Key Points
-
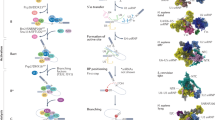
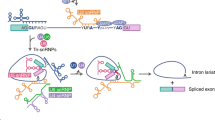
Alternative splicing is a crucial mechanism for gene regulation and for generating proteomic diversity, which allows individual genes to generate multiple mature mRNA isoforms that can be translated into functionally different proteins. Alternative splicing can be regulated at different stages of spliceosome assembly and by different mechanisms.
-
Splice site recognition of alternative exons is frequently regulated by cooperative interactions between factors such as SR (Ser–Arg) proteins and heterogeneous nuclear ribonucleoprotein particles (hnRNPs), which have lower affinities and sequence specificities. Splice site selection is also influenced by the secondary structure of mRNAs.
-
Two models have been proposed to explain the role of RNA polymerase II in alternative splicing regulation: a recruitment model and a kinetic model, and the two models are not mutually exclusive.
-
Alternative splicing, including tissue-specific alternative splicing, is an extremely common regulatory mechanism. However, the number of known sequence-specific alternative splicing factors (<50) is much smaller than that of sequence-specific transcription factors (∼2,500). Although more specific splicing factors will undoubtedly be discovered, this disparity suggests important differences in the pathways regulating transcription and splicing.
-
Core spliceosomal proteins can also regulate tissue-specific alternative splicing. This may reflect differential sensitivity of alternative exons to these factors and/or differential accumulation of the factors in different tissues.
-
Post-translational modifications of splicing factors enable cells to switch between alternative splicing isoforms rapidly after environmental stimuli. Phosphorylation can change the intracellular localization of splicing factor, protein–protein and protein–RNA interactions and even intrinsic splicing factor activity.
Abstract
Alternative splicing of mRNA precursors provides an important means of genetic control and is a crucial step in the expression of most genes. Alternative splicing markedly affects human development, and its misregulation underlies many human diseases. Although the mechanisms of alternative splicing have been studied extensively, until the past few years we had not begun to realize fully the diversity and complexity of alternative splicing regulation by an intricate protein–RNA network. Great progress has been made by studying individual transcripts and through genome-wide approaches, which together provide a better picture of the mechanistic regulation of alternative pre-mRNA splicing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
12 October 2009
In the version of this article initially published online the binding sequences for hnRNP G and hnRNP Q in Table 1 were incorrect. This error has been corrected for the print, HTML and PDF versions of the article.
References
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Wahl, M. C., Will, C. L. & Luhrmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009).
Berglund, J. A., Chua, K., Abovich, N., Reed, R. & Rosbash, M. The splicing factor BBP interacts specifically with the pre-mRNA branchpoint sequence UACUAAC. Cell 89, 781–787 (1997).
Zamore, P. D. & Green, M. R. Identification, purification, and biochemical characterization of U2 small nuclear ribonucleoprotein auxiliary factor. Proc. Natl Acad. Sci. USA 86, 9243–9247 (1989).
Nelson, K. K. & Green, M. R. Mammalian U2 snRNP has a sequence-specific RNA-binding activity. Genes Dev. 3, 1562–1571 (1989).
Graveley, B. R. Sorting out the complexity of SR protein functions. RNA 6, 1197–1211 (2000).
Tacke, R. & Manley, J. L. Determinants of SR protein specificity. Curr. Opin. Cell Biol. 11, 358–362 (1999).
Long, J. C. & Caceres, J. F. The SR protein family of splicing factors: master regulators of gene expression. Biochem. J. 417, 15–27 (2009).
Smith, C. W. & Valcarcel, J. Alternative pre-mRNA splicing: the logic of combinatorial control. Trends Biochem. Sci. 25, 381–388 (2000).
Dreyfuss, G., Kim, V. N. & Kataoka, N. Messenger-RNA-binding proteins and the messages they carry. Nature Rev. Mol. Cell Biol. 3, 195–205 (2002).
Ule, J. et al. An RNA map predicting Nova-dependent splicing regulation. Nature 444, 580–586 (2006).
Hui, J. et al. Intronic CA-repeat and CA-rich elements: a new class of regulators of mammalian alternative splicing. EMBO J. 24, 1988–1998 (2005).
Yeo, G. W. et al. An RNA code for the FOX2 splicing regulator revealed by mapping RNA-protein interactions in stem cells. Nature Struct. Mol. Biol. 16, 130–137 (2009).
Mauger, D. M., Lin, C. & Garcia-Blanco, M. A. hnRNP H and hnRNP F complex with Fox2 to silence fibroblast growth factor receptor 2 exon IIIc. Mol. Cell. Biol. 28, 5403–5419 (2008).
House, A. E. & Lynch, K. W. An exonic splicing silencer represses spliceosome assembly after ATP-dependent exon recognition. Nature Struct. Mol. Biol. 13, 937–944 (2006).
Sharma, S., Kohlstaedt, L. A., Damianov, A., Rio, D. C. & Black, D. L. Polypyrimidine tract binding protein controls the transition from exon definition to an intron defined spliceosome. Nature Struct. Mol. Biol. 15, 183–191 (2008). This study shows that PTB inhibits SRC exon N1 inclusion by preventing the transition from an exon-definition to an intron-definition complex and analyses the protein composition of different complexes.
Lallena, M. J., Chalmers, K. J., Llamazares, S., Lamond, A. I. & Valcarcel, J. Splicing regulation at the second catalytic step by Sex-lethal involves 3′ splice site recognition by SPF45. Cell 109, 285–296 (2002).
Batsche, E., Yaniv, M. & Muchardt, C. The human SWI/SNF subunit Brm is a regulator of alternative splicing. Nature Struct. Mol. Biol. 13, 22–29 (2006). This study shows that BRM promotes the inclusion of variable exons of CD44 pre-mRNA by stalling RNAP II at the variable exon-containing region of the CD44 gene. It also shows that BRM interacts with splicing factor SAM68.
de la Mata, M. & Kornblihtt, A. R. RNA polymerase II C-terminal domain mediates regulation of alternative splicing by SRp20. Nature Struct. Mol. Biol. 13, 973–980 (2006).
Sims, R. J. 3rd et al. Recognition of trimethylated histone H3 lysine 4 facilitates the recruitment of transcription postinitiation factors and pre-mRNA splicing. Mol. Cell 28, 665–676 (2007).
Lin, S., Coutinho-Mansfield, G., Wang, D., Pandit, S. & Fu, X. D. The splicing factor SC35 has an active role in transcriptional elongation. Nature Struct. Mol. Biol. 15, 819–826 (2008).
Moldon, A. et al. Promoter-driven splicing regulation in fission yeast. Nature 455, 997–1000 (2008).
Graveley, B. R. Alternative splicing: increasing diversity in the proteomic world. Trends Genet. 17, 100–107 (2001).
Blencowe, B. J. & Graveley, B. R. (eds) Alternative Splicing in the Postgenomic Era. (Springer, the Netherlands, 2007).
Grabowski, P. J. & Black, D. L. Alternative RNA splicing in the nervous system. Prog. Neurobiol. 65, 289–308 (2001).
Park, J. W., Parisky, K., Celotto, A. M., Reenan, R. A. & Graveley, B. R. Identification of alternative splicing regulators by RNA interference in Drosophila. Proc. Natl Acad. Sci. USA 101, 15974–15979 (2004).
Zhang, Z. et al. SMN deficiency causes tissue-specific perturbations in the repertoire of snRNAs and widespread defects in splicing. Cell 133, 585–600 (2008). This paper shows that SMN deficiency regulates snRNP levels in a tissue-specific manner, and this was reflected in altered alternative splicing patterns in different mouse tissues.
Blencowe, B. J. Alternative splicing: new insights from global analyses. Cell 126, 37–47 (2006).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nature Genet. 40, 1413–1415 (2008).
Castle, J. C. et al. Expression of 24,426 human alternative splicing events and predicted cis regulation in 48 tissues and cell lines. Nature Genet. 40, 1416–1425 (2008). A transcriptome study that analyses alternative splicing events from 48 tissues and identifies tissue-specific regulatory motifs and cognate binding proteins.
Sultan, M. et al. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science 321, 956–960 (2008).
Sterner, D. A., Carlo, T. & Berget, S. M. Architectural limits on split genes. Proc. Natl Acad. Sci. USA 93, 15081–15085 (1996).
Berget, S. M. Exon recognition in vertebrate splicing. J. Biol. Chem. 270, 2411–2414 (1995).
Lim, S. R. & Hertel, K. J. Commitment to splice site pairing coincides with A complex formation. Mol. Cell 15, 477–483 (2004).
Kotlajich, M. V., Crabb, T. L. & Hertel, K. J. Spliceosome assembly pathways for different types of alternative splicing converge during commitment to splice site pairing in the A complex. Mol. Cell. Biol. 29, 1072–1082 (2009). This study shows that the commitment to splicing of some alternative exons occurs during splice site pairing in the A complex and that ATP hydrolysis is required for splice site paring, thereby locking splice sites into a splicing pattern after U2 snRNP binding to the branch site.
Bourgeois, C. F., Popielarz, M., Hildwein, G. & Stevenin, J. Identification of a bidirectional splicing enhancer: differential involvement of SR proteins in 5′ or 3′ splice site activation. Mol. Cell. Biol. 19, 7347–7356 (1999).
Zuo, P. & Maniatis, T. The splicing factor U2AF35 mediates critical protein–protein interactions in constitutive and enhancer-dependent splicing. Genes Dev. 10, 1356–1368 (1996).
Feng, Y., Chen, M. & Manley, J. L. Phosphorylation switches the general splicing repressor SRp38 to a sequence-specific activator. Nature Struct. Mol. Biol. 15, 1040–1048 (2008). This paper shows that phosphorylation switches SRp38 from a general repressor to a sequence-specific activator that functions by recruiting and stabilizing U1 and U2 snRNP at splice sites.
Graveley, B. R., Hertel, K. J. & Maniatis, T. The role of U2AF35 and U2AF65 in enhancer-dependent splicing. RNA 7, 806–818 (2001).
Kohtz, J. D. et al. Protein–protein interactions and 5′-splice-site recognition in mammalian mRNA precursors. Nature 368, 119–124 (1994).
Wu, J. Y. & Maniatis, T. Specific interactions between proteins implicated in splice site selection and regulated alternative splicing. Cell 75, 1061–1070 (1993).
Xiao, S. H. & Manley, J. L. Phosphorylation of the ASF/SF2 RS domain affects both protein-protein and protein-RNA interactions and is necessary for splicing. Genes Dev. 11, 334–344 (1997).
Pacheco, T. R., Coelho, M. B., Desterro, J. M., Mollet, I. & Carmo-Fonseca, M. In vivo requirement of the small subunit of U2AF for recognition of a weak 3′ splice site. Mol. Cell. Biol. 26, 8183–90 (2006).
Longman, D. et al. Multiple interactions between SRm160 and SR family proteins in enhancer-dependent splicing and development of C. elegans. Curr. Biol. 11, 1923–1933 (2001).
Blencowe, B. J. et al. The SRm160/300 splicing coactivator subunits. RNA 6, 111–120 (2000).
Tacke, R. & Manley, J. L. Functions of SR and Tra2 proteins in pre-mRNA splicing regulation. Proc. Soc. Exp. Biol. Med. 220, 59–63 (1999).
Izquierdo, J. M. et al. Regulation of Fas alternative splicing by antagonistic effects of TIA-1 and PTB on exon definition. Mol. Cell 19, 475–484 (2005).
Forch, P., Puig, O., Martinez, C., Seraphin, B. & Valcarcel, J. The splicing regulator TIA-1 interacts with U1-C to promote U1 snRNP recruitment to 5′ splice sites. EMBO J. 21, 6882–6892 (2002).
Tisserant, A. & Konig, H. Signal-regulated pre-mRNA occupancy by the general splicing factor U2AF. PLoS ONE 3, e1418 (2008).
Yang, L., Embree, L. J., Tsai, S. & Hickstein, D. D. Oncoprotein TLS interacts with serine-arginine proteins involved in RNA splicing. J. Biol. Chem. 273, 27761–27764 (1998).
Komatsu, M., Kominami, E., Arahata, K. & Tsukahara, T. Cloning and characterization of two neural-salient serine/arginine-rich (NSSR) proteins involved in the regulation of alternative splicing in neurones. Genes Cells 4, 593–606 (1999).
Cowper, A. E., Caceres, J. F., Mayeda, A. & Screaton, G. R. Serine-arginine (SR) protein-like factors that antagonize authentic SR proteins and regulate alternative splicing. J. Biol. Chem. 276, 48908–48914 (2001).
Shin, C. & Manley, J. L. The SR protein SRp38 represses splicing in M phase cells. Cell 111, 407–417 (2002).
Shin, C., Feng, Y. & Manley, J. L. Dephosphorylated SRp38 acts as a splicing repressor in response to heat shock. Nature 427, 553–558 (2004).
Krainer, A. R., Conway, G. C. & Kozak, D. Purification and characterization of pre-mRNA splicing factor SF2 from HeLa cells. Genes Dev. 4, 1158–1171 (1990).
Singh, R., Valcarcel, J. & Green, M. R. Distinct binding specificities and functions of higher eukaryotic polypyrimidine tract-binding proteins. Science 268, 1173–1176 (1995).
Spellman, R. & Smith, C. W. Novel modes of splicing repression by PTB. Trends Biochem. Sci. 31, 73–76 (2006).
Sauliere, J., Sureau, A., Expert-Bezancon, A. & Marie, J. The polypyrimidine tract binding protein (PTB) represses splicing of exon 6B from the β-tropomyosin pre-mRNA by directly interfering with the binding of the U2AF65 subunit. Mol. Cell. Biol. 26, 8755–8769 (2006).
Tange, T. O., Damgaard, C. K., Guth, S., Valcarcel, J. & Kjems, J. The hnRNP A1 protein regulates HIV-1 tat splicing via a novel intron silencer element. EMBO J. 20, 5748–5758 (2001).
Zhou, H. L. & Lou, H. Repression of prespliceosome complex formation at two distinct steps by Fox-1/Fox-2 proteins. Mol. Cell. Biol. 28, 5507–5516 (2008).
Zhu, H., Hinman, M. N., Hasman, R. A., Mehta, P. & Lou, H. Regulation of neuron-specific alternative splicing of neurofibromatosis type 1 pre-mRNA. Mol. Cell. Biol. 28, 1240–1251 (2008).
Kashima, T. & Manley, J. L. A negative element in SMN2 exon 7 inhibits splicing in spinal muscular atrophy. Nature Genet. 34, 460–463 (2003).
Martins de Araujo, M., Bonnal, S., Hastings, M. L., Krainer, A. R. & Valcarcel, J. Differential 3′ splice site recognition of SMN1 and SMN2 transcripts by U2AF and U2 snRNP. RNA 15, 515–523 (2009).
Sharma, S., Falick, A. M. & Black, D. L. Polypyrimidine tract binding protein blocks the 5′ splice site-dependent assembly of U2AF and the prespliceosomal E complex. Mol. Cell 19, 485–496 (2005).
Damgaard, C. K., Tange, T. O. & Kjems, J. hnRNP A1 controls HIV-1 mRNA splicing through cooperative binding to intron and exon splicing silencers in the context of a conserved secondary structure. RNA 8, 1401–1415 (2002).
Nasim, F. U., Hutchison, S., Cordeau, M. & Chabot, B. High-affinity hnRNP A1 binding sites and duplex-forming inverted repeats have similar effects on 5′ splice site selection in support of a common looping out and repression mechanism. RNA 8, 1078–1089 (2002).
Hutchison, S., LeBel, C., Blanchette, M. & Chabot, B. Distinct sets of adjacent heterogeneous nuclear ribonucleoprotein (hnRNP) A1/A2 binding sites control 5′ splice site selection in the hnRNP A1 mRNA precursor. J. Biol. Chem. 277, 29745–29752 (2002).
Kashima, T., Rao, N., David, C. J. & Manley, J. L. hnRNP A1 functions with specificity in repression of SMN2 exon 7 splicing. Hum. Mol. Genet. 16, 3149–3159 (2007).
Kashima, T., Rao, N. & Manley, J. L. An intronic element contributes to splicing repression in spinal muscular atrophy. Proc. Natl Acad. Sci. USA 104, 3426–3431 (2007).
Chou, M. Y., Underwood, J. G., Nikolic, J., Luu, M. H. & Black, D. L. Multisite RNA binding and release of polypyrimidine tract binding protein during the regulation of c-src neural-specific splicing. Mol. Cell 5, 949–957 (2000).
Mayeda, A., Helfman, D. M. & Krainer, A. R. Modulation of exon skipping and inclusion by heterogeneous nuclear ribonucleoprotein A1 and pre-mRNA splicing factor SF2/ASF. Mol. Cell. Biol. 13, 2993–3001 (1993).
Zahler, A. M., Damgaard, C. K., Kjems, J. & Caputi, M. SC35 and heterogeneous nuclear ribonucleoprotein A/B proteins bind to a juxtaposed exonic splicing enhancer/exonic splicing silencer element to regulate HIV-1 tat exon 2 splicing. J. Biol. Chem. 279, 10077–10084 (2004).
Zhu, J. & Krainer, A. R. Pre-mRNA splicing in the absence of an SR protein RS domain. Genes Dev. 14, 3166–3178 (2000).
Crawford, J. B. & Patton, J. G. Activation of α-tropomyosin exon 2 is regulated by the SR protein 9G8 and heterogeneous nuclear ribonucleoproteins H and F. Mol. Cell. Biol. 26, 8791–802 (2006).
Expert-Bezancon, A. et al. hnRNP A1 and the SR proteins ASF/SF2 and SC35 have antagonistic functions in splicing of β-tropomyosin exon 6B. J. Biol. Chem. 279, 38249–59 (2004).
Charlet, B. N., Logan, P., Singh, G. & Cooper, T. A. Dynamic antagonism between ETR-3 and PTB regulates cell type-specific alternative splicing. Mol. Cell 9, 649–658 (2002).
Blanchette, M. et al. Genome-wide analysis of alternative pre-mRNA splicing and RNA-binding specificities of the Drosophila hnRNP A/B family members. Mol. Cell 33, 438–449 (2009).
Hung, L. H. et al. Diverse roles of hnRNP L in mammalian mRNA processing: a combined microarray and RNAi analysis. RNA 14, 284–296 (2008).
Licatalosi, D. D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008). This paper introduces a method that combines CLIP and high-throughput sequencing to identify targets of NOVA proteins.
Dredge, B. K., Stefani, G., Engelhard, C. C. & Darnell, R. B. Nova autoregulation reveals dual functions in neuronal splicing. EMBO J. 24, 1608–1620 (2005).
Martinez-Contreras, R. et al. Intronic binding sites for hnRNP A/B and hnRNP F/H proteins stimulate pre-mRNA splicing. PLoS Biol. 4, e21 (2006).
Wang, Z. & Burge, C. B. Splicing regulation: from a parts list of regulatory elements to an integrated splicing code. RNA 14, 802–813 (2008).
Dredge, B. K. & Darnell, R. B. Nova regulates GABAA receptor γ2 alternative splicing via a distal downstream UCAU-rich intronic splicing enhancer. Mol. Cell. Biol. 23, 4687–4700 (2003).
Schaub, M. C., Lopez, S. R. & Caputi, M. Members of the heterogeneous nuclear ribonucleoprotein H family activate splicing of an HIV-1 splicing substrate by promoting formation of ATP-dependent spliceosomal complexes. J. Biol. Chem. 282, 13617–13626 (2007).
Caputi, M. & Zahler, A. M. Determination of the RNA binding specificity of the heterogeneous nuclear ribonucleoprotein (hnRNP) H/H′/F/2H9 family. J. Biol. Chem. 276, 43850–43859 (2001).
Sanford, J. R. et al. Identification of nuclear and cytoplasmic mRNA targets for the shuttling protein SF2/ASF. PLoS ONE 3, e3369 (2008).
Sanford, J. R. et al. Splicing factor SFRS1 recognizes a functionally diverse landscape of RNA transcripts. Genome Res. 19, 381–394 (2009).
Graveley, B. R. Mutually exclusive splicing of the insect Dscam pre-mRNA directed by competing intronic RNA secondary structures. Cell 123, 65–73 (2005).
Olson, S. et al. A regulator of Dscam mutually exclusive splicing fidelity. Nature Struct. Mol. Biol. 14, 1134–1140 (2007).
Grover, A. et al. 5′ splice site mutations in tau associated with the inherited dementia FTDP-17 affect a stem-loop structure that regulates alternative splicing of exon 10. J. Biol. Chem. 274, 15134–15143 (1999).
Hiller, M., Zhang, Z., Backofen, R. & Stamm, S. Pre-mRNA secondary structures influence exon recognition. PLoS Genet. 3, e204 (2007).
Camats, M., Guil, S., Kokolo, M. & Bach-Elias, M. P68 RNA helicase (DDX5) alters activity of cis- and trans-acting factors of the alternative splicing of H-Ras. PLoS ONE 3, e2926 (2008).
Libri, D., Balvay, L. & Fiszman, M. Y. In vivo splicing of the beta tropomyosin pre-mRNA: a role for branch point and donor site competition. Mol. Cell. Biol. 12, 3204–3215 (1992).
Henkin, T. M. Riboswitch RNAs: using RNA to sense cellular metabolism. Genes Dev. 22, 3383–3390 (2008).
Cheah, M. T., Wachter, A., Sudarsan, N. & Breaker, R. R. Control of alternative RNA splicing and gene expression by eukaryotic riboswitches. Nature 447, 497–500 (2007).
Kishore, S. & Stamm, S. Regulation of alternative splicing by snoRNAs. Cold Spring Harb. Symp. Quant. Biol. 71, 329–334 (2006).
Kishore, S. & Stamm, S. The snoRNA HBII-52 regulates alternative splicing of the serotonin receptor 2C. Science 311, 230–232 (2006).
Yu, Y. et al. Dynamic regulation of alternative splicing by silencers that modulate 5′ splice site competition. Cell 135, 1224–1236 (2008). The authors screen for splicing silencers that favour the inclusion of a distal 5′ splice site and provide evidence that the silencers work by changing the conformation of the pre-mRNA–U1 snRNP complex and promoting pairing of the distal U1 snRNP with U2 snRNP.
Makeyev, E. V., Zhang, J., Carrasco, M. A. & Maniatis, T. The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 27, 435–448 (2007).
Boutz, P. L. et al. A post-transcriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons. Genes Dev. 21, 1636–1652 (2007). Using siRNA and microarray analysis, the authors show that the PTB-to-nPTB switch provides a post-transcriptional mechanism that is important for programming neuronal differentiation.
Coutinho-Mansfield, G. C., Xue, Y., Zhang, Y. & Fu, X. D. PTB/nPTB switch: a post-transcriptional mechanism for programming neuronal differentiation. Genes Dev. 21, 1573–1577 (2007).
Spellman, R., Llorian, M. & Smith, C. W. Crossregulation and functional redundancy between the splicing regulator PTB and its paralogs nPTB and ROD1. Mol. Cell 27, 420–434 (2007).
Ohi, M. D. et al. Structural and functional analysis of essential pre-mRNA splicing factor Prp19p. Mol. Cell. Biol. 25, 451–460 (2005).
Bessonov, S., Anokhina, M., Will, C. L., Urlaub, H. & Luhrmann, R. Isolation of an active step I spliceosome and composition of its RNP core. Nature 452, 846–850 (2008).
Eldridge, A. G., Li, Y., Sharp, P. A. & Blencowe, B. J. The SRm160/300 splicing coactivator is required for exon-enhancer function. Proc. Natl Acad. Sci. USA 96, 6125–6130 (1999).
Edamatsu, H., Kaziro, Y. & Itoh, H. LUCA15, a putative tumour suppressor gene encoding an RNA-binding nuclear protein, is down-regulated in ras-transformed Rat-1 cells. Genes Cells 5, 849–858 (2000).
Mourtada-Maarabouni, M., Sutherland, L. C. & Williams, G. T. Candidate tumour suppressor LUCA-15 can regulate multiple apoptotic pathways. Apoptosis 7, 421–432 (2002).
Bonnal, S. et al. RBM5/Luca-15/H37 regulates Fas alternative splice site pairing after exon definition. Mol. Cell 32, 81–95 (2008).
Kornblihtt, A. R. Chromatin, transcript elongation and alternative splicing. Nature Struct. Mol. Biol. 13, 5–7 (2006).
Das, R. et al. SR proteins function in coupling RNAP II transcription to pre-mRNA splicing. Mol. Cell 26, 867–881 (2007).
Auboeuf, D. et al. Differential recruitment of nuclear receptor coactivators may determine alternative RNA splice site choice in target genes. Proc. Natl Acad. Sci. USA 101, 2270–2274 (2004).
Cramer, P. et al. Coupling of transcription with alternative splicing: RNA pol II promoters modulate SF2/ASF and 9G8 effects on an exonic splicing enhancer. Mol. Cell 4, 251–258 (1999).
Monsalve, M. et al. Direct coupling of transcription and mRNA processing through the thermogenic coactivator PGC-1. Mol. Cell 6, 307–316 (2000).
de la Mata, M. et al. A slow RNA polymerase II affects alternative splicing in vivo. Mol. Cell 12, 525–532 (2003).
Munoz, M. J. et al. DNA damage regulates alternative splicing through inhibition of RNA polymerase II elongation. Cell 137, 708–720 (2009).
Phatnani, H. P. & Greenleaf, A. L. Phosphorylation and functions of the RNA polymerase II CTD. Genes Dev. 20, 2922–2936 (2006).
Lin, P. S., Marshall, N. F. & Dahmus, M. E. CTD phosphatase: role in RNA polymerase II cycling and the regulation of transcript elongation. Prog. Nucleic Acid Res. Mol. Biol. 72, 333–365 (2002).
David, C. J. & Manley, J. L. The search for alternative splicing regulators: new approaches offer a path to a splicing code. Genes Dev. 22, 279–285 (2008).
Ohno, G., Hagiwara, M. & Kuroyanagi, H. STAR family RNA-binding protein ASD-2 regulates developmental switching of mutually exclusive alternative splicing in vivo. Genes Dev. 22, 360–374 (2008).
Li, Q., Lee, J. A. & Black, D. L. Neuronal regulation of alternative pre-mRNA splicing. Nature Rev. Neurosci. 8, 819–831 (2007).
Ule, J. et al. Nova regulates brain-specific splicing to shape the synapse. Nature Genet. 37, 844–852 (2005).
Perrone-Bizzozero, N. & Bolognani, F. Role of HuD and other RNA-binding proteins in neural development and plasticity. J. Neurosci. Res. 68, 121–126 (2002).
Zhu, H., Hasman, R. A., Barron, V. A., Luo, G. & Lou, H. A nuclear function of Hu proteins as neuron-specific alternative RNA processing regulators. Mol. Biol. Cell 17, 5105–5114 (2006).
Soller, M., Li, M. & Haussmann, I. U. Regulation of the ELAV target ewg: insights from an evolutionary perspective. Biochem. Soc. Trans. 36, 502–504 (2008).
McKee, A. E. et al. A genome-wide in situ hybridization map of RNA-binding proteins reveals anatomically restricted expression in the developing mouse brain. BMC Dev. Biol. 5, 14 (2005).
Yang, Y. Y., Yin, G. L. & Darnell, R. B. The neuronal RNA-binding protein Nova-2 is implicated as the autoantigen targeted in POMA patients with dementia. Proc. Natl Acad. Sci. USA 95, 13254–13259 (1998).
Warzecha, C. C., Sato, T. K., Nabet, B., Hogenesch, J. B. & Carstens, R. P. ESRP1 and ESRP2 are epithelial cell-type-specific regulators of FGFR2 splicing. Mol. Cell 33, 591–601 (2009). The authors identify ESRP1 and ESRP2 as epithelial cell-specific alternative splicing factors and find that they regulate several epithelial cell-specific exons. They also show that ESRP1 and ESRP2 expression levels correlate with the splicing pattern change that is observed during the epithelial-to-mesenchymal cell transition.
Ding, J. H. et al. Dilated cardiomyopathy caused by tissue-specific ablation of SC35 in the heart. EMBO J. 23, 885–896 (2004).
Xu, X. et al. ASF/SF2-regulated CaMKIIδ alternative splicing temporally reprograms excitation-contraction coupling in cardiac muscle. Cell 120, 59–72 (2005).
Feng, Y. et al. SRp38 regulates alternative splicing and is required for Ca2+ handling in the embryonic heart. Dev. Cell 16, 528–538 (2009). This study shows that mice lacking SRp38 die perinatally and have multiple cardiac defects, and that the mRNA encoding cardiac triadin is a direct target of SRp38.
Grosso, A. R. et al. Tissue-specific splicing factor gene expression signatures. Nucleic Acids Res. 36, 4823–4832 (2008).
Massiello, A., Roesser, J. R. & Chalfant, C. E. SAP155 binds to ceramide-responsive RNA cis-element 1 and regulates the alternative 5′ splice site selection of Bcl-x pre-mRNA. FASEB J. 20, 1680–1682 (2006).
Pacheco, T. R., Moita, L. F., Gomes, A. Q., Hacohen, N. & Carmo-Fonseca, M. RNA interference knockdown of hU2AF35 impairs cell cycle progression and modulates alternative splicing of Cdc25 transcripts. Mol. Biol. Cell 17, 4187–4199 (2006).
Pleiss, J. A., Whitworth, G. B., Bergkessel, M. & Guthrie, C. Rapid, transcript-specific changes in splicing in response to environmental stress. Mol. Cell 27, 928–937 (2007).
Monani, U. R. Spinal muscular atrophy: a deficiency in a ubiquitous protein; a motor neuron-specific disease. Neuron 48, 885–896 (2005).
Gabanella, F. et al. Ribonucleoprotein assembly defects correlate with spinal muscular atrophy severity and preferentially affect a subset of spliceosomal snRNPs. PLoS ONE 2, e921 (2007).
Shin, C. & Manley, J. L. Cell signalling and the control of pre-mRNA splicing. Nature Rev. Mol. Cell Biol. 5, 727–738 (2004).
Tarn, W. Y. Cellular signals modulate alternative splicing. J. Biomed. Sci. 14, 517–522 (2007).
Huang, Y., Yario, T. A. & Steitz, J. A. A molecular link between SR protein dephosphorylation and mRNA export. Proc. Natl Acad. Sci. USA 101, 9666–9670 (2004).
van der Houven van Oordt, W. et al. The MKK(3/6)-p38-signaling cascade alters the subcellular distribution of hnRNP A1 and modulates alternative splicing regulation. J. Cell Biol. 149, 307–316 (2000).
Habelhah, H. et al. ERK phosphorylation drives cytoplasmic accumulation of hnRNP-K and inhibition of mRNA translation. Nature Cell Biol. 3, 325–330 (2001).
Daoud, R. et al. Ischemia induces a translocation of the splicing factor tra2-beta 1 and changes alternative splicing patterns in the brain. J. Neurosci. 22, 5889–5899 (2002).
Guil, S., Long, J. C. & Caceres, J. F. hnRNP A1 relocalization to the stress granules reflects a role in the stress response. Mol. Cell. Biol. 26, 5744–5758 (2006).
Huang, C. J., Tang, Z., Lin, R. J. & Tucker, P. W. Phosphorylation by SR kinases regulates the binding of PTB-associated splicing factor (PSF) to the pre-mRNA polypyrimidine tract. FEBS Lett. 581, 223–232 (2007).
Tacke, R., Chen, Y. & Manley, J. L. Sequence-specific RNA binding by an SR protein requires RS domain phosphorylation: creation of an SRp40-specific splicing enhancer. Proc. Natl Acad. Sci. USA 94, 1148–1153 (1997).
Izquierdo, J. M. & Valcarcel, J. Fas-activated serine/threonine kinase (FAST K) synergizes with TIA-1/TIAR proteins to regulate Fas alternative splicing. J. Biol. Chem. 282, 1539–1543 (2007).
Paronetto, M. P., Achsel, T., Massiello, A., Chalfant, C. E. & Sette, C. The RNA-binding protein Sam68 modulates the alternative splicing of Bcl-x. J. Cell Biol. 176, 929–939 (2007).
Ma, S., Liu, G., Sun, Y. & Xie, J. Relocalization of the polypyrimidine tract-binding protein during PKA-induced neurite growth. Biochim. Biophys. Acta 1773, 912–923 (2007).
Xie, J., Lee, J. A., Kress, T. L., Mowry, K. L. & Black, D. L. Protein kinase A phosphorylation modulates transport of the polypyrimidine tract-binding protein. Proc. Natl Acad. Sci. USA 100, 8776–8781 (2003).
Shi, Y. & Manley, J. L. A complex signaling pathway regulates SRp38 phosphorylation and pre-mRNA splicing in response to heat shock. Mol. Cell 28, 79–90 (2007).
Van Eynde, A. et al. Molecular cloning of NIPP-1, a nuclear inhibitor of protein phosphatase-1, reveals homology with polypeptides involved in RNA processing. J. Biol. Chem. 270, 28068–28074 (1995).
Cohen, P. T. Protein phosphatase 1— targeted in many directions. J. Cell Sci. 115, 241–256 (2002).
Kim, E., Goren, A. & Ast, G. Alternative splicing: current perspectives. Bioessays 30, 38–47 (2008).
Hallikas, O. et al. Genome-wide prediction of mammalian enhancers based on analysis of transcription-factor binding affinity. Cell 124, 47–59 (2006).
Babu, M. M., Luscombe, N. M., Aravind, L., Gerstein, M. & Teichmann, S. A. Structure and evolution of transcriptional regulatory networks. Curr. Opin. Struct. Biol. 14, 283–291 (2004).
Tacke, R. & Manley, J. L. The human splicing factors ASF/SF2 and SC35 possess distinct, functionally significant RNA binding specificities. EMBO J. 14, 3540–3551 (1995).
Kent, O. A., Ritchie, D. B. & Macmillan, A. M. Characterization of a U2AF-independent commitment complex (E′) in the mammalian spliceosome assembly pathway. Mol. Cell. Biol. 25, 233–40 (2005).
Shen, H. & Green, M. R. RS domains contact splicing signals and promote splicing by a common mechanism in yeast through humans. Genes Dev. 20, 1755–1765 (2006).
Huang, Y., Gattoni, R., Stevenin, J. & Steitz, J. A. SR splicing factors serve as adapter proteins for TAP-dependent mRNA export. Mol. Cell 11, 837–843 (2003).
Zhang, Z. & Krainer, A. R. Involvement of SR proteins in mRNA surveillance. Mol. Cell 16, 597–607 (2004).
Sanford, J. R., Gray, N. K., Beckmann, K. & Caceres, J. F. A novel role for shuttling SR proteins in mRNA translation. Genes Dev. 18, 755–768 (2004).
Cartegni, L. & Krainer, A. R. Disruption of an SF2/ASF-dependent exonic splicing enhancer in SMN2 causes spinal muscular atrophy in the absence of SMN1. Nature Genet. 30, 377–384 (2002).
Cartegni, L., Hastings, M. L., Calarco, J. A., de Stanchina, E. & Krainer, A. R. Determinants of exon 7 splicing in the spinal muscular atrophy genes, SMN1 and SMN2. Am. J. Hum. Genet. 78, 63–77 (2006).
Acknowledgements
Work from the authors' laboratory was supported in part by grants from the National Institutes of Health. We thank C. David for comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- Small nuclear ribonucleoprotein particle
-
(snRNP). A protein, including U1, U2, U4, U5 and U6, which contains U-rich small nuclear RNAs (snRNAs) and both small nuclear ribonucleoprotein (snRNP)-specific and common proteins, and is a core component of the spliceosome.
- Branch point
-
A nucleotide, usually an adenosine, within a variably conserved branch point sequence upstream of the 3′ splice site, the 2′ hydroxyl group of which attacks the 5′ splice site in the first step of splicing.
- SR (Ser–Arg) protein family
-
A family of nuclear factors that have many important roles in splicing mRNA precursors in metazoan organisms, functioning in both constitutive and alternative splicing.
- Heterologous nuclear RNP
-
(hnRNP). A pre-mRNA- or mRNA-binding protein that associates with transcripts during or after transcription and influences their function and fate. Some hnRNPs shuttle in and out of nuclei, whereas others are constitutively nuclear.
- Alternative exon
-
An exon that is included in mature mRNA in certain cellular contexts but excluded in others.
- RS (Arg–Ser repeat-containing) domain
-
A protein domain that is variable in length and enriched in Arg–Ser dipeptides and seems to be involved in protein–protein and protein–RNA interactions.
- Hu/ELAV family protein
-
A protein belonging to a family of nervous system-specific RNA-binding proteins that specifically bind to AU-rich sequences.
- CLIP
-
A method that combines cross-linking and immunoprecipitation to identify in vivo targets of RNA-binding proteins.
- RRM domain
-
(RNA recognition motif domain). A protein domain that is frequently involved in sequence-specific single-stranded RNA binding. Also known as an RNP-type RNA-binding domain.
- 14-3-3 protein
-
A protein belonging to a family of conserved proteins that bind to phosphorylated serine and threonine residues and that are encoded by seven genes in most mammals. They bind diverse regulatory proteins, including kinases, phosphatases and transmembrane receptors.
- SELEX
-
A technique to determine the DNA or RNA sequence that is specifically recognized by a protein. The method involves multiple rounds of binding to an initially random sequence until a high-affinity consensus sequence emerges.
Rights and permissions
About this article
Cite this article
Chen, M., Manley, J. Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat Rev Mol Cell Biol 10, 741–754 (2009). https://doi.org/10.1038/nrm2777
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm2777