Abstract
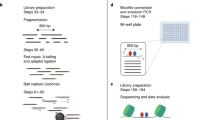
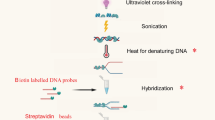
Southwestern blotting is used to investigate DNA–protein interactions. The advantage of this technique over other related methods such as electrophoretic mobility shift assay (EMSA) and DNA footprinting is that it provides information regarding the molecular weight of unknown protein factor. This method combines the features of Southern and Western blotting techniques; a denaturing SDS-PAGE is first employed to separate proteins electrophoretically based on size, and after transferring the proteins to a membrane support, the membrane-bound proteins are renatured and incubated with a 32P-labeled double-stranded oligonucleotide probe of specific DNA sequence. The interaction of the probe with the protein(s) is later visualized by autoradiography. This technique could be combined with database searching (TransFac, http://www.gene-regulation.com/pub/databases.html#transfac), prediction of potential protein factors binding onto a target motif (e.g., Patch search), in vitro supershift EMSA and in vivo chromatin immunoprecipitation (ChIP) assays for effective identification of protein factors. The whole Southwestern blotting procedure takes ∼4 d to complete. In this article, a commonly used protocol and expected results are described and discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bowen, B., Steinberg, J., Laemmli, U.K. & Weintraub, H. The detection of DNA-binding proteins by protein blotting. Nucleic Acids Res. 8, 1–20 (1980).
Jack, R.S., Gehring, W.J. & Brack, C. Protein component from Drosophila larval nuclei showing sequence specificity for a short region near a major heat-shock protein gene. Cell 24, 321–331 (1981).
Anachkova, B. & Russev, G. Differential binding of nonhistone chromosomal proteins to the putative mouse origin of replication. Biochim. Biophys. Acta 740, 369–372 (1983).
Triadou, P., Crepin, M., Gros, F. & Lelong, J.C. Tissue-specific binding of total and beta-globin genomic deoxyribonucleic acid to non-histone chromosomal proteins from mouse erythropoietic cells. Biochemistry 21, 6060–6065 (1982).
Watt, R.A., Shatzman, A.R. & Rosenberg, M. Expression and characterization of the human c-myc DNA-binding protein. Mol. Cell Biol. 5, 448–456 (1985).
Miskimins, W.K., Roberts, M.P., McClelland, A. & Ruddle, F.H. Use of a protein-blotting procedure and a specific DNA probe to identify nuclear proteins that recognize the promoter region of the transferrin receptor gene. Proc. Natl. Acad. Sci. USA 82, 6741–6744 (1985).
Cheng, C.K., Yeung, C.M., Hoo, R.L., Chow, B.K. & Leung, P.C. Oct-1 is involved in the transcriptional repression of the gonadotropin-releasing hormone receptor gene. Endocrinology 143, 4693–4701 (2002).
Lee, L.T., Tan-Un, K.C., Lin, M.C. & Chow, B.K. Retinoic acid activates human secretin gene expression by Sp proteins and nuclear factor I in neuronal SH-SY5Y cells. J. Neurochem. 93, 339–350 (2005).
Hellman, L.M. & Fried, M.G. Electrophoretic mobility shift assay (EMSA) for detecting protein-nucleic acid interactions. Nat. Protoc. 2, 1849–1861 (2007).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Chekmenev, D.S., Haid, C. & Kel, A.E. P-Match: transcription factor binding site search by combining patterns and weight matrices. Nucleic Acids Res. 33, W432–W437 (2005).
Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Lowry, O.H., Rosebrough, N.J., Farr, A.L. & Randall, R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193, 265–275 (1951).
Lee, K.A., Bindereif, A. & Green, M.R. A small-scale procedure for preparation of nuclear extracts that support efficient transcription and pre-mRNA splicing. Gene Anal. Tech. 5, 22–31 (1988).
Acknowledgements
The work was supported by Hong Kong Government RGC Grants HKU7501/05M, HKU7639/07M to B.K.C.C.
Author information
Authors and Affiliations
Contributions
F.K.Y.S. and L.T.O.L. contributed equally to this work, and should be considered as cofirst authors.
Corresponding author
Rights and permissions
About this article
Cite this article
Siu, F., Lee, L. & Chow, B. Southwestern blotting in investigating transcriptional regulation. Nat Protoc 3, 51–58 (2008). https://doi.org/10.1038/nprot.2007.492
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.492
This article is cited by
-
A Region Containing an as-1 Element of Dahlia Mosaic Virus (DaMV) Subgenomic Transcript Promoter Plays a Key Role in Green Tissue- and Root-Specific Expression in Plants
Plant Molecular Biology Reporter (2015)
-
Promoter trapping method: transcription factor purification using human telomerase reverse transcriptase promoter
Proteome Science (2014)
-
L-carnitine and PPARα-agonist fenofibrate are involved in the regulation of Carnitine Acetyltransferase (CrAT) mRNA levels in murine liver cells
BMC Genomics (2014)
-
Comparative analysis of synthetic DNA promoters for high-level gene expression in plants
Planta (2014)
-
DNA–protein interactions: methods for detection and analysis
Molecular and Cellular Biochemistry (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.