small RNAs derived from human protein-coding genes. Nat. Struct. Mol. Biol. 18, 1075–1082 (2011).
O'Carroll, D. et al. A Slicer-independent role for Argonaute-2 in hematopoiesis and the microRNA pathway. Genes Dev. 21, 1999–2004 (2007).
Sinkkonen, L., Hugenschmidt, T., Filipowicz, W. & Svoboda, P. Dicer is associated with ribosomal DNA chromatin in mammalian cells. PLoS ONE 5, e12175 (2010).
Otsuka, M. et al. Hypersusceptibility to vesicular stomatitis virus infection in Dicer1-deficient mice is due to impaired miR24 and miR93 expression. Immunity 27, 123–134 (2007).
Volpe, T. & Martienssen, R.A. RNA interference and heterochromatin assembly. Cold Spring Harb. Perspect. Biol. 3, a003731 (2011).
Schalch, T. et al. High-affinity binding of Chp1 chromodomain to K9 methylated histone H3 is required to establish centromeric heterochromatin. Mol. Cell 34, 36–46 (2009).
Moshkovich, N. et al. RNAi-independent role for Argonaute-2 in CTCF/CP190 chromatin insulator function. Genes Dev. 25, 1686–1701 (2011).
Leung, A.K.L. et al. Genome-wide identification of Ago2 binding sites from mouse embryonic stem cells with and without mature microRNAs. Nat. Struct. Mol. Biol. 18, 237–244 (2011).
Wang, E., Dimova, N. & Cambi, F. PLP/DM20 ratio is regulated by hnRNPH and F and a novel G-rich enhancer in oligodendrocytes. Nucleic Acids Res. 35, 4164–4178 (2007).
Enerly, E., Sheng, Z. & Li, K.B. Natural antisense as potential regulator of alternative initiation, splicing and termination. In Silico Biol. 5, 367–377 (2005).
RIKEN Genome Exploration Research Group and Genome Science Group (Genome Network Project Core Group) and the FANTOM Consortium et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005).
Guang, S. et al. Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription. Nature 465, 1097–1101 (2010).
Cernilogar, F.M. et al. Chromatin-associated RNA interference components contribute to transcriptional regulation in Drosophila. Nature 480, 391–395 (2011).
Nonne, N., Ameyar-Zazoua, M., Souidi, M. & Harel-Bellan, A. Tandem affinity purification of miRNA target mRNAs (TAP-Tar). Nucleic Acids Res. 38, e20 (2010).
Ouararhni, K. et al. The histone variant mH2A1.1 interferes with transcription by downregulating PARP-1 enzymatic activity. Genes Dev. 20, 3324–3336 (2006).
Fritsch, L. et al. A subset of the histone H3 lysine 9 methyltransferases Suv39h1, G9a, GLP, and SETDB1 participate in a multimeric complex. Mol. Cell 37, 46–56 (2010).
Berninger, P., Gaidatzis, D., van Nimwegen, E. & Zavolan, M. Computational analysis of small-RNA cloning data. Methods 44, 13–21 (2008).
Acknowledgements
The authors thank A. Krainer (Cold Spring Harbor Laboratory) for the antibody to SRSF1 (anti-SRSF1) G. Meister (Regensburg University) for anti-AGO1 no. 4B8 and anti-AGO2 no. 11A9, Z. Mourelatos (University of Pennsylvania) for anti-AGO2 no. 2A8, A. Tarakhovsky (Rockefeller University), G. Hannon (Cold Spring Harbor Laboratory) and M. Otsuka (University of Tokyo) for the kind gift of knockout MEFs, A. Polesskaya (Centre National de la Recherche Scientifique) for generating the tagged AGO2 C2C12 cells, M. Zavolan and M. Khorshid (Basel University) for their help with bioinformatics, the Taplin Biological Mass Spectrometry Facility at Harvard Medical School for MS analysis, P. de la Grange (GenoSplice) for help with bioinformatics and J.B. Weitzman (University Paris Diderot), E. Allemand (Institut Pasteur) and L. Pritchard (Centre National de la Recherche Scientifique) for critical reading of the manuscript. This work was supported by the European Commission Sixth Framework Programme (Integrated Project Silencing RNAs: Organisers and Coordinators of Complexity in Eukaryotic Organisms (SIROCCO) contract number LSHG-CT-2006-037900, to A.H.-B.) and by the Agence Nationale de la Recherche (contract number ANR-11-BSV8-0013 to C.M., J.-C.A. and A.H.-B.).
Author information
Authors and Affiliations
Contributions
M.A.-Z. and M.S. conducted the biochemical characterization of AGO complexes; P.R. participated in mass-spectrometry analysis; M.A.-Z. and E.B. conducted functional splicing analyses and ChIP experiments; M.A.-Z. E.B. and C.R. conducted RNA-nChIP experiments; R.Y., E.B., N.M., R.F., N.D. and J.-C.A. analyzed RNA deep-sequencing data; E.B. analyzed exon arrays; J.M. conducted some of the RT-qPCRs; L.F. and S.A.-S.-A. contributed the analysis of AGO-H3K9 KMT interaction; A.H. proposed the tandem affinity purification–tagging procedure on chromatin and participated in the design of that part of the study; E.B. and C.M. initiated and supervised splicing analyses; A.H.-B. initiated and supervised AGO complex characterization; C.R. and S.A.-S.-A. participated in writing the paper; M.A.-Z., E.B., C.M. and A.H.-B. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–6 (PDF 10743 kb)
Rights and permissions
About this article
Cite this article
Ameyar-Zazoua, M., Rachez, C., Souidi, M. et al. Argonaute proteins couple chromatin silencing to alternative splicing. Nat Struct Mol Biol 19, 998–1004 (2012). https://doi.org/10.1038/nsmb.2373
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2373
This article is cited by
-
Aedes aegypti Argonaute 2 controls arbovirus infection and host mortality
Nature Communications (2023)
-
PSMC3 promotes RNAi by maintaining AGO2 stability through USP14
Cellular & Molecular Biology Letters (2022)
-
A nuclear function for an oncogenic microRNA as a modulator of snRNA and splicing
Molecular Cancer (2022)
-
The Heterochromatin protein 1 is a regulator in RNA splicing precision deficient in ulcerative colitis
Nature Communications (2022)
-
The SWI/SNF subunit BRG1 affects alternative splicing by changing RNA binding factor interactions with nascent RNA
Molecular Genetics and Genomics (2022)