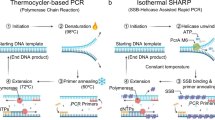
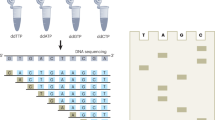
Abstract
This protocol describes the design and execution of monoplex and multiplex linear-after-the-exponential (LATE)-PCR assays using a novel reagent, PrimeSafe, that suppresses all forms of mispriming. LATE-PCR is an advanced form of asymmetric amplification that uses a limiting primer and an excess primer for efficient exponential amplification of double-stranded DNA, followed by linear amplification of one strand. Each single-stranded amplicon can be quantitatively detected in real time or at end point. By separating primer annealing from product detection, LATE-PCR enables product analysis at low temperatures. Alternatively, each single strand can be sequenced by a convenient Dilute-'N'-Go procedure. Amplified samples are diluted with individual sequencing primers without the use of columns or spins. We have amplified and then sequenced 15 different single-stranded products generated in a single multiplexed LATE-PCR comprised of 15 pairs of unrelated primers. Dilute-'N'-Go dideoxy sequencing is more convenient, faster and less expensive than sequencing double-stranded amplicons generated via conventional symmetric PCR. The preparation of LATE-PCR products for Dilute-'N'-Go sequencing takes only 30 seconds.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Surdhar, G.K. Cycle sequencing of PCR products. Methods Mol. Biol. 187, 65–72 (2002).
Gyllensten, U.B. & Erlich, H.A. Generation of single-stranded DNA by the polymerase chain reaction and its application to direct sequencing of the HLA-DQA locus. Proc. Natl. Acad. Sci. USA 85, 7652–7656 (1988).
Shyamala, V. & Ames, G.F. Amplification of bacterial genomic DNA by the polymerase chain reaction and direct sequencing after asymmetric amplification: application to the study of periplasmic permeases. J. Bacteriol. 171, 1602–1608 (1989).
Innis, M.A. et al. DNA sequencing with Thermus aquaticus DNA polymerase and direct sequencing of polymerase chain reaction-amplified DNA. Proc. Natl. Acad. Sci. USA 85, 9436–9440 (1988).
Mazars, G.R. & Theillet, C. Direct sequencing by thermal asymmetric PCR. Methods Mol. Biol. 226, 355–360 (2003).
Sanchez, J.A. et al. Linear-after-the-exponential (LATE)-PCR: an advanced method of asymmetric PCR and its uses in quantitative real-time analysis. Proc. Natl. Acad. Sci. USA 101, 1933–1938 (2004).
Pierce, K.E. et al. Linear-after-the-exponential (LATE)-PCR: primer design criteria for high yields of specific single-stranded DNA and improved real-time detection. Proc. Natl. Acad. Sci. USA 102, 8609–8614 (2005).
Pierce, K.E. & Wangh, L.J. LATE-PCR and allied technologies: real-time detection strategies for rapid, reliable diagnosis from single cells. in Single Cell Diagnostics, Methods in Molecular Medicine Series (ed. Thornhill, A.) (Humana Press, Totowa, New Jersey, USA, 2007) p65–85.
Chou, Q. et al. Prevention of pre-PCR mis-priming and primer dimerization improves low-copy-number amplifications. Nucleic Acids Res. 20, 1717–1723 (1992).
Le Novere, N. MELTING, computing the melting temperature of nucleic acid duplex. Bioinformatics 17, 1226–1227 (2001).
Pierce, K.E. et al. Detection of cystic fibrosis alleles from single cells using molecular beacons and a novel method of asymmetric real-time PCR. Mol. Hum. Reprod. 9, 815–820 (2003).
Bustin, S.A. Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J. Mol. Endocrinol. 25, 169–193 (2000).
Salk, J.J. et al. Direct amplification of single-stranded DNA for pyrosequencing using linear-after-the-exponential (LATE)-PCR. Anal. Biochem. 353, 124–132 (2006).
Henegariu, O. et al. Multiplex PCR: critical parameters and step-by-step protocol. Biotechniques 23, 504–511 (1997).
Elnifro, E.M. et al. Multiplex PCR: optimization and application in diagnostic virology. Clin. Microbiol. Rev. 13, 559–570 (2000).
Markoulatos, P. et al. Multiplex polymerase chain reaction: a practical approach. J. Clin. Lab. Anal. 16, 47–51 (2002).
Tyagi, S. & Kramer, F.R. Molecular beacons: probes that fluoresce upon hybridization. Nat. Biotechnol. 14, 303–308 (1996).
Lee, M.A., Siddle, A.L. & Page, R.H. ResonSense: simple linear fluorescent probes for quantitative homogeneous rapid polymerase chain reaction. Anal. Chim. Acta 457, 61–70 (2002).
Sanchez, J.A. et al. Two-temperature LATE-PCR endpoint genotyping. BMC Biotechnol. 6, 44 (2006).
McCombie, W.R. et al. Rapid and reliable fluorescent cycle sequencing of double-stranded templates. DNA Seq. 2, 289–296 (1992).
Rosenthal, A. et al. Large-scale production of DNA sequencing templates by microtitre format PCR. Nucleic Acids Res. 21, 173–174 (1993).
Dowton, M. & Austin, A.D. Direct sequencing of double-stranded PCR products without intermediate fragment purification; digestion with mung bean nuclease. Nucleic Acids Res. 21, 3599–3600 (1993).
Hanke, M. & Wink, M. Direct DNA sequencing of PCR-amplified vector inserts following enzymatic degradation of primer and dNTPs. Biotechniques 17, 858–860 (1994).
Werle, E. et al. Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Res. 22, 4354–4355 (1994).
Hashimoto, M. et al. On-line integration of PCR and cycle sequencing in capillaries: from human genomic DNA directly to called bases. Nucleic Acids Res. 31, e41 (2003).
Pierce, K.E. et al. QuantiLyse: reliable DNA amplification from single cells. Biotechniques 32, 1106–1111 (2002).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
PrimeSafe® is an independent discovery of the Wangh Laboratory that has been licensed to Smiths Detection by Brandeis University. The inventors of this technology will receive a fraction of any future royalties received by Brandeis University through the sale of PrimeSafe®. The research described here has been supported in part by a grant from Smiths Detection.
Rights and permissions
About this article
Cite this article
Rice, J., Sanchez, J., Pierce, K. et al. Monoplex/multiplex linear-after-the-exponential-PCR assays combined with PrimeSafe and Dilute-'N'-Go sequencing. Nat Protoc 2, 2429–2438 (2007). https://doi.org/10.1038/nprot.2007.362
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.362
This article is cited by
-
Rapid Pyrazinamide Drug Susceptibility Testing using a Closed-Tube PCR Assay of the Entire pncA gene
Scientific Reports (2020)
-
Cancer diagnosis with DNA molecular computation
Nature Nanotechnology (2020)
-
On-site detection of Phytophthora spp.—single-stranded target DNA as the limiting factor to improve on-chip hybridization
Microchimica Acta (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.