Key Points
-
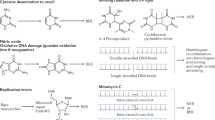
Living cells respond to DNA damage by a variety of mechanisms, including a series of biochemical pathways called DNA repair. These include three discrete pathways for the excision of damaged bases, called base excision repair, mismatch repair and nucleotide excision repair (NER).
-
NER in human cells is a complex biochemical process during which a large multiprotein complex is assembled at several types of base damage. This multiprotein complex (NER machine) catalyses the excision of damaged bases as oligonucleotide fragments.
-
The RNA polymerase II basal transcription factor, TFIIH, is an integral component of the NER multiprotein complex.
-
NER operates somewhat differently on DNA that is transcriptionally active (transcription-coupled repair) and that which is transcriptionally silent (global genome repair).
-
Defective NER in humans caused by genetically inherited mutations in NER genes results in the skin-cancer-prone disease xeroderma pigmentosum.
-
Hereditary defects in transcription-coupled NER can result in a disease called Cockayne syndrome, which is characterized by severe developmental and neurological disorders.
-
Mutational inactivation of certain NER genes can result in a combined syndrome of xeroderma pigmentosum and Cockayne syndrome, or in yet another disease called trichothiodystrophy, which is characterized by brittle hair and nails.
-
Cockayne syndrome, combined xeroderma pigmentosum/Cockayne syndrome complex and trichothiodystrophy are not usually associated with increased cancer risk.
-
Mouse mutant strains generated by targeted gene replacement have been constructed to model these human NER-defective syndromes.
Abstract
Eukaryotic cells can repair many types of DNA damage. Among the known DNA repair processes in humans, one type — nucleotide excision repair (NER) — specifically protects against mutations caused indirectly by environmental carcinogens. Humans with a hereditary defect in NER suffer from xeroderma pigmentosum and have a marked predisposition to skin cancer caused by sunlight exposure. How does NER protect against skin cancer and possibly other types of environmentally induced cancer in humans?
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Friedberg, E. C., Walker, G. C. & Siede, W. DNA Repair and Mutagenesis. (ASM Press, Washington, 1995).
Lindahl, T. Instability and decay of the primary structure of DNA. Nature 362, 709–715 (1993).
Lindahl, T. & Wood, R. D. Quality control by DNA repair. Science 286, 1897–1905 (1999).
Lindahl, T. The Croonian Lecture, 1996: endogenous damage to DNA. Phil. Trans. R. Soc. Lond. B 351, 1529–1538 (1996).
Walker, G. C. Mutagenesis and inducible responses to deoxyribonucleic acid damage in Escherichia coli. Microbiol. Rev. 48, 60–93 (1984).
Kunkel, T. A. & Bebenek, K. DNA replication fidelity. Annu. Rev. Biochem. 69, 497–529 (2000).
Hoeijmakers, J. H. J. Genome maintenance mechanisms for preventing cancer. Nature 411, 366–374 (2001).
Friedberg, E. C. Summary: Biological responses to DNA damage: a perspective in the new millennium. Cold Spring Harbor Symp. Quant. Biol. 65, 593–602 (2000).
Lindahl, T., Karran, P. & Wood, R. D. DNA excision repair pathways. Curr. Opin. Genet. Dev. 7, 158–169 (1997).
Friedberg, E. C. & Wood, R. D. in DNA Replication in Eukaryotic Cells (ed. dePamphilis, M.) 249–269 (Cold Spring Harbor Laboratory Press, Long Island, 1996).
Sancar, A. DNA excision repair. Annu. Rev. Biochem. 65, 43–81 (1996).
Lindahl, T. Suppression of spontaneous mutagenesis in human cells by DNA base excision repair. Mutat. Res. 462, 129–135 (2000).
Memisoglu, A. & Samson, L. Base excision repair in yeast and mammals. Mutat. Res. 451, 39–51 (2000).
Mol, C. D., Parikh, S. S., Putnam, C. D., Lo, T. P. & Tainer, J. A. DNA repair mechanisms for the recognition and removal of damaged DNA bases. Annu. Rev. Biophys. Biomol. Struct. 28, 101–128 (1999).
Krokan, H. E., Nilsen, H., Skorpen, F., Otterlei, M. & Slupphaug, G. Base excision repair of DNA in mammalian cells. FEBS Lett. 476, 73–77 (2000).
McCullough, A. K., Dodson, M. L. & Lloyd, R. S. Initiation of base excision repair: glycosylase mechanisms and structures. Annu. Rev. Biochem. 68, 255–285 (1999).
Parikh, S. S., Mol, C. D., Hosfield, D. J. & Tainer, J. A. Envisioning the molecular choreography of DNA base excision repair. Curr. Opin. Struct. Biol. 9, 37–47 (1999).
de Boer, J. & Hoeijmakers, J. H. J. Nucleotide excision repair and human syndromes. Carcinogenesis 21, 453–460 (2000).
Prakash, S. & Prakash, L. Nucleotide excision repair in yeast. Mutat. Res. 451, 13–24 (2000).
de Laat, W. L., Jaspers, N. G. J. & Hoeijmakers, J. H. J. Molecular mechanism of nucleotide excision repair. Genes Dev. 13, 768–785 (1999).
Petit, C. & Sancar, A. Nucleotide excision repair: from E. coli to man. Biochimie 81, 15–25 (1999).
Wood, R. D. Nucleotide excision repair in mammalian cells. J. Biol. Chem. 272, 23465–23468 (1997).
Buermeyer, A. B., Deschenes, S. M., Baker, S. M. & Liskay, R. M. Mammalian DNA mismatch repair. Annu. Rev. Genet. 33, 533–564 (1999).
Modrich, P. Strand-specific mismatch repair in mammalian cells. J. Biol. Chem. 272, 24727–24730 (1997).
Svejstrup, J. Q. et al. Different forms of TFIIH for transcription and DNA repair: holo-TFIIH and a nucleotide excision repairosome. Cell 80, 21–28 (1995).An early contribution to our understanding of the role of the transcription factor TFIIH in NER in yeast.
Rodriguez, K. et al. Affinity purification and partial characterization of a yeast multiprotein complex for nucleotide excision repair using histidine-tagged Rad14 protein. J. Biol. Chem. 273, 34180–34189 (1998).
Guzder, S. N., Sung, P., Prakash, L. & Prakash, S. Nucleotide excision repair in yeast is mediated by sequential assembly of repair factors and not by a pre-assembled repairosome. J. Biol. Chem. 271, 8903–8910 (1996).
Aboussekhra, A. et al. Mammalian DNA nucleotide excision repair reconstituted with purified protein components. Cell 80, 859–868 (1995).
Wakasugi, M. & Sancar, A. Assembly, subunit composition, and footprint of human DNA repair excision nuclease. Proc. Natl Acad. Sci. USA 95, 6669–6674 (1998).
Guzder, S. N., Habraken, Y., Sung, P., Prakash, L. & Prakash, S. Reconstitution of yeast nucleotide excision repair with purified Rad proteins, replication protein A, and transcription factor TFIIH. J. Biol. Chem. 270, 12973–12976 (1995).References 28–30 are landmark studies on the reconstitution of NER in vitro with purified proteins.
Levin, D. S., McKenna, A. E., Motycka, T. A., Matsumoto, Y. & Tomkinson, A. E. Interaction between PCNA and DNA ligase I is critical for joining of Okazaki fragments and long-patch base excision repair. Curr. Biol. 10, 919–922 (2000).
Kornberg, R. D. & Lorch, Y. Chromatin structure and transcription. Annu. Rev. Cell Biol. 8, 563–587 (1992).
Robertson, K. D. & Jones, P. A. Dynamic interrelationships between DNA replication, methylation and repair. Am. J. Hum. Genet. 61, 1220–1224 (1997).
Thoma, F. Light and dark in chromatin repair: repair of UV-induced DNA lesions by photolyase and nucleotide excision repair. EMBO J. 18, 6585–6598 (1999).
Sugasawa, K. et al. A multistep damage recognition mechanism for global genomic nucleotide excision repair. Genes Dev. 15, 507–521 (2001).
Wood, R. D. DNA damage recognition during nucleotide excision repair in mammalian cells. Biochimie 81, 39–44 (1999).
Naegeli, H. in DNA Recombination and Repair (eds Smith, P. & Jones, C.) 99–137 (Oxford Univ. Press, Oxford, 1999).
Sugasawa, K. et al. Xeroderma pigmentosum group C protein complex is the initiator of global genomic nucleotide excision repair. Mol. Cell 2, 223–232 (1998).An important study, explaining the role of XPC protein in the recognition of base damage during NER.
Masutani, C. et al. Purification and cloning of a nucleotide excision repair complex involving the xeroderma pigmentosum group C protein and a human homolog of yeast RAD23. EMBO J. 13, 1831–1843 (1994).
Sugasawa, K. et al. HHR23B, a human Rad23 homolog, stimulates XPC protein in nucleotide excision repair in vitro. Mol. Cell. Biol. 16, 4852–4861 (1996).
Araki, M. et al. Centrosome protein centrin2/caltractin1 is part of the xeroderma pigmentosum group C complex that initiates global genome nucleotide excision repair. J. Biol. Chem. 276, 18665–18672 (2001).
Hess, M. T., Schwitter, U., Petretta, M., Giese, B. & Naegeli, H. Bipartite substrate discrimination by human nucleotide excision repair. Proc. Natl Acad. Sci. USA 94, 6664–6669 (1997).An excellent contribution to recent models for the molecular basis of damage recognition during NER.
Friedberg, E. C. Relationships between DNA repair and transcription. Annu. Rev. Biochem. 65, 15–42 (1996).
Hanawalt, P. C. Transcription-coupled repair and human disease. Science 266, 1957–1958 (1994).
Hanawalt, P. C. & Spivak, G. in Advances in DNA Repair (eds Dizdaroglu, M. & Karakaya, A.) 169–179 (Academic/Plenum Publishing, New York, 1999).
Tsutakawa, S. E. & Cooper, P. K. Transcription-coupled repair of oxidative DNA damage in human cells: mechanisms and consequences. Cold Spring Harbor Symp. Quant. Biol. 65, 201–215 (2000).
Friedberg, E. C. Cockayne Syndrome: a primary defect in DNA repair, transcription, both or neither? BioEssays 18, 731–738 (1996).
Hebra, F. & Kaposi, M. On Diseases of the Skin, including the Exanthemata. Vol. 16 (translated by W. Tay, London) 252–258 (New Sydenham Society, London, 1874).
Cleaver, J. E. Defective repair replication of DNA in xeroderma pigmentosum. Nature 218, 652–656 (1968).
Setlow, R. B., Regan, J. D., German, J. & Carrier. W. L. Evidence that xeroderma pigmentosum cells do not perform the first step in the repair of ultraviolet damage to their DNA. Proc. Natl Acad. Sci. USA 64, 1035–1041 (1969).
Cleaver, J. E. & Kraemer, K. H. in The Metabolic and Molecular Basis of Inherited Disease (eds Scriver, C. R., Beaudet, A. L., Sly, W. S. & Valle, D.) 4393–4419 (McGraw–Hill, New York, 1995).
Bootsma, D., Kraemer, K. H., Cleaver, J. E. & Hoeijmakers, J. H. H. in The Genetic Basis of Human Cancer (eds Vogelstein, B. & Kinzler, K, W.) 245–274 (McGraw–Hill, New York, 1998).
Hoeijmakers, J. H. J. Human nucleotide excision repair syndromes: molecular clues to unexpected intricacies. Eur. J. Cancer 30A, 1912–1921 (1994).
Masutani, C. et al. Xeroderma pigmentosum variant: from a human genetic disorder to a novel DNA polymerase. Cold Spring Harbor Symp. Quant. Biol. 65, 71–80 (2000).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Lengauer, C., Kinzler, K. W. & Vogelstein, B. Genetic instabilities in human cancers. Nature 396, 643–649 (1998).
Friedberg, E. C. Cancer predisposition associated with defective DNA repair-studies with mutant mouse strains. Cancer J. Sci. Am. 5, 257–263 (1999).
Friedberg, E. C. & Meira, L. B. Database of mouse strains carrying targeted mutations in genes affecting cellular responses to DNA damage. Version 4. Mutat. Res. 459, 243–274 (2000).A comprehensive database of mouse mutants defective in various DNA repair modes.
de Vries, A. & van Steeg, H. Xpa knockout mice. Sem. Can. Biol. 7, 229–240 (1996).
Cheo, D. L., Burns D. K., Meira, L. B., Houle, J. F. & Friedberg, E. C. Mutational inactivation of the xeroderma pigmentosum group C gene confers predisposition to 2-acetylaminofluorene-induced liver and lung cancer and to spontaneous testicular cancer in Trp53−/− mice. Cancer Res. 59, 771–775 (1999).
Hanawalt, P. C. Revisiting the rodent repairadox. Env. Mol. Mutagen. (in the press).
Swift, M. & Chase, C. Cancer in families with xeroderma pigmentosum. J. Natl Cancer Inst. 62, 1415–1421 (1979).
Cheo, D. L., Meira, L. B., Burns, D. K, Reis, A. M. & Friedberg, E. C. UVB radiation-induced skin cancer in mice defective in the Xpc, Trp53 and Apex (HAP1) genes: genotype-specific effects on cancer predisposition and pathology of tumors. Cancer Res. 60, 1580–1584 (2000).
Wijnhoven, S. W. P. et al. Age-dependent spontaneous mutagenesis in Xpc mice defective in nucleotide excision repair. Oncogene 19, 5034–5037 (2000).
McWhir, J., Selfridge, J., Harrison, D. J., Squires, S. & Melton, D. W. Mice with DNA repair gene (ERCC-1) deficiency have elevated levels of p53, liver nuclear abnormalities and die before weaning. Nature Genet. 5, 217–224 (1993).
Weeda, G. et al. Disruption of the mouse ERCC1 gene results in a novel repair syndrome with growth failure, nuclear abnormalities and senescence. Curr. Biol. 7, 427–439 (1997).
Taylor, E. M. et al. Xeroderma pigmentosum and trichothiodystrophy are associated with different mutations in the XPD(ERCC2) repair/transcription gene. Proc. Natl Acad. Sci. USA 94, 8658–8663 (1997).
Lehmann, A. R. The xeroderma pigmentosum group D (XPD) gene: one gene, two functions, three diseases. Genes Dev. 15, 15–23 (2000).
Vermeulen, W. et al. Three unusual repair deficiencies associated with transcription factor BTF2 (TFIIH). Evidence for the existence of a transcription syndrome. Cold Spring Harb, Symp. Quant. Biol. 54, 317–329 (1994).
de Boer, J. et al. A mouse model for the basal transcription/DNA repair syndrome trichothiodystrophy. Mol. Cell 1, 981–990 (1998).
Vermeulen, W. et al. Sublimiting concentration of TFIIH transcription/DNA repair factor causes TTD-A trichothiodystrophy disorder. Nature Genet. 26, 307–313 (2000).
Harada, Y.-N. et al. Post-natal growth failure, a short life span, and early onset of cellular senescence and subsequent immortalization in mice lacking the xeroderma pigmentosum group G gene. Mol. Cell. Biol. 19, 2366–2372 (1999).
Nouspiekel, T., Lalle, P., Leadon, S. A., Cooper, P. K. & Clarkson, S. G. A common mutational pattern in Cockayne syndrome patients from xeroderma pigmentosum group G: implications for a second XPG function. Proc. Natl Acad. Sci. USA 94, 3116–3121 (1997).
Le Page, F. et al. Transcription-coupled repair of 8-oxoguanine: requirement for XPG, TFIIH, and CSB and implications for Cockayne syndrome. Cell 101, 159–171 (2000).An excellent recent contribution to the complexities of transcription-coupled DNA repair.
van der Horst, G. T. J. et al. Defective transcription-coupled repair in Cockayne syndrome B mice is associated with skin cancer predisposition. Cell 89, 425–435 (1997).
Lu, Y. et al. Disruption of the Cockayne syndrome B gene impairs spontaneous tumorigenesis in cancer-predisposed Ink4a/ARF knockout mice. Mol. Cell. Biol. 21, 1810–1818 (2001).
Stillman, B. Foreword. Cold Spring Harbor Symp. Quant. Biol. 65, 21 (2000).
Gillette, T. et al. The proteasome 19S complex regulates nucleotide excision repair in yeast. Genes Dev. 15, 1528–1539 (2001).
Yasuhira, S. & Yasui, A. Alternative excision repair pathway of UV-damaged DNA in Schizosaccharomyces pombe operates both in nucleus and in mitochondria. J. Biol. Chem. 275, 11824–11828 (2000).
Kaur, B. & Doetsch, P. Ultraviolet damage endonuclease (Uve1p): a structure and strand-specific DNA endonuclease. Biochemistry 39, 5788–5796 (2000).
Lunn, R. M. et al. XPD polymorphisms: effects on DNA repair proficiency. Carcinogenesis 21, 551–555 (2000).
Tomescu, D., Kavanagh, G., Ha, T., Campbell, H. & Melton, D. W. Nucleotide excision repair gene XPD polymorphisms and genetic predisposition to melanoma. Carcinogenesis 22, 403–408 (2001).
Shen, H. et al. An intronic poly (AT) polymorphism of the DNA repair gene XPC and risk of squamous cell carcinoma of the head and neck: a case-control study. Cancer Res. 61, 3321–3325 (2001).
Zhou, B.-B. S. & Elledge, S. J. The DNA damage response: putting checkpoints in perspective. Nature 408, 433–439 (2000).
Walker, G. C. Understanding the complexity of an organism's responses to DNA damage. Cold Spring Harb. Symp. Quant. Biol. 65, 1–10 (2000).
Jiricny, J. & Nystrom-Lahti, M. Mismatch repair defects in cancer. Curr. Opin. Genet. Dev. 10, 157–161 (2000).
Robbins, J. H., Kraemer, K. H., Lutzner, M. A., Festoff, B. W. & Coon, G. Xeroderma pigmentosum. An inherited disease with sun sensitivity, multiple cutaneous neoplasms and abnormal DNA repair. Ann. Intern. Med. 80, 221–248 (1974).
Takebe, H. et al. DNA repair characteristics and skin cancers of xeroderma pigmentosum patients in Japan. Cancer Res. 367, 490–495 (1977).
Kraemer, K. H., Myung, M. L. & Scotto, J. Xeroderma pigmentosum. Cutaneous, ocular, and neurologic abnormalities in 830 published cases. Arch. Derm. 123, 241–250 (1987).
Prakash, S. et al. Role of yeast and human DNA polymerase η in error-free replication of damaged DNA. Cold Spring Harb. Symp. Quant. Biol. 65, 51–59 (2000).
Gerlach, V. L. et al. Human DNA polymerase κ: a novel DNA polymerase in search of a biological function. Cold Spring Harbor Symp. Quant. Biol. 65, 41–49 (2000).
Cockayne, E. A. Dwarfism with retinal atrophy and deafness. Arch. Dis. Child. 11, 1–8 (1936).
Nance, M. A. & Berry, S. A. Cockayne syndrome: review of 140 cases. Am. J. Med. Genet. 42, 68–84 (1992).
Schmickel, R. D., Chu, E. H. Y., Trosko, J. E. & Chang, C. C. Cockayne syndrome: a cellular sensitivity to ultraviolet light. Pediatrics 60, 135–139 (1977).
Rapin, I., Lindenbaum, Y., Dickson, D. W., Kraemer, K. H. & Robbins, J. H. Cockayne syndrome and xeroderma pigmentosum. DNA repair disorders with overlaps and paradoxes. Neurology 55, 1442–1449 (2000).
Berneburg, M. & Lehmann, A. R. Xeroderma pigmentosum and related disorders: defects in DNA repair and transcription. Adv. Genet. 43, 71–102 (2001).
Lehmann, A. R., Kirk-Bell, S. and Mayne, L. Abnormal kinetics of DNA synthesis in ultraviolet light-irradiated cells from patients with Cockayne's syndrome. Cancer Res. 39, 4237–4241 (1979).
Venema, J., Mullenders, L. H. F., Natarajan, A. T., van Zeeland, A. A. & Mayne, L. V. The genetic defect in Cockayne's syndrome is associated with a defect in repair of UV-induced DNA damage in transcriptionally active DNA. Proc. Natl Acad. Sci. USA 87, 4704–4711 (1990).
van Gool, A. J., van der Horst, T. J., Citterio, E. & Hoeijmakers, J. H. J. Cockayne syndrome: defective repair or transcription? EMBO J. 14, 4155–4162 (1997).
Hanawalt, P. C. DNA repair. The bases for Cockayne syndrome. Nature 405, 415–416 (2000).
Itin, P. H., Sarasin, A. & Pittelkow, M. R. Trichothiodystrophy: update on the sulfur-deficient brittle hair syndrome. J. Am. Acad. Dermatol. 44, 891–920 (2001).
Weeda, G. et al. A mutation in the XPB/ERCC3 DNA repair transcription gene, associated with trichothiodystrophy. Am. J. Hum. Genet. 60, 320–329 (1997).
Stefanini, M. et al. A new nucleotide-excision-repair gene associated with the disorder trichothiodystrophy. Am. J. Hum. Genet. 53, 817–821 (1993).
Acknowledgements
I apologize to numerous colleagues for the omission of many pertinent references owing to space constraints. I also gratefully acknowledge B. Stewart for outstanding assistance with the illustrations, and L. Queimado, R. Rolig, M. Liskay and P. Modrich for critical reading of some or all of the manuscript.
Author information
Authors and Affiliations
Supplementary information
Related links
Glossary
- BASE DAMAGE
-
Any change in the chemistry of the nitrogenous bases in DNA (adenine, thymine, guanine or cytosine), as well as the presence of inappropriate bases such as uracil, which is normally only in RNA.
- REPLICATION FORK
-
The region of replicating DNA in which active DNA synthesis is occurring to generate two copies of the parental DNA.
- REPAIR SYNTHESIS
-
A mode of DNA synthesis that is associated with DNA repair rather than with replication. Sometimes called conservative DNA synthesis to distinguish it from the semi-conservative DNA synthesis that characterizes replication.
- SEMI-CONSERVATIVE
-
This term refers to the way in which double-stranded DNA is replicated in all cells. During replication, each of the parental DNA strands is partitioned to a new double-stranded DNA molecule. So one half of the parental DNA is conserved, hence semi-conservative.
- REACTIVE OXYGEN SPECIES
-
(ROS). Highly reactive chemical radicals that are generated as products of oxygen degradation.
- COMPLEMENTATION GROUP
-
This refers to a genetic grouping of individuals, based on the fact that cells from individuals in any complementation group cannot correct a phenotypic defect among themselves, but can correct it in cells from other groups. Individuals in a given complementation group usually carry mutations in the same gene.
- BASAL TRANSCRIPTION FACTORS
-
A set of protein complexes that associate with RNA polymerase II during the initiation of all mRNA synthesis. Sometimes called general transcription factors.
- ENDONUCLEASES
-
Enzymes that cut the sugar–phosphate backbone of DNA (or RNA) at sites other than the free ends of linear DNA (which are degraded by exonucleases).
- SOMATIC MUTATION THEORY
-
A theory on the pathogenesis of cancer that essentially postulates that cancer in somatic cells arises from mutations. More recently, the theory has been adapted to specifically implicate mutations in oncogenes or tumour suppressor genes.
- HAPLOINSUFFICIENCY
-
A phenotypic state that results from loss of one functional allele of any given gene in diploid cells. Sometimes also called allelic insufficiency.
- HPRT GENE
-
A gene that encodes hypoxanthine phosphoribosyltransferase. It is a convenient marker for measuring mutation frequency in cells, because cells with a mutant HPRT gene require supplementation with 6-thioguanine to grow.
Rights and permissions
About this article
Cite this article
Friedberg, E. How nucleotide excision repair protects against cancer. Nat Rev Cancer 1, 22–33 (2001). https://doi.org/10.1038/35094000
Issue Date:
DOI: https://doi.org/10.1038/35094000
This article is cited by
-
UV-induced G4 DNA structures recruit ZRF1 which prevents UV-induced senescence
Nature Communications (2023)
-
Genetic variability in cisplatin metabolic pathways and outcome of locally advanced head and neck squamous cell carcinoma patients
Scientific Reports (2023)
-
Genetic variability of oxidative stress and DNA repair genes associated with pre-treatment cancer-related fatigue in women with breast cancer
Supportive Care in Cancer (2023)
-
Inflammatory landscape in Xeroderma pigmentosum patients with cutaneous melanoma
Scientific Reports (2022)
-
Side effect prediction based on drug-induced gene expression profiles and random forest with iterative feature selection
The Pharmacogenomics Journal (2021)