Key Points
-
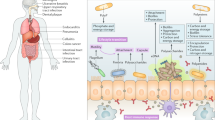
Bacteria are capable of producing a vast diversity of polymers serving a range of biological functions, such as reserve material, protective capsules and slimes, and biofilm matrix components. Several of these polymers have chemical and material properties that are suitable for industrial and medical applications, and many are therefore commercially produced, providing a valuable source for renewable, biodegradable and biocompatible materials.
-
Bacterial polymers can be categorized into polysaccharides, polyamides, polyesters and the inorganic polyanhydrides. Although biosynthesis of the various activated precursors (for example, nucleoside diphosphate sugars and hydroxyacyl-CoA thioesters) is well understood, the molecular mechanisms of polymerisation and, when relevant, secretion remain elusive.
-
A central goal in the field is to obtain a deeper understanding of the operation and three-dimensional architecture of polymerases and synthases, or the respective polymerisation and secretion multiprotein complexes, in order to increase the efficiency of engineering experiments. Random mutagenesis approaches have already proved to be successful, generating biosynthesis enzymes with a change in substrate specificity and activity and enabling the efficient production of modified polymers.
-
The increasing knowledge with respect to biosynthesis pathways and key biosynthesis enzymes has enabled metabolic engineering and protein engineering, leading to the production of modified and new polymers with unique and valuable material properties. These engineering approaches are informed by recent advances in the analysis of the polymer structure–material property relationship.
-
Aside from the possibility of easily engineering bacterial production hosts for the production of modified polymers, in vitro enzymatic and chemical modifications offer additional design space to the manufacture of bacterial polymers. The purification of polymerases and synthases or subcellular fractions containing the respective enzyme activities from bacteria allows controlled processive in vitro synthesis of defined polymers.
-
Whereas several bacterial exopolysaccharides and polyhydroxyalkanoates are already commercially produced, the polyamides and polyphosphates are not commercially produced by bacterial fermentation. However, recent advances in polyamide biosynthesis hold promise for future commercial synthesis.
-
The economics of the bacterial production of polymers is largely driven by the cost of the precursor substrate, the productivity of the host organisms, the upstream and downstream processing costs and the application value of the polymer. In addition, when bacterial polymers such as the polyhydroxyalkanoates compete with oil-based polymers, crude oil prices substantially affect polymer economics.
-
Notable progress has been made in all of the areas mentioned above, and exploitation of metabolic engineering in particular has led to the generation of exciting new production hosts able to synthesize even unnatural polymers such as polythioesters and polylactic acid. An interesting new development considers the intracellularly formed spherical polyhydroxyalkanoate granules as versatile biobeads, the surface proteins of which can be engineered to display a range of protein functions for applications in bioseparation, protein production, biocatalysis, diagnostics and antigen delivery.
Abstract
Bacteria can synthesize a wide range of biopolymers that serve diverse biological functions and have material properties suitable for numerous industrial and medical applications. A better understanding of the fundamental processes involved in polymer biosynthesis and the regulation of these processes has created the foundation for metabolic- and protein-engineering approaches to improve economic-production efficiency and to produce tailor-made polymers with highly applicable material properties. Here, I summarize the key aspects of bacterial biopolymer production and highlight how a better understanding of polymer biosynthesis and material properties can lead to increased use of bacterial biopolymers as valuable renewable products.
Similar content being viewed by others
Main
Recent discoveries in the field of bacterial polymer biosynthesis have opened up new avenues for the rational engineering of bacteria towards the production of tailor-made biopolymers suitable for industrial and medical applications. Over the past decade, a better understanding of the molecular mechanisms and regulatory processes underlying the synthesis of biopolymers has emerged. This knowledge has provided us with powerful tools to engineer bacteria that are capable of not only efficient biopolymer production but also the production of modified and even unnatural polymers exhibiting unique material properties for specific high-value applications, all at a viable economic cost. Production at a viable economic cost refers to the market value of the respective biopolymer substantially exceeding the production costs in order to generate profit. The market value depends on the material properties that impact on the field of application. However, low-value applications of biopolymers become economically viable when they can be produced at low costs.
Bacteria efficiently convert different carbon sources into a diverse range of polymers with varying chemical and material properties (Box 1; Fig. 1). Although bacteria synthesize only a few intracellular polymers, the range of extracellular polymers that they can synthesize is vast. Some of these polymers serve the same function in a wide range of prokaryotes1, whereas other polymers can be specific for certain bacterial taxa and serve distinct biological functions2. Many bacterial species are able to synthesize several polymers3,4.
a | Glycogen, an intracellular polysaccharide. b | Xanthan, a secreted polysaccharide. c | K30 antigen, a capsular polysaccharide. d | Alginate, a secreted and/or cyst cell wall polysaccharide. e | Dextran, an extracellularly synthesized polysaccharide. f | Cyanophycin, an intracellular polyamide. g | Poly-γ-glutamate, a secreted polyamide. h | An intracellular polyhydroxyalkanoate. i | Polyphosphate, an intracellular polyanhydride. Numbers below the chemical structure brackets indicate the polymerization degree most commonly found for each polymer; note that the polymerization degree can vary substantially under different growth conditions and in different species.
Four major classes of polymers are produced by bacteria: polysaccharides, polyesters, polyamides and inorganic polyanhydrides (such as polyphosphates). These polymers serve various biological functions, for example as reserve material or as part of a protective structure, and can provide a substantial advantage for bacteria under certain environmental conditions. Therefore, complex regulatory pathways exist to control the biosynthesis and even the material properties of these polymers in response to external stimuli.
Several bacterial polymers are already produced commercially through medium- to large-scale fermentations, with annual world production volumes of around 2,000 tonnes and 100,000 tonnes for the polysaccharides dextran5 and xanthan (D. Seisun, personal communication), respectively, and up to 100,000 tonnes for the polyesters6 (Table 1). When bacterial polymers are required to compete with oil-based non-renewable polymers, the cost of production is a crucial parameter. However, biopolymers derived from natural resources have a competitive advantage, owing to their sustainable production from renewable resources, their biodegradability and, often, their biocompatibility. Biopolymers are, by definition, biodegradable, and so their application as commodity products becomes increasingly attractive in view of the desire to avoid the use of recalcitrant oil-based polymers that will accumulate in the environment. When exposed to the microbial flora present in a given environment (for example, in soil or water), biopolymers are fully degraded and mineralized to CO2 and H2O. Secreted depolymerases and hydrolases attack the biopolymer backbone, leading to lower-molecular-mass degradation products, which can then be taken up by the microbial cell to be used as carbon and energy sources. Furthermore, biopolymers are composed of natural non-toxic constituents and are considered to be inherently biocompatible. This biocompatibility has been harnessed in numerous medical applications, in which biopolymers have been used as scaffolds or matrices in tissue engineering, wound dressing and drug delivery. Some biopolymers are gradually degraded in vivo, making them well suited for use in tissue replacement and controlled drug release.
This Review highlights recent advances in our understanding of in vivo polymer biosynthesis pathways and then considers how this knowledge has enabled the design and metabolic engineering of biopolymers with specific material properties for production on a commercial scale. Owing to the ever-growing number of bacterial biopolymers that have been characterized, detailed descriptions are limited to only representative model polymers. These model polymers have been selected on the basis of the characteristics of their biosynthesis pathways, the extent of knowledge available to describe their formation, and their commercial relevance and applied potential.
Polysaccharides
Given the overwhelming diversity of bacterial polysaccharides, this section focuses on only commercially relevant examples of this class of polymer. The polysaccharides produced by bacteria can be subdivided into the exopolysaccharides (for example, xanthan, dextran, alginate, cellulose, hyaluronic acid (HA) and colanic acid), which can be either secreted or synthesized extracellularly by cell wall-anchored enzymes, the capsular polysaccharides (for example, the K30 antigen) and the intracellular polysaccharide (glycogen). Structural cell wall polysaccharides have been recently discussed elsewhere7 and will not be considered in this Review. Further categorization divides the polysaccharides into repeat unit polymers (for example, xanthan and the K30 antigen), repeating polymers (for example, cellulose) and non-repeating polymers (for example, alginate)8,9,10,11 (Table 1). The formation of polysaccharides with such varied structures and compositions requires the recruitment of different enzymes and proteins, which is reflected in the varied organizations of the biosynthesis gene clusters (Fig. 2).
Triangles represent promoters; triangles labelled P are the key regulatory promoters for polymer biosynthesis. Single-letter gene names take the operon name, such that D in the gum operon represents gumD. Precursor biosynthesis genes that are outside of the main gene clusters are not shown for the gum, kps, alg and pgs operons. Question marks indicate genes with unconfirmed functions. Region 2 is a central group of N-acetylneuraminic-acid biosynthesis genes. β-glu, β-glucosidase; asd, aspartate semi-aldehyde dehydrogenase; gntU, gluconate uptake; orf, open reading frame encoding a protein of unknown function; wzb, protein tyrosine phosphatase.
The exopolysaccharide and capsular-polysaccharide biosynthesis gene clusters are subject to extensive transcriptional regulation involving two-component signal transduction pathways, quorum sensing, alternative RNA polymerase σ-factors and anti-σ-factors, as well as integration host factor (IHF)-dependent and cyclic di-GMP-dependent processes12,13,14,15,16,17. Induction of exopolysaccharide biosynthesis is often correlated with establishment of the biofilm growth mode, during which exopolysaccharides are important matrix components18.
Nucleoside diphosphate sugars (such as ADP–glucose), nucleoside diphosphate sugar acids (such as GDP–mannuronic acid) and nucleoside diphosphate sugar derivatives (such as UDP–N-acetyl glucosamine) are direct precursors for bacterial polysaccharide biosynthesis (Fig. 3; Table 1). Polymer-specific biosynthesis enzymes (for example, pyrophosphorylases and dehydrogenases) are required for synthesis of the activated polymer precursor, which is the first committed biosynthesis step, and have been targeted by metabolic engineering to enhance polymer production and to allow the synthesis of tailor-made polysaccharides19.
The major metabolic routes towards the synthesis of the various polymer precursors are summarized. Solid lines indicate either linking of primary metabolic pathways with intermediates of polymer biosynthesis or direct enzyme-catalysed conversions towards the immediate polymer precursor. Dashed lines indicate multiple enzymatic steps. Polysaccharides are shown in light green boxes, polyesters are shown in pink boxes, polyamides are shown in blue boxes, and the inorganic polyanhydride is shown in a purple box. FAB, fatty acid de novo biosynthesis; FAD, fatty acid β-oxidation; KDPG, the 2-keto-3-deoxy-6-phosphogluconate pathway; NDP, nucleoside 5′-diphosphate; P, phosphate; PGI, phosphoglucoisomerase; PGM, phosphoglucomutase; Pha, polyhydroxyalkanoate synthesis enzyme; PHAscl, short-chain-length PHAs; PHAmcl, medium-chain-length PHAs; TCA cycle, tricarboxylic acid cycle.
Polymerization and secretion of exopolysaccharides and capsular polysaccharides are often rate limiting and can substantially affect the flux of carbon towards the processive formation of high-molecular-mass exopolymers. The synthases or catalytic subunits of synthases are mostly localized in the cytoplasmic membrane and are often associated with proteins that are required for translocation of the polymer across the cytoplasmic membrane and, if needed, the outer membrane (Fig. 4).
The biosynthesis and secretion (where relevant) of many bacterial polymers requires the assembly of multiprotein complexes. Some representative and advanced multiprotein complex models are depicted here. It should be noted that these model are supported by experimental data, but the extent of supporting data varies substantially from polymer to polymer. Protein colours indicate proteins with similar functions. See main text for details. a | The alginate biosynthesis model from Pseudomonas aeruginosa. b | A model of the biosynthesis of the K30 capsular polysaccharide from Escherichia coli. c | The poly-γ-glutamate (PGA) synthesis pathways from Bacillus spp. A, B, C, and E represent either CapA, CapB, CapC and CapE for the synthesis of capsular PGA in Bacillus anthracis or PgsA (also known as PgsAA and CapA), PgsB (also known as CapB), PgsC (also known as CapC) and PgsE for the synthesis of released extracellular PGA in Bacillus licheniformis and Bacillus subtilis. d | The structure of polyhydroxyalkanoate (PHA) granules from Ralstonia eutropha. AlgT, RNA polymerase factor σ22; c-di-GMP, cyclic di-GMP; Pi, inorganic phosphate; Wzb, protein tyrosine phosphatase.
Exopolysaccharides. Exopolysaccharides are produced by a wide range of bacteria and some archaea20. Depending on their subunit composition, structure and molecular mass, exopolysaccharides can have commercially relevant material properties that are attractive for industrial and medical applications. These properties range from forming viscous solutions to exhibiting a pseudoplastic material nature (Table 1). Dextran and xanthan, which are commercially produced, and alginate, another potentially applicable exopolysaccharide, are described in more detail below; owing to similarities between the biosynthesis pathways for xanthan and capsular polysaccharides, the xanthan biosynthesis pathway is discussed in the section concerning capsular polysaccharides.
Dextrans are soluble in water, and dextran solutions behave as Newtonian fluids and have a viscosity that changes as a function of concentration, temperature and average molecular mass21. Native dextrans are poly-disperse (that is, their molecular masses typically range from 106 to 109 daltons). Acid hydrolysis of dextrans generates fractions of defined molecular masses. This property, in addition to their low immunogenicity, has led to numerous clinical and pharmaceutical applications; for example, dextrans are used as blood plasma extenders and as chromatography media22.
Dextransucrase is a glucansucrase belonging to the glycoside hydrolase superfamily and is the key enzyme for dextran synthesis. It is secreted and anchored to the cell wall, and it has an average molecular weight of 160 kDa23 (Table 1). Glucansucrases have been extensively studied, leading to a detailed understanding of their catalytic mechanism (reviewed in Refs 23, 24, 25). These enzymes catalyse the hydrolysis of the glycosidic bond in sucrose and the transfer of glucose to the growing reducing end of the covalently linked glucan chain by an insertion mechanism that relies on two separate catalytic sites in the same active site24; the hydrolysis of the glycosidic bond produces the energy required for the glucose transfer reaction. Dextransucrase is encoded by dsrS, the expression of which is induced in the presence of sucrose26.
The production of HA requires only a single protein, HA synthase (HasA), for polymerization and secretion27,28; however, most exopolysaccharides are polymerized and secreted by membrane-spanning multiprotein complexes (Fig. 4). These complex-mediated biosynthesis processes can be subdivided into two general pathways. One pathway is exemplified by the Wzy-dependent polymerization and secretion mechanism used in xanthan biosynthesis, which requires a lipid carrier for transfer of the repeat unit oligosaccharide across the cytoplasmic membrane. The second pathway is independent of both Wzy and a lipid carrier and is proposed for the production of exopolysaccharides such as alginate and cellulose, which are not composed of repeat units (Fig. 4a; Table 1).
Industrial and medical applications of alginates are linked to their stabilizing, viscosifying and gelling properties and their ability to retain water. Although alginates are commercially produced from seaweeds, the ability to genetically engineer alginate-producing bacteria such as Azotobacter vinelandii and Pseudomonas fluorescens to produce tailor-made high-value alginates holds great promise (Figs 5, 6). The in vitro synthesis of alginate requires the presence of an intact envelope, providing evidence that a multiprotein complex spanning the cytoplasmic membrane, the periplasm and the outer membrane is required for coordinated polymerization and secretion26 (Fig. 4a). The roles of the individual proteins in the complex as well as their regulation have been extensively investigated (for reviews, see Refs 28, 29, 30), but the underlying molecular mechanisms of polymerization and secretion remain elusive. One component of the biosynthesis complex, Alg8, has substantial sequence similarities to processive β-glycosyl transferases of the GT2 family and was recently identified as a key membrane protein for the production of alginate, with multiple copies of Alg8 contributing to substantial overproduction of alginate in Pseudomonas aeruginosa 26,29,31,32,33. The material properties of alginates depend mainly on the composition (that is, the molar ratio and sequence of mannuronic acid and guluronic acid residues), degree of acetylation and molecular mass of the molecules, and the enzymes that control these parameters — which include C5 epimerase, lyase and acetyltransferase — have been studied in great detail with respect to their protein properties and enzyme characteristics11,34. These studies have provided a foundation for the development of bacterial strains that produce tailor-made alginates (Fig. 6). For example, it was shown that inactivation of the C5 epimerase activity in P. fluorescens led to the production of the homopolymer polymannuronate35. Further inactivation of genes encoding other alginate-modifying enzymes and the controlled expression of those same genes in trans would provide a molecular toolbox for tailor-made alginate production.
Blue boxes represent metabolic-engineering targets to increase production or to enable the production of tailor-made polymers. a | Production strains with improved yields and/or that are capable of producing modified or tailor-made polymers have been developed for in vivo production of polymers. This approach applies metabolic engineering, implementing our knowledge about polymer biosynthesis pathways, metabolic flux and key enzymes to modify the biosynthesis pathway appropriately. b | In vitro, there are two basic strategies that can be adopted. First, in vitro synthesis using polymerizing enzymes or engineered enzymes exposed to selected substrates can lead to new biopolymers. Second, isolated biopolymers can be upgraded by exposure to enzymatic or chemical modifications. PHA, polyhydroxyalkanoate.
a | The precursor substrate provided to a bacterial culture can substantially affect the composition of the resulting polymer. This has been found to be particularly useful for the production of modified polyhydroxyalkanoates (PHAs) such as poly(3-hydroxybutylate-co-3-hydroxyvalerate) (poly(3HB-co-3HV)), sold under the trade name Biopol (Metabolix). b | The C5 epimerase that introduces G residues into preformed alginates has been used as an example for in vitro enzyme-catalysed modifications and in vivo metabolic engineering towards the production of tailor-made alginates. G, α-L-guluronic acid; M, β-D-mannuronic acid; poly(3HB), poly(3-hydroxybutyrate).
Capsular polysaccharides. Capsular polysaccharides (CPSs) are secreted but remain attached to the cell and often function as major surface antigens and virulence factors. Their biosynthesis and assembly have been extensively studied in Escherichia coli 36, and the protective CPSs produced by pathogenic bacteria such as Klebsiella pneumoniae and Streptococcus pneumoniae have been shown to contribute to the pathogens' ability to evade phagocytosis by macrophages37,38. Commercial applications of CPSs as valuable materials have not yet been developed, but their function as virulence factors has motivated much research into these polymers. An understanding of the biosynthesis process could identify targets for the treatment of infections, and CPSs or their derivatives might serve as potential vaccine candidates. However, as CPSs provide a paradigm for repeat unit polysaccharide biosynthesis, characterization of their biosynthesis pathways has also contributed substantially towards understanding the synthesis of commercially relevant exopolysaccharides such as xanthan and gellan (Fig. 4b). Indeed, several oligosaccharide repeat unit polymers require the lipid carrier undecaprenyl phosphate for translocation of the repeat unit across the cytoplasmic membrane following the Wzy-dependent pathway, and there is a high level of amino acid sequence similarity between the protein subunits that are involved in the polymerization and secretion of these oligosaccharides8,39,40,41 (Fig. 4b). Previous studies on the lipopolysaccharide O antigen polymerization and secretion processes laid the foundation for our current understanding of what is thought to be the most widely distributed Wzy-dependent polysaccharide biosynthesis pathway42.
During the assembly of K30 antigen, a well-studied example for repeat unit polysaccharide biosynthesis, transfer of the sugar phosphate from the respective nucleotide sugar to undecaprenyl phosphate is catalysed by the initiating glycosyl transferase, WbaP, a membrane-anchored polyisoprenyl sugar phosphate transferase8 and an orthologue of GumD (the glycosyl transferase involved in xanthan synthesis) (Fig. 4b). After a series of further sugar transfers to undecaprenyl phosphate, catalysed by monofunctional glycosyl transferases located in or at the inner leaflet of the cytoplasmic membrane, the repeat unit at the cytosolic side of the membrane is completed.
In xanthan biosynthesis, GumK has been identified as one of these monofunctional glycosyl transferases, providing a potential target for rational protein engineering towards the production of tailor-made modified xanthans43.
The polymerization reaction occurs at the periplasmic side of the cytoplasmic membrane and requires transfer of the undecaprenyl phosphate-linked repeat unit across the membrane, mediated by Wzx, a putative polysaccharide-specific transport protein (a so-called 'flippase') that also interacts with WbaP44,45 (Fig. 4b). The integral membrane protein Wzy has been proposed to be the polymerase that catalyses the transfer of the nascent polymer from its undecaprenyl phosphate carrier to the new lipid-linked repeat unit46. Wzc belongs to the polysaccharide co-polymerase (PCP) family of proteins, which not only assist polymerization but also control polymer length and guide the nascent polymer chain through the periplasm to the outer-membrane auxiliary protein Wza47 (Fig. 4b). It was proposed that PCPs share common structural features, such as transmembrane domains and extended periplasmic domains with surface properties that allow not only the assembly of PCPs into oligomers but also a defined interaction with Wzy protomers, which was suggested to mediate control of polymer chain length47. A model was developed suggesting that polysaccharide synthesis occurs by transferring the nascent chain from one Wzy to an adjacent Wzy, with the PCP controlling chain length by affecting assembly of Wzy (that is, chain growth would be terminated in the absence of an adjacent Wzy). Recent X-ray structures of the PCP ferric enterobactin transport protein E (FepE) (Protein Data Bank accession number 3B8M) and of Wza from E. coli, combined with mutational analyses, have contributed towards our understanding of the structural requirements for translocation of nascent polysaccharide from the periplasm through the outer membrane39,40,48.
Storage polysaccharides. Glycogen is the only intracellular storage polysaccharide found in bacteria and archaea. Glycogen synthases belong to the GT-B superfamily of retaining glycosyl transferases, which retain the anomeric stereochemistry of the donor sugar in the resulting polysaccharide. Resolution of the crystal structures of one archaeal and two bacterial glycogen synthases has improved our mechanistic understanding of their activity as retaining glycosyl transferases49,50,51. ADP–glucose and an α-(1,4)-glucan acceptor bind to the enzyme and, through an induced fit, activate it to catalyse transfer of the glucose moiety to the acceptor molecule49. How the α-(1,4)-glucan acceptor (known as the primer) is synthesized remains unknown. Despite this mechanistic understanding, bacterial glycogen has not been considered for commercial applications to date.
Polyamides
The non-ribosomally synthesized polyamides are a distinct group of biopolymers consisting of only two extracellular polyamides, poly-γ-glutamate (PGA) and ɛ-poly-L-lysine (PL), and the intracellular cyanophycin granule peptide (CGP)52 (Fig. 1; Table 1). PL has been shown to exhibit antibacterial properties. PGA can exist in either capsular or released forms; in Bacillus anthracis, capsular PGA is an important virulence factor that has a low immunogenicity and therefore does not trigger a humoral immune response in the infected host. Released PGA might serve as a nitrogen or carbon source and has also been implicated as a water-binding component of the biofilm matrix. The material and chemical properties of PGA and of CGPs with a chemically reduced arginine content resemble the properties of chemically synthesized and extensively applied polyacrylates; for example they can be used as dispersants, antiscalants or superabsorbers52,53,54. Hence, bacterial polyamides could provide renewable, non-toxic and biodegradable alternatives to polyacrylates.
Intracellular polyamides. CGP polymerization is catalysed by CGP synthetase (CphA), and its mobilization (that is, its breakdown for use as a nitrogen and carbon source) is catalysed by cyanophicinase (CphB) (Fig. 2; Table 1). CGP synthesis has been proposed to resemble an amide ligase-dependent reaction55. In vitro synthesis of CGP requires ATP, K+, Mg2+, a CGP primer and a thiol reagent56,57,58, and recent biochemical studies suggest that CphA forms a homodimer and possesses two different substrate- and ATP-binding sites to accommodate the incorporation of both aspartate and arginine residues into the nascent polymer58,59. A lack of structural data for CphA still hampers elucidation of the reaction mechanism of this enzyme.
Released extracellular and capsular polyamides. In addition to being produced by Gram-positive bacteria (Fig. 2) and archaea, PGA was recently found to be produced by the Gram-negative bacterium Fusobacterium nucleatum60,61 (Table 1). The membrane-anchored protein PGA synthase B (PgsB; also known as CapB), which has similarities to the amide ligases, is the catalytic subunit of the PGA synthetase62. PgsB and PgsC (also known as CapC) both have ATPase activity that is increased by the addition of PgsA (also known as CapA and PgsAA), suggesting that all three proteins might constitute the PGA synthetase multiprotein complex in vivo60,63,64 (Fig. 4c). The proposed PGA synthesis reaction is based on the amide ligase mechanism, such that a primer (oligo-(γ-glutamate) or glutamate) is phosphorylated at its carboxyl terminus and one glutamate residue is then added. PgdS, a γ-glutamyl hydrolase, might hydrolyse the nascent PGA, inducing release of the polymer and controlling its molecular mass65. For PGA capsule formation, CapD (a γ-glutamyl transpeptidase) is required to catalyse covalent anchoring of the PGA chain to the cell wall66.
The biosynthesis of the third natural polyamide, PL, which consists of only 25–35 L-lysine residues, has been intensively studied with respect to the reaction mechanism of the single subunit enzyme PL synthetase (Pls). Only recently, Pls was purified from Streptomyces albulus and the respective gene was cloned67. Pls localizes to the cytoplasmic membrane and contains domains characteristic of non-ribosomal peptide synthetases but lacks the conserved condensation or thioesterase domains required for release of the final product67. PL chain-length diversity was proposed to be the result of an inherent property of Pls, which acts iteratively for the growth of the PL chain residing in a proposed slender cavity. A reaction mechanism similar to an amino acid ligase mechanism has been proposed67; however, PL synthesis is clearly distinguishable from PGA and CGP synthesis in that it does not require phosphorylation of the carboxyl terminus of the growing chain. Pls therefore represents a new single-module non-ribosomal peptide synthetase.
Polyesters
Polyhydroxyalkanoates (PHAs; also known as bacterial bioplastics) accumulate as carbon reserve material in response to the availability of excess carbon source when growth is limited owing to starvation of other nutrients such as nitrogen and phosphorus. This accumulation is subject to extensive regulation by biosynthesis genes1,68. PHA is deposited as spherical intracellular inclusions with an amorphous, hydrophobic PHA core that is mainly surrounded by proteins involved in PHA metabolism69,70 (Fig. 4d). PHAs can vary substantially in composition, as there are over 150 known constituents, resulting in an enormous diversity of material properties. PHAs exhibit a crystallinity ranging from 30% to 70% and a melting temperature of 50 °C to 180 °C; these thermoplastic material properties make PHAs commercially relevant as renewable and biodegradable alternatives to oil-based plastics.
Thermoplastic resins represent around two-thirds of the current global production of oil-based commodity materials, amounting to 170 million tonnes per year, with their global usage growing at about 5% per year71. Bacterial bioplastics can be processed into materials that are suitable to replace oil-based materials in many applications, including film, fibres, moulded products, extruded goods, coatings and adhesives. Currently, the production capacity for generating bioplastics through large-scale bacterial fermentation has reached ∼100,000 tonnes per year6. Production costs of bioplastics synthesis are currently 5–10 times the cost of synthesis for plastics derived from petrochemicals, which is a major hindrance for the successful commercialization of bioplastics.
Owing to the broad substrate specificity of PHA synthase (PhaC), any organic molecules containing a carboxyl and a hydroxyl group that can be converted to the respective CoA thioester can, in principle, be incorporated into a high-molecular-mass PHA72. The biosynthesis pathways of the activated PHA precursor, (R)-3-hydroxyacyl-CoA, have been extensively studied and exploited through metabolic engineering, leading to the production of modified PHAs — that is, a range of heteropolymers and homopolymers containing (R)-3-hydroxy fatty acids and/or (R)-4-hydroxy fatty acids with different carbon chain lengths — that show more favourable material properties (for example, melting temperature, glass transition temperature and elongation at break) for industrial and medical applications72,73 (Figs 5, 6).
PHAs can be classified by chain length, with medium-chain-length PHAs (which have constituent C6–C14 chains) being produced mainly by pseudomonads and short-chain-length PHAs (which have constituent C3–C5 chains) being produced by a wide range of bacteria and archaea1.
Biosynthesis of medium-chain-length PHAs recruits PHA-specific enzymes such as (R)-specific enoyl-CoA hydratase (PhaJ) and (R)-3-hydroxyacyl ACP:CoA transacylase (PhaG) to divert intermediates of fatty acid metabolism (such as enoyl-CoA and (R)-3-hydroxyacyl acyl carrier protein (ACP)) towards biosynthesis pathways of precursors74,75 (Fig. 3).
Biosynthesis of short-chain-length PHAs involves a β-ketothiolase-catalysed condensation of two acetyl-CoA monomers (or an acetyl-CoA and a propionyl-CoA monomer), with subsequent (R)-specific reduction being catalysed by acetoacetyl-CoA reductase, leading to formation of (R)-3-hydroxybutyryl-CoA (or (R)-3-hydroxyvaleryl-CoA)76,77,78 (Fig. 3).
PhaC belongs to the α/β-hydrolase fold family of enzymes and catalyses the stereoselective conversion of the activated precursor (R)-3-hydroxyacyl-CoA to polyoxoesters, with the concomitant release of CoA79,80. Initiation of PHA synthesis requires activation of the thiol group of the cysteine residue in the PhaC active site by the conserved histidine in the same active site, enabling a nucleophilic attack on the thioester bond of the (R)-3-hydroxyacyl-CoA substrate, concomitantly releasing CoA and forming a covalent enzyme–substrate intermediate81,82. A conserved PhaC aspartic acid residue presumably activates the hydroxyl group of the bound 3-hydroxy fatty acid, which then attacks the thioester bond between a second hydroxyl fatty acid unit and the active site cysteine of a second PhaC subunit, joining the two fatty acids together. Incoming substrate is then covalently bound to the free cysteine and, after activation of its hydroxyl group, the next nucleophilic attack will extend the polyester chain by another unit. Experimental evidence was obtained for chain termination occurring by transferring most of the polyester chain to a second, surface-exposed amino acid, which hydrolyses the chain83. The remaining primed PhaC then started a new cycle of PHA synthesis. Experimental evidence was obtained showing that an increased PhaC copy number causes a decrease in PHA chain length, suggesting that the amount of PhaC in a host cell has a role in controlling PHA chain length84. The crystal structure of PhaC remains elusive.
Polyanhydrides
Inorganic polyphosphate is the only polyanhydride found in all living cells (Fig. 1; Table 1). In bacteria, polyphosphate can form intracellular storage particles but may also form a membrane-anchored complex with low-molecular-mass polyhydroxybutyrate, facilitating the uptake of DNA and various ions85. Furthermore, it has been shown that polyphosphate affects numerous aspects of bacterial physiology, such as survival during the stationary growth phase, response to stress, motility, quorum sensing, biofilm formation and pathogenicity. Biodegradable polyphosphates are used in various industrial applications ranging from the replacement of asbestos as a flame retardant to their use as flavour enhancers in food. As polyphosphates can be easily produced by dehydration of rock phosphate, commercial production of bacterial polyphosphates is not economically feasible. However, the ability of bacteria to accumulate polyphosphate while removing phosphate from the environment could make bacterial polyphosphate production a useful biproduct of commercial waste water treatment.
The key enzyme for polyphosphate biosynthesis is the highly conserved polyphosphate kinase (PPK). In addition to synthesizing polyphosphate from ATP, PPK catalyses the reverse reaction, phosphorylating ADP to produce ATP. This activity has been used to recycle costly ATP from cheap polyphosphate in ATP-dependent enzymatic synthesis reactions86,87. The reaction mechanism of PPK has been elucidated by obtaining the crystal structure of PPK from E. coli in complex with the inhibitor, β-γ-imidoadenosine-5-phosphate88. The first step in polyphosphate synthesis is the autophosphorylation of a conserved histidine residue. His435 of E. coli PPK presumably acts as a nucleophile, attacking the phosphodiester bond of the γ-phosphate group of ATP, whereas His592 is a general acid catalyst, donating a proton to the oxygen atom between the β-phosphate and the γ-phosphate88,89. The chain elongation reaction mechanism leading to the generation of polyphosphate remains to be elucidated. Recently, a second PPK, PPK2, has been identified in P. aeruginosa; PPK2 catalyses the synthesis of polyphosphate from GTP and prefers GDP to ADP as a phosphate group acceptor90. This GDP kinase activity, leading to GTP formation, substantially affects the biosynthesis of the alginate precursor91 (Table 1). PPK2 homologues have been found to be widely spread among bacteria92.
Tailor-made biopolymers
Genome sequencing, functional genomics and the cloning and characterization of biosynthesis genes have all had a substantial impact on our understanding of biosynthesis pathways in organisms that produce commercially relevant polymers, as well as leading to the discovery of new biopolymer-producing bacteria93,94,95,96. This knowledge has been applied to pathway reconstruction and engineering towards improved production and the synthesis of tailor-made polymers (Fig. 5). In particular, recombinant production of the less complex polymers, such as PHA, CGP, HA and PGA, through the establishment of biosynthesis pathways in non-polymer-producing heterologous hosts has proven to be a powerful development97,98,99,100,101,102,103. One example is the large-scale commercial production of PHA by fermentation of recombinant E. coli6. In addition, E. coli harbouring the polyhydroxybutyrate (PHB) biosynthesis genes from Ralstonia eutropha (and, if relevant, genes from other bacterial species) has been commonly used for the production of PHA composed of (R)-3-hydroxybutyrate and (R)-3-hydroxyvalerate and/or (R)-3-hydroxyhexanoate, and these polymers show material properties (such as increased elasticity and decreased brittleness) that are preferred for various industrial applications68,103.
More recently, metabolic engineering exploiting bacterial biosynthesis pathways led to the production of new unnatural polymers, including polythioesters and lactate-based polyesters, in recombinant E. coli104,105,106. Homopolythioesters were produced from the precursor substrate 3-mercaptoalkanoate by recombinant E. coli harbouring genes encoding phosphotransbutyrylase (Ptb) and butyrate kinase (Buk1) from Clostridium acetobutylicum and the promiscuous PHA synthase from Thiocapsa pfennigii. These new polymers showed unique properties when compared with PHAs and petrochemical-derived polymers104. Direct production of polylactic acid, a bioplastic that is commercially produced in a two-step process comprising the fermentative production of lactic acid and its subsequent chemical polymerization, was achieved using E. coli harbouring the genes encoding engineered propionate-CoA transferase and PhaC. In silico genome-scale metabolic-flux analysis was applied to inform further genetic-engineering approaches, and productivity was notably enhanced by knocking out the acetate kinase (ackA), phosphoenolpyruvate carboxylase (ppc) and aldehyde–alcohol dehydrogenase (adhE) genes as well as by replacing the promoters of the D-lactate dehydrogenase (ldhA) and acetyl-CoA synthetase (acs) genes with the strong trc promoter (Invitrogen)107,108.
The identification of key biosynthesis enzymes (for example, synthases, synthetases and polymerases) and polymer-modifying enzymes (for example, epimerases, acetyltransferases and lyases) and an increased understanding of their reaction mechanisms as well as their structure–function relationships promise not only to improve polymer production but also to allow us obtain new and tailor-made polymers34,43,84,105,109. Besides rational design of biosynthesis pathways, including engineering of key enzymes, random mutagenesis and site-directed evolution are valid strategies towards the development of polymer production strains26,110,111,112,113,114.
Whereas the cell, as a biosynthesis machine, can use cheap carbon sources as precursor substrates, such as waste products (glycerol, whey and so on), the in vitro synthesis of biopolymers using purified key enzymes or subcellular fractions relies on costly precursor molecules such as ATP, CoA, CoA thioesters and nucleotide sugars or sugar acids. In vitro synthesis has been achieved for various polymers (for example, PHA, cellulose, alginate and PGA), and it allows the ultimate control of polymer composition, but it has only limited commercial applicability owing to the very high production costs. For example, in vitro synthesis of the bioplastic PHB, which requires the costly precursor (R)-3-hydroxybutyryl-CoA and purified PHB synthase, would amount production costs of around US$286,000 per gram of PHB. By contrast, bacterial production of PHB at a scale of thousands of tonnes per year has been estimated to cost about $0.0025 per gram of PHB, and this is still 5–10 times as expensive to produce as the respective petroleum-based polymers.
The strategies outlined above show the enormous polymer design space available for bacterial polymer production, and this could be even further extended by exposing isolated biopolymers to further chemical or enzymatic modifications (Fig. 5). One example of enzymatic modification is the use of alginate epimerases with different substrate specificities to introduce guluronic acid residues at specific sites in alginates, thereby altering their material properties115 (Fig. 6). All the strategies for the production of tailor-made biopolymers that are mentioned above are strongly informed by knowledge of the structure–material property relationships of the respective biopolymers.
An exciting and recent development is the potential use of PHA granules, which are formed inside recombinant bacterial cells, as tailor-made functionalized micro- or nano-beads in which specific proteins attached to the PHA core have been engineered to display various protein functions (Fig. 4d). The application performance of engineered PHA beads in high-affinity bioseparation116,117,118, enzyme immobilization119, protein production120, diagnostics121 and as an antigen delivery system122 has been demonstrated, and the technology is now being commercialized (see Ref. 69 for a review).
Industrial production: where are the bottlenecks?
The ability to develop bacterial production strains by random approaches or by engineering biosynthesis pathways and key enzymes makes bacteria ideal hosts for the production of tailor-made polymers for use either as commodity products or in the high-value medical field. The production costs are driven by the yield of polymer relative to the amount of carbon source required, as well as by downstream processing requirements, which are strongly informed by the targeted application field. This is particularly relevant for intracellular polymers such as PHAs, as cells need to be lysed to release the polymer, which will subsequently be subjected to various separation processes. These separation processes become especially challenging when polymers are considered for medical applications. Depending on the production scale, which can range from kilograms to tonnes, biopolymer production by bacterial fermentation under current good manufacturing practice requires a substantial investment into capital equipment — from $1 million for kilogram yields to $10–100 million for tonne yields. Capital equipment costs are determined by the culture volume used, which in turn dictates the dimensions of the bioreactor and downstream processing equipment. Downstream processing involves the separation of biomass and culture supernatant, usually by filtration, followed by product-specific separation processes (for example, micro- or nanofiltration, solvent extraction, precipitation, chromatography or crystallization). When compared with the production process for synthetic polymers, the biotechnological process provides a notable reduction in capital equipment and operating costs and, moreover, waste disposal costs are lower and toxic metal catalysts are not used. However, the production costs of synthetic polymers depend on the price of crude oil, whereas bioprocesses are dependent on the price of feedstock such as sugars, starch, vegetable oils and glycerol.
Quo vadis? Biopolymers produced by bacteria
Although plant products or plant-derived biopolymers served as commodity materials before the 1920s, petrochemically synthesized polymers (including thermoplastics, thermoset plastics, adhesives and coatings) now dominate the commodity materials market, with an annual production estimated at 260 million tonnes in 2007 (Ref. 123). This suggests a previous growth rate of approximtely 9% per annum.
Owing to an increasing environmental awareness and the limitation of fossil resources, it is anticipated that renewable biopolymers will replace a substantial fraction of the market for synthetic polymers. There will also probably be a growing demand for bacterial production of polymers with material properties that are specifically tailored for applications in various fields of daily life. The competitive advantage of the environmentally friendly and highly processive synthesis of biodegradable and, in many cases, biocompatible polymers will become increasingly attractive for industry. Bacteria remain ideal production organisms for tailor-made polymers owing to the availability of genetic systems and techniques for engineering metabolic pathways, which provide an ever-growing design space for the production of biopolymers with material properties of interest. In addition, further knowledge of the molecular mechanisms of biopolymer biosynthesis could be used for the design of specific biosynthesis enzyme inhibitors that will be useful when the respective biopolymer is an important virulence factor.
References
Anderson, A. J., Haywood, G. W. & Dawes, E. A. Biosynthesis and composition of bacterial poly(hydroxyalkanoates). Int. J. Biol. Macromol. 12, 102–105 (1990).
Rehm, B. H. A. & Valla, S. Bacterial alginates: biosynthesis and applications. Appl. Microbiol. Biotechnol. 48, 281–288 (1997).
Ryder, C., Byrd, M. & Wozniak, D. J. Role of polysaccharides in Pseudomonas aeruginosa biofilm development. Curr. Opin. Microbiol. 10, 644–648 (2007).
Pham, T. H., Webb, J. S. & Rehm, B. H. A. The role of polyhydroxyalkanoate biosynthesis by Pseudomonas aeruginosa in rhamnolipid and alginate production as well as stress tolerance and biofilm formation. Microbiology 150, 3405–3413 (2004).
Vandamme, E. J., Bruggeman, G., De Baets, S. & Vanhooren, P. T. Useful polymers of microbial origin. Agro Food Industry Hi-Tech. 7, 21–25 (1996).
Chen, G. Q. A microbial polyhydroxyalkanoates (PHA) based bio- and materials industry. Chem. Soc. Rev. 38, 2434–2446 (2009).
Vollmer, W., Blanot, D. & de Pedro, M. A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 (2008).
Whitfield, C. Biosynthesis and assembly of capsular polysaccharides in Escherichia coli. Annu. Rev. Biochem. 75, 39–68 (2006).
Becker, A., Katzen, F., Puhler, A. & Ielpi, L. Xanthan gum biosynthesis and application: a biochemical/genetic perspective. Appl. Microbiol. Biotechnol. 50, 145–152 (1998).
Valla, S., Ertesvag, H., Tonouchi, N. & Fjaervig, E. in Microbial Production of Biopolymers and Polymer Precursors: Applications and Perspectives (ed. Rehm, B. H. A.) 43–78 (Caister Academic, Norwich, UK, 2009).
Remminghorst, U. & Rehm, B. H. A. Bacterial alginates: from biosynthesis to applications. Biotechnol. Lett. 28, 1701–1712 (2006).
Hickman, J. W. & Harwood, C. S. Identification of FleQ from Pseudomonas aeruginosa as a c-di-GMP-responsive transcription factor. Mol. Microbiol. 69, 376–389 (2008).
Leech, A. J., Sprinkle, A., Wood, L., Wozniak, D. J. & Ohman, D. E. The NtrC family regulator AlgB, which controls alginate biosynthesis in mucoid Pseudomonas aeruginosa, binds directly to the algD promoter. J. Bacteriol. 190, 581–589 (2008).
Sakuragi, Y. & Kolter, R. Quorum-sensing regulation of the biofilm matrix genes (pel) of Pseudomonas aeruginosa. J. Bacteriol. 189, 5383–5386 (2007).
Ionescu, M. & Belkin, S. Overproduction of exopolysaccharides by an Escherichia coli K-12 rpoS mutant in response to osmotic stress. Appl. Environ. Microbiol. 75, 483–492 (2009).
Weber, H., Pesavento, C., Possling, A., Tischendorf, G. & Hengge, R. Cyclic-di-GMP-mediated signalling within the σS network of Escherichia coli. Mol. Microbiol. 62, 1014–1034 (2006).
Castaneda, M., Sanchez, J., Moreno, S., Nunez, C. & Espin, G. The global regulators GacA and σS form part of a cascade that controls alginate production in Azotobacter vinelandii. J. Bacteriol. 183, 6787–6793 (2001).
Hall-Stoodley, L., Costerton, J. W. & Stoodley, P. Bacterial biofilms: from the natural environment to infectious diseases. Nature Rev. Microbiol. 2, 95–108 (2004).
Ruffing, A. & Chen, R. R. Metabolic engineering of microbes for oligosaccharide and polysaccharide synthesis. Microb. Cell Fact. 5, 25 (2006).
Parolis, H. et al. The structure of the exopolysaccharide produced by the halophilic Archaeon Haloferax mediterranei strain R4 (ATCC 33500). Carbohydr. Res. 295, 147–156 (1996).
Carrasco, F., Chornet, E., Overend, R. P. & Costa, J. A generalized correlation for the viscosity of dextrans in aequeous solutions as a function of temperature, concentration, and molecular weight at low shear rates. J. Appl. Polym. Sci. 37, 2087–2098 (1989).
Leathers, T. D. in Biopolymers for Medical and Pharmaceutical Applications (eds Steinbüchel, A. & Marchessault, R. H.) 537–560 (Wiley-VCH, Weinheim, 2005).
van Hijum, S. A., Kralj, S., Ozimek, L. K., Dijkhuizen, L. & van Geel-Schutten, I. G. Structure-function relationships of glucansucrase and fructansucrase enzymes from lactic acid bacteria. Microbiol. Mol. Biol. Rev. 70, 157–176 (2006).
Robyt, J. F., Yoon, S. H. & Mukerjea, R. Dextransucrase and the mechanism for dextran biosynthesis. Carbohydr. Res. 343, 3039–3048 (2008). This study demonstrates the involvement of conserved amino acids in the dextransucrase active site (Asp551, Glu589 and Asp622) in the polymerization of dextran, as well as showing that a single active site mediates dextran synthesis by a two-catalytic-site insertion mechanism.
Moulis, C. et al. Understanding the polymerization mechanism of glycoside-hydrolase family 70 glucansucrases. J. Biol. Chem. 281, 31254–31267 (2006).
Kim, D. & Robyt, J. F. Production and selection of mutants of Leuconostoc mesenteroides constitutive for glucansucrases. Enzyme Microb. Technol. 16, 659–664 (1994).
Brown, S. H. & Pummill, P. E. Recombinant production of hyaluronic acid. Curr. Pharm. Biotechnol. 9, 239–241 (2008).
Dougherty, B. A. & van de Rijn, I. Molecular characterization of hasA from an operon required for hyaluronic acid synthesis in group A streptococci. J. Biol. Chem. 269, 169–175 (1994).
Rehm, B. H. A. in Microbiology Monographs, Alginates: Biology and Applications (ed. Rehm, B. H. A.) 55–72 (Springer, Berlin, 2009).
Ohman, D. E. in Microbiology Monographs, Alginates: Biology and Applications (ed. Rehm, B. H. A.) 117–134 (Springer, Berlin, 2009).
Remminghorst, U., Hay, I. D. & Rehm, B. H. A. Molecular characterization of Alg8, a putative glycosyltransferase, involved in alginate polymerisation. J. Biotechnol. 140, 176–183 (2009).
Remminghorst, U. & Rehm, B. H. A. In vitro alginate polymerization and the functional role of Alg8 in alginate production by Pseudomonas aeruginosa. Appl. Environ. Microbiol. 72, 298–305 (2006). The first in vitro synthesis of alginate, providing evidence for a multiprotein polymerization and secretion complex spanning the envelope and identifying Alg8 as a key protein for alginate overproduction.
Saxena, I. M., Brown, R. M. Jr, Fevre, M., Geremia, R. A. & Henrissat, B. Multidomain architecture of β-glycosyl transferases: implications for mechanism of action. J. Bacteriol. 177, 1419–1424 (1995).
Steigedal, M. et al. The Azotobacter vinelandii AlgE mannuronan C-5-epimerase family is essential for the in vivo control of alginate monomer composition and for functional cyst formation. Environ. Microbiol. 10, 1760–1770 (2008).
Gimmestad, M. et al. The Pseudomonas fluorescens AlgG protein, but not its mannuronan C-5-epimerase activity, is needed for alginate polymer formation. J. Bacteriol. 185, 3515–3523 (2003). This article describes the production of modified alginate (polymannuronate) by engineering a P. fluorescens strain lacking mannuronan C5 epimerase activity.
Whitfield, C. & Paiment, A. Biosynthesis and assembly of Group 1 capsular polysaccharides in Escherichia coli and related extracellular polysaccharides in other bacteria. Carbohydr. Res. 338, 2491–2502 (2003).
Rahn, A., Drummelsmith, J. & Whitfield, C. Conserved organization in the cps gene clusters for expression of Escherichia coli group 1 K antigens: relationship to the colanic acid biosynthesis locus and the cps genes from Klebsiella pneumoniae. J. Bacteriol. 181, 2307–2313 (1999).
Arakawa, Y. et al. Genomic organization of the Klebsiella pneumoniae cps region responsible for serotype K2 capsular polysaccharide synthesis in the virulent strain Chedid. J. Bacteriol. 177, 1788–1796 (1995).
Cuthbertson, L., Mainprize, I. L., Naismith, J. H. & Whitfield, C. Pivotal roles of the outer membrane polysaccharide export and polysaccharide copolymerase protein families in export of extracellular polysaccharides in Gram-negative bacteria. Microbiol. Mol. Biol. Rev. 73, 155–177 (2009). An outstanding review describing the role of the conserved proteins involved in the final steps of exopolysaccharide formation.
Ford, R. C. et al. Structure-function relationships of the outer membrane translocon Wza investigated by cryo-electron microscopy and mutagenesis. J. Struct. Biol. 166, 172–182 (2009). This paper provides an insight into the structure–function relationship of an outer-membrane protein complex involved in export of expolysaccharides.
Whitfield, C. & Larue, K. Stop and go: regulation of chain length in the biosynthesis of bacterial polysaccharides. Nature Struct. Mol. Biol. 15, 121–123 (2008).
Raetz, C. R. & Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 71, 635–700 (2002).
Barreras, M., Salinas, S. R., Abdian, P. L., Kampel, M. A. & Ielpi, L. Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase. J. Biol. Chem. 283, 25027–25035 (2008).
Wang, L., Liu, D. & Reeves, P. R. C-terminal half of Salmonella enterica WbaP (RfbP) is the galactosyl-1-phosphate transferase domain catalyzing the first step of O-antigen synthesis. J. Bacteriol. 178, 2598–2604 (1996).
Wang, L. & Reeves, P. R. Involvement of the galactosyl-1-phosphate transferase encoded by the Salmonella enterica rfbP gene in O-antigen subunit processing. J. Bacteriol. 176, 4348–4356 (1994).
Drummelsmith, J. & Whitfield, C. Gene products required for surface expression of the capsular form of the group 1 K antigen in Escherichia coli (O9a:K30). Mol. Microbiol. 31, 1321–1332 (1999).
Tocilj, A. et al. Bacterial polysaccharide co-polymerases share a common framework for control of polymer length. Nature Struct. Mol. Biol. 15, 130–138 (2008).
Collins, R. F. & Derrick, J. P. Wza: a new structural paradigm for outer membrane secretory proteins? Trends Microbiol. 15, 96–100 (2007).
Sheng, F., Jia, X., Yep, A., Preiss, J. & Geiger, J. H. The crystal structures of the open and catalytically competent closed conformation of Escherichia coli glycogen synthase. J. Biol. Chem. 284, 17796–17807 (2009).
Horcajada, C., Guinovart, J. J., Fita, I. & Ferrer, J. C. Crystal structure of an archaeal glycogen synthase: insights into oligomerization and substrate binding of eukaryotic glycogen synthases. J. Biol. Chem. 281, 2923–2931 (2006).
Buschiazzo, A. et al. Crystal structure of glycogen synthase: homologous enzymes catalyze glycogen synthesis and degradation. EMBO J. 23, 3196–3205 (2004). A seminal paper reporting the first crystal structure of glycogen synthase in the presence and absence of adenosine diphosphate.
Oppermann-Sanio, F. B. & Steinbüchel, A. Occurrence, functions and biosynthesis of polyamides in microorganisms and biotechnological production. Naturwissenschaften 89, 11–22 (2002).
Joentgen, W., Groth, T., Steinbüchel, A., Hai, T. & Oppermann, F. B. Polyaspartic acid homopolymers and copolymers: biotechnical production and use thereof. International patent application WO 98/39090 (1998).
Schwammborn, M. Chemical synthesis of polyaspartates: a biodegradable alternative to currently used polycarboxylate homo- and copolymers. Polym. Degrad. Stab. 59, 39–45 (1998).
Berg, H. et al. Biosynthesis of the cyanobacterial reserve polymer multi-L-arginyl-poly-L-aspartic acid (cyanophycin): mechanism of the cyanophycin synthetase reaction studied with synthetic primers. Eur. J. Biochem. 267, 5561–5570 (2000).
Aboulmagd, E., Oppermann-Sanio, F. B. & Steinbüchel, A. Purification of Synechocystis sp. strain PCC6308 cyanophycin synthetase and its characterization with respect to substrate and primer specificity. Appl. Environ. Microbiol. 67, 2176–2182 (2001).
Simon, R. D. The biosynthesis of multi-L-arginyl-poly(L-aspartic acid) in the filamentous cyanobacterium Anabaena cylindrica. Biochim. Biophys. Acta 422, 407–418 (1976). This investigation demonstrates the isolation and purification of an enzyme that synthesizes cyanophycin in vitro and analyses the basic properties of the biosynthesis reaction.
Ziegler, K. et al. Molecular characterization of cyanophycin synthetase, the enzyme catalyzing the biosynthesis of the cyanobacterial reserve material multi-L-arginyl-poly-L-aspartate (cyanophycin). Eur. J. Biochem. 254, 154–159 (1998). The identification of the cyanophycin synthetase gene mediating recombinant cyanophycin production in E. coli and a biochemical characterization of the encoded cyanophycin synthetase.
Krehenbrink, M. & Steinbüchel, A. Partial purification and characterization of a non-cyanobacterial cyanophycin synthetase from Acinetobacter calcoaceticus strain ADP1 with regard to substrate specificity, substrate affinity and binding to cyanophycin. Microbiology 150, 2599–2608 (2004).
Buescher, J. M. & Margaritis, A. Microbial biosynthesis of polyglutamic acid biopolymer and applications in the biopharmaceutical, biomedical and food industries. Crit. Rev. Biotechnol. 27, 1–19 (2007).
Candela, T., Moya, M., Haustant, M. & Fouet, A. Fusobacterium nucleatum, the first Gram-negative bacterium demonstrated to produce polyglutamate. Can. J. Microbiol. 55, 627–632 (2009). The first description of poly-γ-glutamate biosynthesis by a Gram-negative bacterium.
Eveland, S. S., Pompliano, D. L. & Anderson, M. S. Conditionally lethal Escherichia coli murein mutants contain point defects that map to regions conserved among murein and folyl poly-γ-glutamate ligases: identification of a ligase superfamily. Biochemistry 36, 6223–6229 (1997).
Urushibata, Y., Tokuyama, S. & Tahara, Y. Characterization of the Bacillus subtilis ywsC gene, involved in γ-polyglutamic acid production. J. Bacteriol. 184, 337–343 (2002).
Ashiuchi, M., Soda, K. & Misono, H. A poly-γ-glutamate synthetic system of Bacillus subtilis IFO 3336: gene cloning and biochemical analysis of poly-γ-glutamate produced by Escherichia coli clone cells. Biochem. Biophys. Res. Commun. 263, 6–12 (1999).
Candela, T. & Fouet, A. Poly-γ-glutamate in bacteria. Mol. Microbiol. 60, 1091–1098 (2006).
Candela, T. & Fouet, A. Bacillus anthracis CapD, belonging to the γ-glutamyltranspeptidase family, is required for the covalent anchoring of capsule to peptidoglycan. Mol. Microbiol. 57, 717–726 (2005).
Yamanaka, K., Maruyama, C., Takagi, H. & Hamano, Y. ɛ-poly-L-lysine dispersity is controlled by a highly unusual nonribosomal peptide synthetase. Nature Chem. Biol. 4, 766–772 (2008). A seminal paper describing the cloning of the ɛ-poly- L -lysine synthetase gene and the enzymatic characterization of the encoded protein, revealing an amino acid ligase-like catalytic activity for peptide bond formation.
Kessler, B. & Witholt, B. Factors involved in the regulatory network of polyhydroxyalkanoate metabolism. J. Biotechnol. 86, 97–104 (2001).
Grage, K. et al. Bacterial polyhydroxyalkanoate granules: biogenesis, structure, and potential use as nano-/micro-beads in biotechnological and biomedical applications. Biomacromolecules 10, 660–669 (2009).
Jendrossek, D. Polyhydroxyalkanoate granules are complex subcellular organelles (carbonosomes). J. Bacteriol. 191, 3195–3202 (2009).
Andrade, A. L. in Plastics and the Environment (ed. Andrady, A.) 3–76 (Wiley Interscience, Hoboken, 2003).
Rehm, B. H. A. & Steinbüchel, A. Biochemical and genetic analysis of PHA synthases and other proteins required for PHA synthesis. Int. J. Biol. Macromol. 25, 3–19 (1999).
Aldor, I. S. & Keasling, J. D. Process. design for microbial plastic factories: metabolic engineering of polyhydroxyalkanoates. Curr. Opin. Biotechnol. 14, 475–483 (2003). An excellent review on metabolic-engineering approaches towards the production of modified and new PHAs in the context of biotechnological production processes.
Fukui, T., Shiomi, N. & Doi, Y. Expression and characterization of (R)-specific enoyl coenzyme A hydratase involved in polyhydroxyalkanoate biosynthesis by Aeromonas caviae. J. Bacteriol. 180, 667–673 (1998).
Rehm, B. H. A., Krüger, N. & Steinbüchel, A. A new metabolic link between fatty acid de novo synthesis and polyhydroxyalkanoic acid synthesis. The phaG gene from Pseudomonas putida KT2440 encodes a 3-hydroxyacyl-acyl carrier protein-coenzyme A transferase. J. Biol. Chem. 273, 24044–24051 (1998).
Schubert, P., Steinbüchel, A. & Schlegel, H. G. Cloning of the Alcaligenes eutrophus genes for synthesis of poly-β-hydroxybutyric acid (PHB) and synthesis of PHB in Escherichia coli. J. Bacteriol. 170, 5837–5847 (1988).
Slater, S. C., Voige, W. H. & Dennis, D. E. Cloning and expression in Escherichia coli of the Alcaligenes eutrophus H16 poly-β-hydroxybutyrate biosynthetic pathway. J. Bacteriol. 170, 4431–4436 (1988).
Peoples, O. P. & Sinskey, A. J. Poly-β-hydroxybutyrate biosynthesis in Alcaligenes eutrophus H16. Characterization of the genes encoding β-ketothiolase and acetoacetyl-CoA reductase. J. Biol. Chem. 264, 15293–15297 (1989).
Rehm, B. H. A. Polyester synthases: natural catalysts for plastics. Biochem. J. 376, 15–33 (2003).
Steinbüchel, A. et al. Molecular basis for biosynthesis and accumulation of polyhydroxyalkanoic acids in bacteria. FEMS Microbiol. Rev. 104, 347–350 (1993).
Wodzinska, J. et al. Polyhydroxybutyrate synthase: evidence for covalent catalysis. J. Am. Chem. Soc. 118, 6319–6320 (1996).
Tian, J., Sinskey, A. J. & Stubbe, J. Detection of intermediates from the polymerization reaction catalyzed by a D302A mutant of class III polyhydroxyalkanoate (PHA) synthase. Biochemistry 44, 1495–1503 (2005). This study provides insight into the reaction mechanism of the PHA synthase.
Tian, J., Sinskey, A. J. & Stubbe, J. Class III polyhydroxybutyrate synthase: involvement in chain termination and reinitiation. Biochemistry 44, 8369–8377 (2005).
Sim, S. J. et al. PHA synthase activity controls the molecular weight and polydispersity of polyhydroxybutyrate in vivo. Nature Biotech. 15, 63–67 (1997). This work provides evidence that PHA chain length is controlled by the copy number of the PHA synthase.
Reusch, R. N. & Sadoff, H. L. Putative structure and functions of a poly-β-hydroxybutyrate/calcium polyphosphate channel in bacterial plasma membranes. Proc. Natl Acad. Sci. USA 85, 4176–4180 (1988).
Kameda, A. et al. A novel ATP regeneration system using polyphosphate-AMP phosphotransferase and polyphosphate kinase. J. Biosci. Bioeng. 91, 557–5563 (2001).
Noguchi, T. & Shiba, T. Use of Escherichia coli polyphosphate kinase for oligosaccharide synthesis. Biosci. Biotechnol. Biochem. 62, 1594–1596 (1998).
Zhu, Y., Huang, W., Lee, S. S. & Xu, W. Crystal structure of a polyphosphate kinase and its implications for polyphosphate synthesis. EMBO Rep. 6, 681–687 (2005).
Kumble, K. D., Ahn, K. & Kornberg, A. Phosphohistidyl active sites in polyphosphate kinase of Escherichia coli. Proc. Natl Acad. Sci. USA 93, 14391–14395 (1996).
Ishige, K., Zhang, H. & Kornberg, A. Polyphosphate kinase (PPK2), a potent, polyphosphate-driven generator of GTP. Proc. Natl Acad. Sci. USA 99, 16684–16688 (2002).
Kim, H. Y. et al. Alginate, inorganic polyphosphate, GTP and ppGpp synthesis co-regulated in Pseudomonas aeruginosa: implications for stationary phase survival and synthesis of RNA/DNA precursors. Mol. Microbiol. 27, 717–725 (1998).
Rao, N. N., Gomez-Garcia, M. R. & Kornberg, A. Inorganic polyphophate: essential for growth and survival. Annu. Rev. Biochem. 78, 605–647 (2009). An outstanding review describing the synthesis of polyphosphates and their biological functions.
Pohlmann, A. et al. Genome sequence of the bioplastic-producing “Knallgas” bacterium Ralstonia eutropha H16. Nature Biotech. 24, 1257–1262 (2006). This study unravels the genome sequence of R. eutropha and reflects on the status of this species as a model bioplastics producer.
Vorholter, F. J. et al. The genome of Xanthomonas campestris pv. campestris B100 and its use for the reconstruction of metabolic pathways involved in xanthan biosynthesis. J. Biotechnol. 134, 33–45 (2008).
Lee, S. Y. Deciphering bioplastic production. Nature Biotech. 24, 1227–1229 (2006).
Kalia, V. C., Chauhan, A., Bhattacharyya, G. & Rashmi . Genomic databases yield novel bioplastic producers. Nature Biotech. 21, 845–846 (2003).
Peoples, O. P. & Sinskey, A. J. Poly-β-hydroxybutyrate (PHB) biosynthesis in Alcaligenes eutrophus H16. Identification and characterization of the PHB polymerase gene (phbC). J. Biol. Chem. 264, 15298–15303 (1989).
Chien, L. J. & Lee, C. K. Enhanced hyaluronic acid production in Bacillus subtilis by coexpressing bacterial hemoglobin. Biotechnol. Prog. 23, 1017–1022 (2007).
Jiang, H., Shang, L., Yoon, S. H., Lee, S. Y. & Yu, Z. Optimal production of poly-γ-glutamic acid by metabolically engineered Escherichia coli. Biotechnol. Lett. 28, 1241–1246 (2006).
Park, S. J., Lee, S. Y. & Lee, Y. Biosynthesis of (R)-3-hydroxyalkanoic acids by metabolically engineered Escherichia coli. Appl. Biochem. Biotechnol. 114, 373–379 (2004).
Lee, S. Y. & Choi, J. I. Production of microbial polyester by fermentation of recombinant microorganisms. Adv. Biochem. Eng. Biotechnol. 71, 183–207 (2001).
Frey, K. M., Oppermann-Sanio, F. B., Schmidt, H. & Steinbüchel, A. Technical-scale production of cyanophycin with recombinant strains of Escherichia coli. Appl. Environ. Microbiol. 68, 3377–3384 (2002).
Mao, Z., Shin, H. D. & Chen, R. A recombinant E. coli bioprocess for hyaluronan synthesis. Appl. Microbiol. Biotechnol. 84, 63–69 (2009).
Lutke-Eversloh, T. et al. Biosynthesis of novel thermoplastic polythioesters by engineered Escherichia coli. Nature Mater. 1, 236–240 (2002). The first description of the production of unnatural polythioesters using metabolically engineered E. coli as the producing species and mercaptoalkanoates as precursor substrates.
Taguchi, S. et al. A microbial factory for lactate-based polyesters using a lactate-polymerizing enzyme. Proc. Natl Acad. Sci. USA 105, 17323–17327 (2008). The first paper to describe the incorporation of lactate into PHA by metabolically engineered E. coli , providing an important step towards one-step production of polylactic acid.
Doi, Y. Biotechnology: unnatural biopolymers. Nature Mater. 1, 207–208 (2002).
Jung, Y. K., Kim, T. Y., Park, S. J. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of polylactic acid and its copolymers. Biotechnol. Bioeng. 105, 161–171 (2010).
Yang, T. H. et al. Biosynthesis of polylactic acid and its copolymers using evolved propionate CoA transferase and PHA synthase. Biotechnol. Bioeng. 105, 150–160 (2010).
Amara, A. A. & Rehm, B. H. A. Replacement of the catalytic nucleophile cysteine-296 by serine in class II polyhydroxyalkanoate synthase from Pseudomonas aeruginosa-mediated synthesis of a new polyester: identification of catalytic residues. Biochem. J. 374, 413–421 (2003).
Rehm, B. H. A., Antonio, R. V., Spiekermann, P., Amara, A. A. & Steinbüchel, A. Molecular characterization of the poly(3-hydroxybutyrate) (PHB) synthase from Ralstonia eutropha: in vitro evolution, site-specific mutagenesis and development of a PHB synthase protein model. Biochim. Biophys. Acta 1594, 178–190 (2002).
Kichise, T., Taguchi, S. & Doi, Y. Enhanced accumulation and changed monomer composition in polyhydroxyalkanoate (PHA) copolyester by in vitro evolution of Aeromonas caviae PHA synthase. Appl. Environ. Microbiol. 68, 2411–2419 (2002).
Monchois, V., Vignon, M. & Russell, R. R. Mutagenesis of asp-569 of glucosyltransferase I glucansucrase modulates glucan and oligosaccharide synthesis. Appl. Environ. Microbiol. 66, 1923–1927 (2000).
Füser, G. & Steinbüchel, A. Analysis of genome sequences for genes of cyanophycin metabolism: identifying putative cyanophycin metabolizing prokaryotes. Macromol. Biosci. 7, 278–296 (2007).
Schallmey, M., Singh, A. & Ward, O. P. Developments in the use of Bacillus species for industrial production. Can. J. Microbiol. 50, 1–17 (2004).
Valla, S., Li, J., Ertesvag, H., Barbeyron, T. & Lindahl, U. Hexuronyl C5-epimerases in alginate and glycosaminoglycan biosynthesis. Biochimie 83, 819–830 (2001).
Lewis, J. G. & Rehm, B. H. A. ZZ polyester beads: an efficient and simple method for purifying IgG from mouse hybridoma supernatants. J. Immunol. Methods 346, 71–74 (2009).
Grage, K. & Rehm, B. H. A. In vivo production of scFv-displaying biopolymer beads using a self-assembly-promoting fusion partner. Bioconjug. Chem. 19, 254–262 (2008).
Brockelbank, J. A., Peters, V. & Rehm, B. H. A. Recombinant Escherichia coli strain produces a ZZ domain displaying biopolyester granules suitable for immunoglobulin G purification. Appl. Environ. Microbiol. 72, 7394–7397 (2006).
Rasiah, I. A. & Rehm, B. H. A. One-step production of immobilized α-amylase in recombinant Escherichia coli. Appl. Environ. Microbiol. 75, 2012–2016 (2009).
Banki, M. R., Gerngross, T. U. & Wood, D. W. Novel and economical purification of recombinant proteins: intein-mediated protein purification using in vivo polyhydroxybutyrate (PHB) matrix association. Protein Sci. 14, 1387–1395 (2005).
Baeckstroem, T. B., Brockelbank, J. A. & Rehm, B. H. A. Recombinant Escherichia coli produces tailor-made biopolyester granules for applications in fluorescence activated cell sorting: functional display of the mouse interleukin-2 and myelin oligodendrocyte glycoprotein. BMC Biotechnol. 7, 3 (2007).
Parlane, N. A., Wedlock, D. N., Buddle, B. M. & Rehm, B. H. A. Bacterial polyester inclusions engineered to display vaccine candidate antigens for use as a novel class of safe and efficient vaccine delivery agents. Appl. Environ. Microbiol. 75, 7739–7744 (2009). The first description of the application of engineered bacterial-polymer inclusions as antigen delivery systems.
Hopewell, J., Dvorak, R. & Kosior, E. Plastics recycling: challenges and opportunities. Philos. Trans R. Soc. Lond. B Biol. Sci. 364, 2115–2126 (2009).
Pasteur, L. On the viscous fermentation and the butyrous fermentation. Bull. Soc. Chim. Paris 11, 30–31 (1861) (in French).
Van Tieghem, P. On sugar mill gum. Ann. Sci. Nature Bot. Biol. Veg. 7, 180–203 (1878).
Brown, A. J. On an acetic ferment which forms cellulose. J. Chem. Soc. 49, 432–439 (1886).
Borzi, A. Le communicazioni intracellulari delle Nostochinae. Malpighia 1, 28–74 (1887) (in Italian).
Lemoigne, M. Produits de deshydration et de polymerisation de lácide β-oxybutyrique. Bull. Soc. Chim. Bio. 8, 770–782 (1926) (in French).
Linker, A. & Jones, R. S. A new polysaccharide resembling alginic acid isolated from pseudomonads. J. Biol. Chem. 241, 3845–3851 (1966).
Leach, J. G., Lilly, V. G., Wilson, H. A. & Purvis, M. R. Bacterial polysaccharides: the nature and the function of the exudate produced by Xanthomonas phaseoli. Phytopathology 47, 113–120 (1957).
Ivanovics, G. & Erdos, L. Ein beitrag zum wesen der kapselsubstanz des milzbrandbazillus. Z. Immunitatsforsch. 90, 5–19 (1937) (in German).
Kornberg, A., Kornberg, S. R. & Simms, E. S. Metaphosphate synthesis by an enzyme from Escherichia coli. Biochim. Biophys. Acta 20, 215–227 (1956).
Pindar, D. F. & Bucke, C. The biosynthesis of alginic acid by Azotobacter vinelandii. Biochem. J. 152, 617–622 (1975).
Couperwhite, I. & McCallum, M. F. Polysaccharide production and the possible occurrence of GDP-D-mannose dehydrogenase in Azotobacter vinelandii. Antonie Van Leeuwenhoek 41, 25–32 (1975).
Hestrin, S., Aschner, M. & Mager, J. Synthesis of cellulose by resting cells of Acetobacter xylinum. Nature 159, 64–65 (1947).
Hestrin, S. & Schramm, M. Synthesis of cellulose by Acetobacter xylinum. II. Preparation of freeze-dried cells capable of polymerizing glucose to cellulose. Biochem. J. 58, 345–352 (1954).
Glaser, L. The enzymic synthesis of cellulose by Acetobacter xylinum. Biochim. Biophys. Acta 25, 436 (1957).
Ielpi, L., Couso, R. O. & Dankert, M. A. Pyruvic acid acetal residues are transferred from phosphoenolpyruvate to the pentasaccharide-P-P-lipid. Biochem. Biophys. Res. Commun. 102, 1400–1408 (1981).
Oeding, V. & Schlegel, H. G. β-ketothiolase from Hydrogenomonas eutropha H16 and its significance in the regulation of poly-β-hydroxybutyrate metabolism. Biochem. J. 134, 239–248 (1973).
Greenberg, E. & Preiss, J. The occurrence of adenosine diphosphate glucose: glycogen transglucosylase in bacteria. J. Biol. Chem. 239, 4314–4315 (1964).
Thorne, L., Tansey, L. & Pollock, T. J. Clustering of mutations blocking synthesis of xanthan gum by Xanthomonas campestris. J. Bacteriol. 169, 3593–3600 (1987).
Darzins, A., Wang, S. K., Vanags, R. I. & Chakrabarty, A. M. Clustering of mutations affecting alginic acid biosynthesis in mucoid Pseudomonas aeruginosa. J. Bacteriol. 164, 516–524 (1985).
Wong, H. C. et al. Genetic organization of the cellulose synthase operon in Acetobacter xylinum. Proc. Natl Acad. Sci. USA 87, 8130–8134 (1990).
Hara, T., Nagatomo, S., Ogata, S. & Ueda, S. The DNA sequence of γ-glutamyltranspeptidase gene of Bacillus subtilis (natto) plasmid pUH1. Appl. Microbiol. Biotechnol. 37, 211–215 (1992).
Remminghorst, U. & Rehm, B. H. A. in Microbial Production of Biopolymers and Polymer Precursors: Applications and Perspectives (ed. Rehm, B. H. A.) 13–42 (Caister Academic, Norwich, UK, 2009).
Acknowledgements
I am indebted to all former and current co-workers who have contributed substantially to the biopolymer research presented in this Review, and regret that I could not refer to all of the studies that contributed to the bacterial polymer research field owing to space constraints. This work was supported by the Deutsche Forschungsgemeinschaft, Massey University (Palmerston North, New Zealand), PolyBatics Ltd (Palmerston North) and the Foundation of Research, Science and Technology in New Zealand.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Related links
DATABASES
Entrez Genome Project
Protein Data Bank
FURTHER INFORMATION
Glossary
- Biocompatibility
-
In a biomaterials context, the ability of a material or polymer that is non-toxic to avoid eliciting an immune response.
- Two-component signal transduction pathway
-
A regulatory pathway found in most bacterial and archaeal species that uses phosphotransfer schemes involving two conserved components, a histidine protein kinase and a response regulator protein.
- Quorum sensing
-
The regulation of bacterial gene expression in response to fluctuations in cell population density. Quorum sensing is mediated by the release of chemical signal molecules (such as homoserine lactones) called autoinducers.
- Pseudoplastic
-
A material that exhibits so- called shear thinning, which is a decrease in viscosity with an increase in the rate of shear stress.
- Newtonian fluid
-
A fluid exhibiting a linear relationship between the shear stress and the strain rate, with the proportionality being the coefficient of viscosity.
- Immunogenicity
-
The potential of a compound to elicit an immune response.
- Lipid carrier
-
An amphipathic molecule, with a hydrophobic polyisoprenoid moiety and a hydrophilic phosphate residue, that is embedded in the cytoplasmic membrane for the transport of sugar repeat units.
- Crystallinity
-
The degree of highly ordered structures inside a polymer. In the polymer context, the glass transition temperature and the melting temperature increase with increasing crystallinity, whereas the elongation-at-break value decreases.
- Rock phosphate
-
A natural inorganic-phosphate resource that, it is assumed, was prebiotically present on earth.
Rights and permissions
About this article
Cite this article
Rehm, B. Bacterial polymers: biosynthesis, modifications and applications. Nat Rev Microbiol 8, 578–592 (2010). https://doi.org/10.1038/nrmicro2354
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2354
This article is cited by
-
3D bioprinting of microorganisms: principles and applications
Bioprocess and Biosystems Engineering (2024)
-
Exploring polyhydroxyalkanoates biosynthesis using hydrocarbons as carbon source: a comprehensive review
Biodegradation (2024)
-
Development and Characterization of New Environmentally Friendly Polylactide Formulations with Terpenoid-Based Plasticizers with Improved Ductility
Journal of Polymers and the Environment (2024)
-
Selective isolation and genomic characterization of biopolymer producer—a novel feature of halophile Brachybacterium paraconglomeratum MTCC 13074
Journal of Genetic Engineering and Biotechnology (2023)
-
Spiro-salen catalysts enable the chemical synthesis of stereoregular polyhydroxyalkanoates
Nature Catalysis (2023)