Abstract
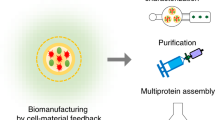
Bacteria can be programmed to create engineered living materials (ELMs) with self-healing and evolvable functionalities. However, further development of ELMs is greatly hampered by the lack of engineerable nonpathogenic chassis and corresponding programmable endogenous biopolymers. Here, we describe a technological workflow for facilitating ELMs design by rationally integrating bioinformatics, structural biology and synthetic biology technologies. We first develop bioinformatics software, termed Bacteria Biopolymer Sniffer (BBSniffer), that allows fast mining of biopolymers and biopolymer-producing bacteria of interest. As a proof-of-principle study, using existing pathogenic pilus as input, we identify the covalently linked pili (CLP) biosynthetic gene cluster in the industrial workhorse Corynebacterium glutamicum. Genetic manipulation and structural characterization reveal the molecular mechanism of the CLP assembly, ultimately enabling a type of programmable pili for ELM design. Finally, engineering of the CLP-enabled living materials transforms cellulosic biomass into lycopene by coupling the extracellular and intracellular bioconversion ability.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All produced data that support the findings of this study are included in this published article. The internal bacterial database used for BBSniffer is available at GuitHub (https://github.com/xbiome/BBSniffer/blob/publish/Database/Bacteria_database.xlsx). The structural factors and coordinates of the Spa2 have been deposited in the PDB under ID no. 7WOI. Source data are provided with this paper.
Code availability
The code of BBSniffer software has been deposited in GitHub: https://github.com/xbiome/BBSniffer/.
References
Tang, T. C. et al. Materials design by synthetic biology. Nat. Rev. Mater. 6, 332–350 (2021).
Huang, Y. et al. Engineering microbial systems for the production and functionalization of biomaterials. Curr. Opin. Microbiol. 68, 102154 (2022).
Gilbert, C. et al. Living materials with programmable functionalities grown from engineered microbial co-cultures. Nat. Mater. 20, 691–700 (2021).
Cao, Y. et al. Programmable assembly of pressure sensors using pattern-forming bacteria. Nat. Biotechnol. 35, 1087–1093 (2017).
Pu, J. et al. Virus disinfection from environmental water sources using living engineered biofilm materials. Adv. Sci. 7, 1903558 (2020).
Liu, A. P. et al. The living interface between synthetic biology and biomaterial design. Nat. Mater. 21, 390–397 (2022).
Praveschotinunt, P. et al. Engineered E. coli Nissle 1917 for the delivery of matrix-tethered therapeutic domains to the gut. Nat. Commun. 10, 5580 (2019).
Dai, Z. et al. Versatile biomanufacturing through stimulus-responsive cell-material feedback. Nat. Chem. Biol. 15, 1017–1024 (2019).
Dai, Z. et al. Living fabrication of functional semi-interpenetrating polymeric materials. Nat. Commun. 12, 3422 (2021).
Nguyen, P. Q. et al. Wearable materials with embedded synthetic biology sensors for biomolecule detection. Nat. Biotechnol. 39, 1366–1374 (2021).
Bird, L. J. et al. Engineered living conductive biofilms as functional materials. MRS Commun. 9, 505–517 (2019).
Rodrigo-Navarro, A., Sankaran, S., Dalby, M. J., Del Campo, A. & Salmeron-Sanchez, M. Engineered living biomaterials. Nat. Rev. Mater. 6, 1175–1190 (2021).
Bracha, D., Walls, M. T. & Brangwynne, C. P. Probing and engineering liquid-phase organelles. Nat. Biotechnol. 37, 1435–1445 (2019).
Bourdeau, R. W. et al. Acoustic reporter genes for noninvasive imaging of microorganisms in mammalian hosts. Nature 553, 86–90 (2018).
Nguyen, P. Q., Botyanszki, Z., Tay, P. K. R. & Joshi, N. S. Programmable biofilm-based materials from engineered curli nanofibres. Nat. Commun. 5, 4945 (2014).
Huang, J. et al. Programmable and printable Bacillus subtilis biofilms as engineered living materials. Nat. Chem. Biol. 15, 34–41 (2019).
McBee, R. M. et al. Engineering living and regenerative fungal-bacterial biocomposite structures. Nat. Mater. 21, 471–478 (2021).
An, B. et al. Programming living glue systems to perform autonomous mechanical repairs. Matter 3, 2080–2092 (2020).
Wang, Y. et al. Living materials fabricated via gradient mineralization of light-inducible biofilms. Nat. Chem. Biol. 17, 351–359 (2021).
Caro-Astorga, J., Walker, K. T., Herrera, N., Lee, K. Y. & Ellis, T. Bacterial cellulose spheroids as building blocks for 3D and patterned living materials and for regeneration. Nat. Commun. 12, 5027 (2021).
Charrier, M. et al. Engineering the S-layer of Caulobacter crescentus as a foundation for stable, high-density, 2D living materials. ACS Synth. Biol. 8, 181–190 (2018).
Kim, L. J. et al. Prospecting for natural products by genome mining and microcrystal electron diffraction. Nat. Chem. Biol. 17, 872–877 (2021).
Ramirez, N. A., Das, A. & Ton-That, H. New paradigms of pilus assembly mechanisms in gram-positive actinobacteria. Trends Microbiol. 28, 999–1009 (2020).
Moradali, M. F. & Rehm, B. H. Bacterial biopolymers: from pathogenesis to advanced materials. Nat. Rev. Microbiol. 18, 195–210 (2020).
Blin, K. et al. antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res. 49, W29–W35 (2021).
Skinnider, M. A., Merwin, N. J., Johnston, C. W. & Magarvey, N. A. PRISM 3: expanded prediction of natural product chemical structures from microbial genomes. Nucleic Acids Res. 45, W49–W54 (2017).
Weber, T. et al. CLUSEAN: a computer-based framework for the automated analysis of bacterial secondary metabolite biosynthetic gene clusters. J. Biotechnol. 140, 13–17 (2009).
McConnell, S. A. et al. Protein labeling via a specific lysine-isopeptide bond using the pilin polymerizing sortase from Corynebacterium diphtheriae. J. Am. Chem. Soc. 140, 8420–8423 (2018).
Zhao, N. et al. Development of a transcription factor-based diamine biosensor in Corynebacterium glutamicum. ACS Synth. Biol. 10, 3074–3083 (2021).
Mandlik, A., Swierczynski, A., Das, A. & Ton-That, H. Pili in gram-positive bacteria: assembly, involvement in colonization and biofilm development. Trends Microbiol. 16, 33–40 (2008).
Ton‐That, H., Marraffini, L. A. & Schneewind, O. Sortases and pilin elements involved in pilus assembly of Corynebacterium diphtheriae. Mol. Microbiol. 53, 251–261 (2004).
Kang, H. J., Paterson, N. G., Gaspar, A. H., Ton-That, H. & Baker, E. N. The Corynebacterium diphtheriae shaft pilin SpaA is built of tandem Ig-like modules with stabilizing isopeptide and disulfide bonds. Proc. Natl Acad. Sci. USA 106, 16967–16971 (2009).
Kang, H. J. et al. A slow-forming isopeptide bond in the structure of the major pilin SpaD from Corynebacterium diphtheriae has implications for pilus assembly. Acta Crystallogr. D. Biol. Crystallogr. 70, 1190–1201 (2014).
Kang, H. J., Coulibaly, F., Clow, F., Proft, T. & Baker, E. N. Stabilizing isopeptide bonds revealed in gram-positive bacterial pilus structure. Science 318, 1625–1628 (2007).
Budzik, J. M. et al. Intramolecular amide bonds stabilize pili on the surface of bacilli. Proc. Natl Acad. Sci. USA 106, 19992–19997 (2009).
Wang, X. et al. Photocatalyst-mineralized biofilms as living bio-abiotic interfaces for single enzyme to whole-cell photocatalytic applications. Sci. Adv. 8, eabm7665 (2022).
Reddington, S. C. & Howarth, M. Secrets of a covalent interaction for biomaterials and biotechnology: SpyTag and SpyCatcher. Curr. Opin. Chem. Biol. 29, 94–99 (2015).
Lau, Y. H., Giessen, T. W., Altenburg, W. J. & Silver, P. A. Prokaryotic nanocompartments form synthetic organelles in a eukaryote. Nat. Commun. 9, 1311 (2018).
Tang, H. et al. Efficient yeast surface-display of novel complex synthetic cellulosomes. Microb. Cell. Fact. 17, 122 (2018).
Kodama, Y. & Hu, C. D. An improved bimolecular fluorescence complementation assay with a high signal-to-noise ratio. BioTechniques 49, 793–805 (2010).
Lin, K., Han, S. & Zheng, S. Application of Corynebacterium glutamicum engineering display system in three generations of biorefinery. Microb. Cell. Fact. 21, 14 (2022).
Anusree, M., Wendisch, V. F. & Nampoothiri, K. M. Co-expression of endoglucanase and β-glucosidase in Corynebacterium glutamicum DM1729 towards direct lysine fermentation from cellulose. Bioresour. Technol. 213, 239–244 (2016).
Heider, S. A., Peters-Wendisch, P. & Wendisch, V. F. Carotenoid biosynthesis and overproduction in Corynebacterium glutamicum. BMC Microbiol. 12, 198 (2012).
Li, C. et al. Heterologous production of α-Carotene in Corynebacterium glutamicum using a multi-copy chromosomal integration method. Bioresour. Technol. 341, 125782 (2021).
Wang, Y. et al. Directed evolution: methodologies and applications. Chem. Rev. 121, 12384–12444 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Lu, H. et al. Machine learning-aided engineering of hydrolases for PET depolymerization. Nature 604, 662–667 (2022).
Ma, X. et al. Upcycling chitin-containing waste into organonitrogen chemicals via an integrated process. Proc. Natl Acad. Sci. USA 117, 7719–7728 (2020).
Anishchenko, I. et al. De novo protein design by deep network hallucination. Nature 600, 547–552 (2021).
Cai, P. et al. SynBioStrainFinder: a microbial strain database of manually curated CRISPR/Cas genetic manipulation system information for biomanufacturing. Microb. Cell. Fact. 21, 87 (2022).
Criscuolo, A. A fast alignment-free bioinformatics procedure to infer accurate distance-basedphylogenetic trees from genome assemblies. Res. Ideas Outcomes 5, e36178 (2019).
Yu, G., Lam, T. T.-Y., Zhu, H. & Guan, Y. Two methods for mapping and visualizing associated data on phylogeny using ggtree. Mol. Biol. Evol. 35, 3041–3043 (2018).
Huerta-Cepas, J. et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol. Biol. Evol. 34, 2115–2122 (2017).
Römling, U. & Galperin, M. Y. Bacterial cellulose biosynthesis: diversity of operons, subunits, products, and functions. Trends Microbiol. 23, 545–557 (2015).
Kennedy, N. W., Mills, C. E., Nichols, T. M., Abrahamson, C. H. & Tullman-Ercek, D. Bacterial microcompartments: tiny organelles with big potential. Curr. Opin. Microbiol. 63, 36–42 (2021).
Budzik, J. M., Marraffini, L. A. & Schneewind, O. Assembly of pili on the surface of Bacillus cereus vegetative cells. Mol. Microbiol. 66, 495–510 (2007).
Mccoy, A. J. et al. PHASER crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Dong, C. et al. Engineering Pichia pastoris with surface-display minicellulosomes for carboxymethyl cellulose hydrolysis and ethanol production. Biotechnol. Biofuels 13, 108 (2020).
Acknowledgements
This work was partially sponsored by the National Key R&D Program of China (grant no. 2020YFA0908100) (C.Z.), the National Science Fund for Distinguished Young Scholars (grant no. 32125023) (C.Z.), the Shenzhen Science and Technology Program (grant no. ZDSYS20220606100606013) (C.Z.), the National Natural Science Foundation of China (grant no. U1932204) (C.Z.), the Guangdong Basic and Applied Basic Research Foundation (grant no. 2021A1515110149) (Y.H.), the National Natural Science Foundation of China (grant no. 32301226) (Y.H.), the Shenzhen Science and Technology Program (grant no. ZDSYS20210623091810032) (J.Z.) and the National Natural Science Foundation of China (grant nos. 32271501 and 31971354) (N.L.). We thank the Electron Microscopy Center and the Analytical Instrumentation Center at the School of Physical Science and Technology, ShanghaiTech University; the Core Facility at Shenzhen Institute of Advanced Technology, Chinese Academy of Sciences. We also thank the staff of beamlines BL18U1 of Shanghai Synchrotron Radiation Facility for access and help with the X-ray data collection. We also thank S. Zheng (South China University of Technology) for the kind gift of the C. glutamicum ATCC 14067 strain and the RecET/Cre-loxP system related plasmids.
Author information
Authors and Affiliations
Contributions
C.Z. and Y.H. conceived the concept and directed the research. C.Z. and Y.H. designed and conducted the experiments and data analysis. J.Z. and Y.W. carried out crystallographic studies. Y.T., H.H. and B.T. performed the development of software. J.W., Y.M.W. and J.P. participated in plasmids and strains construction. S.Z. contributed to AFM imaging experiments. Y.Y.W., J.Z. and B.A. assisted with TEM imaging experiments. N.L., C.P., Y.Y., S.D. and W.Z. helped perform the mass spectrometry analysis. C.Z. and Y.H. wrote the paper with help from all authors.
Corresponding authors
Ethics declarations
Competing interests
C.Z. and Y.H. are co-inventors on patent applications (no. PCT/CN2022/130033) filed by Shenzhen Institute of Advanced Technology, based on the Spa2-based work covered in this article. C.Z., Y.H., Y.T., H.H. and B.T. are co-inventors on patent applications (no. 202310091110.2) and software copyright applications (no. 2023SR0280226) filed by Shenzhen Institute of Advanced Technology and Shenzhen Xbiome Biotech Co. Ltd., based on the BBSniffer software reported in this article. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Anna Duraj-Thatte and the other anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The use of the BBSniffer software for mining biopolymer-producers of bacterial cellulose (BC), gas vesicle (GV) and bacterial microcompartment (BMC).
(a) For bacteria cellulose-producer mining, 7,109 strains were detected, and 1,767 strains are grouped in industrial microorganisms based on our internal bacterial database for classification. BBSniffer scored the distance between the mined industrial microorganisms and the reference strains of Komagataeibacter xylinus E25 using a distance-based phylogenetic tree. The top five mined industrial workhorses along with recommended information about culture conditions and a gene editing tool of CRISPRi system were provided for further engineering. (b) For gas vesicle-producer mining, 489 strains were detected, and 191 strains are industrial microorganisms. BBSniffer scored the distance between the mined industrial microorganisms and the reference strains of Halobacterium salinarum 91-R6 using a distance-based phylogenetic tree. The top five mined industrial workhorses along with recommended information about culture conditions and a gene editing tool of CRISPR-Cas system were provided for further engineering of gas vesicle. (c) For bacterial microcompartment mining, 4,241 strains were detected, and 1,381 strains are industrial microorganisms. BBSniffer scored the distance between the mined industrial microorganisms and the reference strains of pathogenic Salmonella typhimurium LT2 by a distance-based phylogenetic tree. The top five mined industrial workhorses along with recommended information about culture conditions and a gene editing tool of CRISPR-Cas system were provided for further engineering of bacterial microcompartment.
Extended Data Fig. 2 Quantitative analysis of the amount of CgCLP fiber for engineered C. glutamicum using the whole-cell filtration ELISA.
(a) ELISA quantification analysis (detection by anti-Spa2 antibody) of the cells defective for Spa1 (Δspa1 strain), Spa3 (Δspa3 strain), or both (Δspa1Δspa3 strain) and the cells overexpression of Spa2 (Spa2 strain). P values are indicated above the bars (Not significant (NS) P > 0.05, **P < 0.01, ***P < 0.001; engineered C. glutamicum versus wild-type C. glutamicum using two-tailed t-test, from three biologically independent samples, mean ± s.d.). (b) ELISA quantification analysis of the engineered cells of Spa2 fused with various domains. (c) ELISA quantification analysis of the engineered cells for validation of the co-assembly of split-Venus components into the CgCLP fibers. (d) ELISA quantification analysis of the engineered cells of TrEgl+SdBgl (the case of simultaneously secreted free enzymes of TrEgl and SdBgl) and TrEgl-Spa2+SdBgl-Spa2 (the case of the TrEgl and SdBgl enzymes were coassembled into the CgCLP fibers). All samples were cultivated in an incubator at 30 °C without shaking for 2 days in M63 medium and measured the normalized CLP production for each engineered C. glutamicum (OD600 = 0.1, 25 μL), using the amount of CLP produced by wild-type C. glutamicum as a benchmark. N=3 biologically independent experiments were performed in b, c and d; data are presented as mean ± s.d.
Extended Data Fig. 3 Probing the structure and essential residues of Spa2 for CLP assembly in C. glutamicum.
(a) The X-ray crystal structure of Spa2 is arranged in three tandem Ig-like domains, N-domain (pink), M-domain (blue), and C-domain (green). Residues involved in the formation of three intramolecular isopeptide bonds (yellow) and two disulfide bonds (red) are shown as sticks. b, c, Three intramolecular isopeptide bonds (b) and two disulfide bonds (c) in Spa2 monomer are rendered from the Spa2 structure with a 2Fo-Fc electron-density map contoured at 1.0 σ. Hydrogen bonds are shown as black dashed lines. d, e, Genetic manipulation in Δspa2 strains (harboring a plasmid that expressed Spa2 or Spa2 variants of K194A, LPLTG474LALAA478, E158A, D246A, E435A, D246A/E435A, C97A, C380A, and C97A/C380A, respectively) to assess the key residues promoting the formation of inter- and intramolecular isopeptide bonds, and disulfide bonds, in Spa2 by TEM bio-imaging (d) and quantitative analysis of the amount of CgCLP fiber by whole-cell filtration ELISA (detection by anti-Spa2 antibody) (e). P values are indicated above the bars. (Not significant (NS) P > 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; Spa2 mutated strains versus the Spa2 using two-tailed t-test, from three biologically independent samples, mean ± s.d.). The bars in the TEM images are 200 nm.
Extended Data Fig. 4 Identification of the disulfide bonds and intramolecular isopeptide bonds formation at appropriate sequence locations in Spa2 by LC–MS/MS analysis.
(a) The cartoon shows the critical features in Spa2, including three intramolecular isopeptide bonds in individual domains, two disulfide bonds in the N-domain (C97-C128) and the C-domain (C380-C432), the pilin motif of YPKN in N-domain, and the sortase cleavage sorting signal motif of LPLTG in C-domain. (b) MS/MS spectrum of the peptide with m/z 1407.44+ generated from pepsin digest of Spa2 containing the disulfide bond between Cys97 and Cys128. (c) MS/MS spectrum of the peptide with m/z 1583.72+ generated from pepsin digest of Spa2 containing the disulfide bond between Cys380 and Cys432. (d) MS/MS spectrum of the peptide with m/z 1326.94+ generated from pepsin digest of Spa2 containing the Internal isopeptide bond between Lys57 and Asn195. (e) MS/MS spectrum of the peptide with m/z 1324.63+ generated from pepsin digest of Spa2 containing the Internal isopeptide bond between Lys203 and Asn318. (f) MS/MS spectrum of the peptide with m/z 754.64+ generated from pepsin digest of Spa2 containing the Internal isopeptide bond between Lys355 and Asp466. For (b)–(f), predicted b- and y-type ions (not all included) are listed above and below the peptide sequence, respectively; the disulfide bonds and intramolecular isopeptide bonds are shown as red and yellow bars, respectively.
Extended Data Fig. 5 Characterization of the CgCLP fibers mineralized with CdS nanoparticles.
(a) The immunogold labelling and TEM images of CLP fibers. (b) TEM images of the CdS-mineralized CLP fibers. (c) HRTEM images of the nanofiber/CdS composites in the mineralized CLP fibers. The mineralized CLP fibers exhibited a clear space lattice of 0.36 nm for the CdS nanoparticles. The bars are shown in the panels.
Extended Data Fig. 6 Functional characterization of engineered CgCLP with various fusion domains.
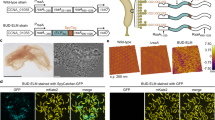
(a)TEM images showed that Ni-NTA-decorated AuNPs were anchored onto 6His-Spa2 fibers. (b) Confocal microscopic images showed the green fluorescence emitted from SpyTag-Spa2 cells to which SpyCatcher-EGFP protein binding partners were covalently attached via Spytag-SpyCatcher interaction pairs. (c) Confocal microscopic images show the green fluorescence emitted from SpyCatcher-Spa2 cells to which SpyTag-EGFP protein binding partners were covalently attached via Spytag-SpyCatcher interaction pairs. (d) Fluorescent images and quantification analysis of the immobilization ability of Mfp3Spep-Spa2 cells. Immobilized microspheres (left) on the substrates before (top) and after (bottom) challenge with water jetting at a constant discharge pressure of 5 psi. Quantification analysis of the relative capabilities of different cells (right) with immobilized PS microspheres on the substrate. The number of immobilized microspheres was set to 100% before subjecting them to mechanical challenge with water jetting. P values are indicated above the bars. Not significant (NS) P > 0.05, ***P < 0.001; Mfp3Spep-Spa2 strain versus Spa2 strain using two-tailed t-test, from three biologically independent samples, mean ± s.d. (e) Confocal microscopic images show the green fluorescence emitted from Venus-Spa2 cells. (f) Endo-1,4-β-glucanase activity of CcEgl-Spa2 cells. The P value of CcEgl-Spa2 strain versus Spa2 strain is P = 7.0E-10. ****P < 0.0001. Statistically significant differences were calculated by using a two-tailed t-test, from three biologically independent samples, mean ± s.d. Scale bars, 200 nm in a, 2 μm in b, c, and e, 100μm in d.
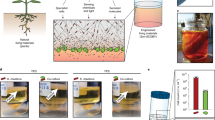
Extended Data Fig. 7 Engineering C. glutamicum as living material for cellulose degradation and lycopene production.
(a) Construction of Δspa2Δdec chassis in which both spa2 gene (defective for CgCLP formation) and gene fragment between CEY17_RS03380 and CEY17_RS03560 (with a function of producing the precursor for lycopene production, resulting in the color change of the cells from yellow to white) were deleted. The colony PCR identification indicates that the spa2 gene and genes between CEY17_RS03380 and CEY17_RS03560 were markerless deleted. (b) Construction of P1 plasmid for lycopene production, and P2 and P3 plasmids for cellulose degradation. P1 plasmid co-expressed the dxs gene and crtEBI gene cluster under the control of an IPTG-inducible promoter; P2 plasmid simultaneously expressed Spa2 pilin fusion proteins of TrEgl-Spa2 and SdBgl-Spa2 under two independent expression cassettes with the same transcription and translation elements; P3 plasmid simultaneously expressed proteins of TrEgl and SdBgl under the same genetic parts in the P2 plasmid. (c) The red color of different engineered cells indicates the lycopene accumulation. The C001 strain showed white due to the deletion region between CEY17_RS03380 and CEY17_RS03560, which lacked the synthesis of decaprenoxanthin. The red cells of C002, C003, and C004 indicated that lycopene was successfully accumulated. The C002 cells harbor P1 plasmid; the C003 cells harbor both P1 and P2 plasmids; the C004 cells have both P1 and P3 plasmids. (d) TEM images show that cells of C003, which contain the P2 plasmid, enabled co-assembly of TrEgl and SdBgl into CgCLP structure, while the cells of C001, C002, and C004 did not. CgCLP was labeled with 10 nm gold particles by immunogold labelling. Scale bars, 200 nm.
Supplementary information
Supplementary Information
Supplementary Text, Figs. 1–21 and Tables 1–6.
Supplementary Data 1
Files including detailed information for BBSniffer-detected CLP-BGC-containing strains. Files as follows: Sheet 1, strain classification for the BBSniffer-detected CLP-BGC-containing strains, indicated as ‘Strain classification’; Sheet 2, BBSniffer-detected industrial microorganisms with information on distance scores, condition of growth and genetic manipulation tools, indicated as ‘Candidate strains’; Sheet 3, annotation results of BBSniffer predicted CLP-BGC in various strains, indicated as ‘BBSniffer predicted strains’; Sheet 4, annotation results of BBSniffer for the experimentally characterized CLP-BGC strains, indicated as ‘Identified strains’; Sheet 5, calculation the success rate of the established workflow via the results detected by BBSniffer for predicted CLP-BGC in various strains and the experimentally characterized CLP-BGC strains, indicated as ‘Calculation of success rate’.
Supplementary Data 2
Files including detailed information for BBSniffer-detected BC-BGC-containing strains. Files as follows: Sheet 1, strain classification for the BBSniffer-detected BC-BGC-containing strains, indicated as ‘Strain classification’; Sheet 2, BBSniffer-detected industrial microorganisms with information on distance scores, condition of growth and genetic manipulation tools, indicated as ‘Candidate strains’; Sheet 3, annotation results of BBSniffer predicted BC-BGC in various strains, indicated as ‘BBSniffer predicted strains’; Sheet 4, annotation results of BBSniffer for the experimentally characterized BC-BGC strains, indicated as ‘Identified strains’; Sheet 5, calculation the success rate of the established workflow via the results detected by BBSniffer for predicted BC-BGC in various strains and the experimentally characterized BC-BGC strains, indicated as ‘Calculation of success rate’.
Supplementary Data 3
Files including detailed information for BBSniffer-detected GV-BGC-containing strains. Files as follows: Sheet 1, strain classification for the BBSniffer-detected GV-BGC-containing strains, indicated as ‘Strain classification’; Sheet 2, BBSniffer-detected industrial microorganisms with information on distance scores, condition of growth and genetic manipulation tools, indicated as ‘Candidate strains’; Sheet 3, annotation results of BBSniffer predicted GV-BGC in various strains, indicated as ‘BBSniffer predicted strains’; Sheet 4, annotation results of BBSniffer for the experimentally characterized GV-BGC strains, indicated as ‘Identified strains’; Sheet 5, calculation the success rate of the established workflow via the results detected by BBSniffer for predicted GV-BGC in various strains and the experimentally characterized GV-BGC strains, indicated as ‘Calculation of success rate’.
Supplementary Data 4
Files including detailed information for BBSniffer-detected BMC-BGC-containing strains. Files as follows: Sheet 1, strain classification for the BBSniffer-detected BMC-BGC-containing strains, indicated as ‘Strain classification’; Sheet 2, BBSniffer-detected industrial microorganisms with information on distance scores, condition of growth, and genetic manipulation tools, indicated as ‘Candidate strains’; Sheet 3, annotation results of BBSniffer predicted BMC-BGC in various strains, indicated as ‘BBSniffer predicted strains’; Sheet 4, annotation results of BBSniffer for the experimentally characterized BMC-BGC strains, indicated as ‘Identified strains’; Sheet 5, calculation the success rate of the established workflow via the results detected by BBSniffer for predicted BMC-BGC in various strains and the experimentally characterized BMC-BGC strains, indicated as ‘Calculation of success rate’.
Source data
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed gel.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, Y., Wu, Y., Hu, H. et al. Accelerating the design of pili-enabled living materials using an integrative technological workflow. Nat Chem Biol 20, 201–210 (2024). https://doi.org/10.1038/s41589-023-01489-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01489-x