Abstract
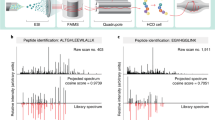
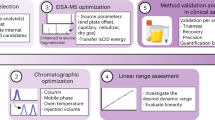
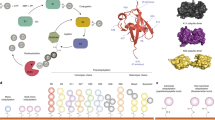
Measuring enzymatic activities in biological fluids is a form of activity-based proteomics and may be utilized as a means of developing disease biomarkers. Activity-based assays allow amplification of output signals, thus potentially visualizing low-abundant enzymes on a virtually transparent whole-proteome background. The protocol presented here describes a semiquantitative in vitro assay of proteolytic activities in complex proteomes by monitoring breakdown of designer peptide substrates using robotic extraction and a matrix-assisted laser desorption/ionization–time of flight (MALDI-TOF) mass spectrometric readout. Relative quantitation of the peptide metabolites is carried out by comparison with spiked internal standards, followed by statistical analysis of the resulting mini-peptidome. Partial automation provides reproducibility and throughput essential for comparing large sample sets. The approach may be used for diagnostic or predictive purposes and it enables profiling of 96 samples in 30 h. It could be tailored to many diagnostic and pharmaco-dynamic purposes as a readout of catalytic and metabolic activities in body fluids or tissues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lopez-Otin, C. & Bond, J.S. Proteases: multifunctional enzymes in life and disease. J. Biol. Chem. 283, 30433–30437 (2008).
Overall, C.M. & Blobel, C.P. In search of partners: linking extracellular proteases to substrates. Nat. Rev. 8, 245–257 (2007).
Palermo, C. & Joyce, J.A. Cysteine cathepsin proteases as pharmacological targets in cancer. Trends Pharmacol. Sci. 29, 22–28 (2008).
Lopez-Otin, C. & Matrisian, L.M. Emerging roles of proteases in tumour suppression. Nat. Rev. Cancer 7, 800–808 (2007).
Egeblad, M. & Werb, Z. New functions for the matrix metalloproteinases in cancer progression. Nat. Rev. Cancer 2, 161–174 (2002).
Leiting, B. et al. Catalytic properties and inhibition of proline-specific dipeptidyl peptidases II, IV and VII. Biochem. J. 371, 525–532 (2003).
Martinez, J.M. et al. Aminopeptidase activities in breast cancer tissue. Clin. Chem. 45, 1797–1802 (1999).
Villanueva, J. et al. Serum peptide profiling by magnetic particle-assisted, automated sample processing and MALDI-TOF mass spectrometry. Anal. Chem. 76, 1560–1570 (2004).
Villanueva, J. et al. Differential exoprotease activities confer tumor-specific serum peptidome patterns. J. Clin. Invest. 116, 271–284 (2006).
Villanueva, J. et al. Serum peptidome patterns that distinguish metastatic thyroid carcinoma from cancer-free controls are unbiased by gender and age. Mol. Cell. Proteomics 5, 1840–1852 (2006).
Villanueva, J. et al. A sequence-specific exopeptidase activity test (SSEAT) for 'functional' biomarker discovery. Mol. Cell. Proteomics 7, 509–518 (2008).
Villanueva, J., Lawlor, K., Toledo-Crow, R. & Tempst, P. Automated serum peptide profiling. Nat. Protoc. 1, 880–891 (2006).
Villanueva, J., Philip, J., DeNoyer, L. & Tempst, P. Data analysis of assorted serum peptidome profiles. Nat. Protoc. 2, 588–602 (2007).
Acknowledgements
This work was supported by US National Institutes of Health Grants R33 CA111942 and U24 CA126485. It is part of National Cancer Institute's Clinical Proteomic Technologies Initiative and Clinical Proteomic Technology Assessment for Cancer (CPTAC) consortium (Broad Institute of MIT and Harvard, Memorial Sloan-Kettering Cancer Center, Purdue University, University of California, San Francisco and Vanderbilt University School of Medicine).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Fig. 1
Design diagram of the custom aliquoting rack. Custom aliquoting rack designed to hold sixteen rows of nine 0.5-mL eppendorf tubes for use with a liquid handler robot of variable pipette separation (Tecan Genesis Freedom). At the end of each row is a sample source vial kept in a separate rack. The rack is machined in aluminum and placed on a base with inserts that match the work surface of the liquid handler. The component parts of the rack: (a) 9 × 16 aliquot vial tray (yellow), (b) 16 source vial block (green), mounted on (c) the matching base for the Tecan handling robot (gray, red). (PDF 113 kb)
Supplementary Fig. 2
Design diagrams of the custom side-magnet rack. Panel A shows the exploded assembly view of the top part, magnets and bottom part of the rack (and a 96-well vial plate). Panel B shows the projected design views of the top and bottom parts of the rack. The machined material used is clear Lexan and all dimensions are in millimeters. (PDF 238 kb)
Supplementary Fig. 3
Design diagrams of the custom bottom-magnet rack. Panel A shows the exploded assembly view of the top part, magnets, and bottom part of the rack (and a 96-well vial plate). Panel B shows the orthogonal projected design views of the top and bottom parts of the rack. The machined material used is clear Lexan and all dimensions are in millimeters. (PDF 270 kb)
Supplementary Manual 1
Instructions for the use of the Mass Spectra Viewer (MSV). (PDF 894 kb)
Rights and permissions
About this article
Cite this article
Villanueva, J., Nazarian, A., Lawlor, K. et al. Monitoring peptidase activities in complex proteomes by MALDI-TOF mass spectrometry. Nat Protoc 4, 1167–1183 (2009). https://doi.org/10.1038/nprot.2009.88
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2009.88
This article is cited by
-
Functional protease profiling with reporter peptides in serum specimens of colorectal cancer patients: demonstration of its routine diagnostic applicability
Journal of Experimental & Clinical Cancer Research (2012)
-
Early diagnostic protein biomarkers for breast cancer: how far have we come?
Breast Cancer Research and Treatment (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.