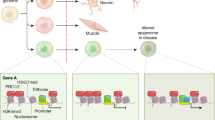
Abstract
To celebrate the first 10 years of Nature Reviews Genetics, we asked eight leading researchers for their views on the key developments in genetics and genomics in the past decade and the prospects for the future. Their responses highlight the incredible changes that the field has seen, from the explosion of genomic data and the many possibilities it has opened up to the ability to reprogramme adult cells to pluripotency. The way ahead looks similarly exciting as we address questions such as how cells function as systems and how complex interactions among genetics, epigenetics and the environment combine to shape phenotypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Margueron, R. & Reinberg, D. Chromatin structure and the inheritance of epigenetic information. Nature Rev. Genet. 11, 285–296 (2010).
Jenuwein, T. & Allis. C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Cloos, P. A., Christensen, J., Agger, K. & Helin, K. Erasing the methyl mark: histone demethylases at the center of cellular differentiation and disease. Genes Dev. 22, 1115–1140 (2008).
Baumann, K. Epigenetics: Unravelling demethylation. Nature Rev. Mol. Cell Biol. 11, 87 (2010).
Kloc, A. & Martienssen, R. RNAi, heterochromatin and the cell cycle. Trends Genet. 24, 511–517 (2008).
Okamoto, I. & Heard, E. Lessons from comparative analysis of X-chromosome inactivation in mammals. Chromosome Res. 17, 659–669 (2009).
Sulem, P. et al. Genetic determinants of hair, eye and skin pigmentation in Europeans. Nature Genet. 39, 1443–1452 (2007).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nature Genet. 42, 565–569 (2010).
Bersaglieri, T. et al. Genetic signatures of strong recent positive selection at the lactase gene. Am. J. Hum. Genet. 74, 1111–1120 (2004).
Tishkoff, S. A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nature Genet. 39, 31–40 (2007).
Pickrell, J. K. et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 19, 826–837 (2009).
Simonson, T. S. et al. Genetic evidence for high-altitude adaptation in Tibet. Science 329, 72–75 (2010).
Sabeti, P. C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Tishkoff, S. A. et al. Haplotype diversity and linkage disequilibrium at human G6PD: recent origin of alleles that confer malarial resistance. Science 293, 455–462 (2001).
Genovese, G. et al. Association of trypanolytic APOL1 variants with kidney disease in African-Americans. Science 329, 841–845 (2010).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Rosenberg, N. A. et al. Genetic structure of human populations. Science 298, 2381–2385 (2002).
Novembre, J. et al. Genes mirror geography within Europe. Nature 456, 98–101 (2008).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Wheeler, D. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008).
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Nejentsev, S., Walker, N., Riches, D., Egholm, M. & Todd, J. A. Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324, 387–389 (2009).
Lupski, J. R. Retrotransposition and structural variation in the human genome. Cell 141, 1110–1112 (2010).
Todd, J. A. D'oh! genes and environment cause Crohn's disease. Cell 141, 1114–1116 (2010).
Clayton, D. G. Prediction and interaction in complex disease genetics: experience in type 1 diabetes. PLoS Genet. 5, e1000540 (2009).
Purcell, S. M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Nadeau, J. H. Transgenerational genetic effects on phenotypic variation and disease risk. Hum. Mol. Genet. 18, R202–R210 (2009).
Amos, C. I., Spitz, M. R. & Cinciripini, P. Chipping away at the genetics of smoking behavior. Nature Genet. 42, 366–368 (2010).
Brown, M. S., Ye, J. & Goldstein, J. L. Medicine. HDL miR-ed down by SREBP introns. Science 328, 1495–1496 (2010).
Poliseno, L. et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465, 1033–1038 (2010).
Athanasopoulos, V. et al. The ROQUIN family of proteins localizes to stress granules via the ROQ domain and binds target mRNAs. FEBS J. 277, 2109–2127 (2010).
Petronis, A. Epigenetics as a unifying principle in the aetiology of complex traits and diseases. Nature 465, 721–727 (2010).
The C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Somerville, C. & Dangl, J. Plant biology in 2010. Science 290, 2077–2078 (2000).
Falk, R. The gene in search of an identity. Hum. Genet. 68, 195–204 (1984).
Johannsen, W. Elemente der exakten Erblichkeitslehre (Gustav Fischer, Jena, 1909).
Beadle, G. W. & Tatum, E. L. Genetic control of biochemical reactions in Neurospora. Proc. Natl Acad. Sci. USA 27, 499–506 (1941).
Crick, F. Central dogma of molecular biology. Nature 227, 561–563 (1970).
Feschotte, C. Transposable elements and the evolution of regulatory networks. Nature Rev. Genet. 9, 397–405 (2008).
Wagner, G. P. & Lynch, V. J. Evolutionary novelties. Current Biol. 20, R48–R52 (2010).
Oliver, K. R. & Greene, W. K. Transposable elements: powerful facilitators of evolution. BioEssays 31, 703–714 (2009).
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Waterston, R. H. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Elsik, C. G. et al. The genome sequence of taurine cattle: a window to ruminant biology and evolution. Science 324, 522–528 (2009).
Yu, J. et al. A draft sequence of the rice genome (Oryza sativa, L. ssp. indica). Science 296, 79–92 (2002).
International HapMap Consortium. The International HapMap Project. Nature 426, 789–796 (2003).
Gerdsen, F., Mueller, S., Jablonski, S. & Prokosch, H. U. Standardized exchange of medical data between a research database, an electronic patient record and an electronic health record using CDA/SCIPHOX. AMIA Annu. Symp. Proc. 2005, 963 (2005).
Wang, W. Y., Barratt, B. J., Clayton, D. G. & Todd, J. A. Genome-wide association studies: theoretical and practical concerns. Nature Rev. Genet. 6, 109–118 (2005).
Andersson, L. et al. Genetic mapping of quantitative trait loci for growth and fatness in pigs. Science 263, 1771–1774 (1994).
Li, C., Zhou, A. & Sang, T. Rice domestication by reducing shattering. Science 311, 1936–1939 (2006).
West, M. A. et al. Global eQTL mapping reveals the complex genetic architecture of transcript-level variation in Arabidopsis. Genetics 175, 1441–1450 (2007).
Xiang, H. et al. Single base-resolution methylome of the silkworm reveals a sparse epigenomic map. Nature Biotech. 28, 516–520 (2010).
Allen, E., Xie, Z., Gustafson, A. M. & Carrington, J. C. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121, 207–221 (2005).
Franco-Zorrilla, J. M. et al. Target mimicry provides a new mechanism for regulation of microRNA activity. Nature Genet. 39, 1033–1037 (2007).
Lu, C. et al. Elucidation of the small RNA component of the transcriptome. Science 309, 1567–1569 (2005).
Slotkin, R. K. et al. Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136, 461–472 (2009).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Liu, F., Marquardt, S., Lister, C., Swiezewski, S. & Dean, C. Targeted 3′ processing of antisense transcripts triggers Arabidopsis FLC chromatin silencing. Science 327, 94–97 (2010).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Wang, Z. Y., Seto, H., Fujioka, S., Yoshida, S. & Chory, J. BRI1 is a critical component of a plasma-membrane receptor for plant steroids. Nature 410, 380–383 (2001).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Yamanaka, S. A fresh look at iPS cells. Cell 137, 13–17 (2009).
Daley, G. Q. & Scadden, D. T. Prospects for stem cell-based therapy. Cell 132, 544–548 (2008).
Murry, C. E. & Keller, G. Differentiation of embryonic stem cells to clinically relevant populations: lessons from embryonic development. Cell 132, 661–680 (2008).
Saha, K. & Jaenisch, R. Technical challenges in using human induced pluripotent stem cells to model disease. Cell Stem Cell 5, 584–595 (2009).
Zhou, Q. et al. In vivo reprogramming of adult pancreatic exocrine cells to β-cells. Nature 455, 627–632 (2008).
Vierbuchen, T. et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463, 1035–1041 (2010).
Graf, T. & Enver, T. Forcing cells to change lineages. Nature 462, 587–594 (2009).
Jaenisch, R. & Young, R. A. Stem cells, the molecular circuitry of pluripotency and nuclear reprogramming. Cell 132, 567–582 (2008).
Janicki, S. M. & Spector, D. L. Nuclear choreography: interpretations from living cells. Curr. Opin. Cell Biol. 15, 149–157 (2003).
Probst, A. V., Dunleavy, E. & Almouzni, G. Epigenetic inheritance during the cell cycle. Nature Rev. Mol. Cell Biol. 10, 192–206 (2009).
Johannes, F., Colot, V. & Jansen, R. C. Epigenome dynamics: a quantitative genetics perspective. Nature Rev. Genet. 9, 883–890 (2008).
Teixeira, F. K. & Colot, V. Repeat elements and the Arabidopsis DNA methylation landscape. Heredity 105, 14–23 (2010).
Eichler, E. E. et al. Missing heritability and strategies for finding the underlying causes of complex disease. Nature Rev. Genet. 11, 446–450 (2010).
Heinig, M. et al. A trans-acting locus regulates an anti-viral expression network and type 1 diabetes risk. Nature (in the press).
McKinney, E. F. et al. A CD8+ T cell transcription signature predicts prognosis in autoimmune disease. Nature Med. 16, 586–591, 1p following 591 (2010).
Pham, M. X. et al. Gene-expression profiling for rejection surveillance after cardiac transplantation. N. Engl. J. Med. 362, 1890–1900 (2010).
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Manolio, T. A. Genomewide association studies and assessment of the risk of disease. N. Engl. J. Med. 363, 166–176 (2010).
Barabási, A. L. & Oltvai, Z. N. Network biology: understanding the cell's functional organization. Nature Rev. Genet. 5, 101–113 (2004).
Delbrück, M. in Unités Biologiques Douées De Continuité Génétique. 33–35 (Editions du Centre National de la Recherche Scientifique Paris, 1949).
Waddington, C. H. The Strategy of the Genes (Geo. Allen & Unwin, London, 1957).
Monod, J. & Jacob, F. Teleonomic mechanisms in cellular metabolism, growth, and differentiation. Cold Spring Harb. Symp. Quant. Biol. 26, 389–401 (1961).
Nurse, P. The great ideas of biology. Clin. Med. 3, 560–568 (2003).
Wagner, G. P. The developmental genetics of homology. Nature Rev. Genet. 8, 473–479 (2007).
Arendt, D. The evolution of cell types in animals: emerging principles from molecular studies. Nature Rev. Genet. 9, 868–882 (2008).
Lynch, V. J. et al. Adaptive changes in the transcription factor HoxA-11 are essential for the evolution of pregnancy in mammals. Proc. Natl Acad. Sci. USA 105, 14928–14933 (2008).
Patel, C. J., Bhattacharya, J. & Butte, A. J. An environment-wide association study (EWAS) on type 2 diabetes mellitus. PLoS ONE 5, e10746 (2010).
Buckler, E. S. et al. The genetic architecture of maize flowering time. Science 325, 714–718 (2009).
Schneeberger, K. et al. SHOREmap: simultaneous mapping and mutation identification by deep sequencing. Nature Methods 6, 550–551 (2009).
Ehrenreich, I. M. et al. Dissection of genetically complex traits with extremely large pools of yeast segregants. Nature 464, 1039–1042 (2010).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Dangl, J. L. & Jones, J. D. Plant pathogens and integrated defence responses to infection. Nature 411, 826–833 (2001).
Brady, S. M. et al. A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 318, 801–806 (2007).
Chickarmane, V. et al. Computational morphodynamics: a modeling framework to understand plant growth. Annu. Rev. Plant Biol. 61, 65–87 (2010).
Guenther, M. & Young, R. A. Repressive transcription. Science 329, 150–151 (2010).
Jacob, F. & Monod, J. Genetic regulatory mechanisms in the synthesis of proteins. J. Mol. Biol. 3, 318–356 (1961).
Gurdon, J. B. Adult frogs derived from the nuclei of single somatic cells. Dev. Biol. 4, 256–273 (1962).
Tang, F. et al. Tracing the derivation of embryonic stem cells from the inner cell mass by single-cell RNA-seq analysis. Cell Stem Cell 6, 468–478 (2010).
Acknowledgements
S.T. thanks members of her laboratory for helpful discussions.J.A.T. thanks the Wellcome Trust, the Juvenile Diabetes Research Foundation International and the National Institute for Health Research Cambridge Biomedical Research Centre for funding, and J. Nadeau for sharing unpublished information. The Cambridge Institute for Medical Research is a recipient of a Wellcome Trust Strategic Award (079895). M.V. would like to thank M. Walhout, J. Dekker and J. Vandenhaute for helpful conversations on the subject discussed here.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Rights and permissions
About this article
Cite this article
Heard, E., Tishkoff, S., Todd, J. et al. Ten years of genetics and genomics: what have we achieved and where are we heading?. Nat Rev Genet 11, 723–733 (2010). https://doi.org/10.1038/nrg2878
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg2878
This article is cited by
-
Multilevel comparative bioinformatics to investigate evolutionary relationships and specificities in gene annotations: an example for tomato and grapevine
BMC Bioinformatics (2018)
-
Happiness in Behaviour Genetics: An Update on Heritability and Changeability
Journal of Happiness Studies (2017)
-
ComPlEx: conservation and divergence of co-expression networks in A. thaliana, Populus and O. sativa
BMC Genomics (2014)
-
Isoform level expression profiles provide better cancer signatures than gene level expression profiles
Genome Medicine (2013)