Abstract
Multiple displacement amplification (MDA) is a recently described method of whole-genome amplification (WGA) that has proven efficient in the amplification of small amounts of DNA, including DNA from single cells. Compared with PCR-based WGA methods, MDA generates DNA with a higher molecular weight and shows better genome coverage. This protocol was developed for preimplantation genetic diagnosis, and details a method for performing single-cell MDA using the φ29 DNA polymerase. It can also be useful for the amplification of other minute quantities of DNA, such as from forensic material or microdissected tissue. The protocol includes the collection and lysis of single cells, and all materials and steps involved in the MDA reaction. The whole procedure takes 3 h and generates 1–2 μg of DNA from a single cell, which is suitable for multiple downstream applications, such as sequencing, short tandem repeat analysis or array comparative genomic hybridization.
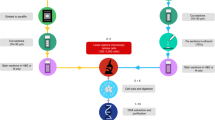
Note: In the version of this article initially published online, the third and fourth panels of Figure 1 (p. 1965) were incorrectly drawn. Figure 1 has been corrected in all versions of the article.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
21 December 2006
In the version of this article initially published online, the third and fourth panels of Figure 1 (p. 1965) were incorrectly drawn. Figure 1 has been corrected in all versions of the article.
References
Telenius, H. et al. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13, 718–725 (1992).
Zhang, L. et al. Whole genome amplification from a single cell: implications for genetic analysis. Proc. Natl. Acad. Sci. USA 89, 5847–5851 (1992).
Saunders, R.D. et al. PCR amplification of DNA microdissected from a single polytene chromosome band: a comparison with conventional microcloning. Nucleic Acids Res. 17, 9027–9037 (1989).
Cheung, V.G. & Nelson, S.F. Whole genome amplification using a degenerate oligonucleotide primer allows hundreds of genotypes to be performed on less than one nanogram of genomic DNA. Proc. Natl. Acad. Sci. USA 93, 14676–14679 (1996).
Paunio, T., Reima, I. & Syvanen, A.C. Preimplantation diagnosis by whole-genome amplification, PCR amplification, and solid-phase minisequencing of blastomere DNA. Clin. Chem. 42, 1382–1390 (1996).
Lizardi, P.M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nature Genet. 19, 225–232 (1998).
Dean, F.B., Nelson, J.R., Giesler, T.L. & Lasken, R.S. Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 11, 1095–1099 (2001).
Blanco, L. et al. Highly efficient DNA synthesis by the phage phi 29 DNA polymerase. Symmetrical mode of DNA replication. J. Biol. Chem. 264, 8935–8940 (1989).
Lage, J.M. et al. Whole genome analysis of genetic alterations in small DNA samples using hyperbranched strand displacement amplification and array-CGH. Genome Res. 13, 294–307 (2003).
Hellani, A. et al. Multiple displacement amplification on single cell and possible PGD applications. Mol. Hum. Reprod. 10, 847–852 (2004).
Handyside, A.H. et al. Isothermal whole genome amplification from single and small numbers of cells: a new era for preimplantation genetic diagnosis of inherited disease. Mol. Hum. Reprod. 10, 767–772 (2004).
Spits, C. et al. Optimization and evaluation of single-cell whole-genome multiple displacement amplification. Hum. Mutat. 27, 496–503 (2006).
Renwick, P.J. et al. Proof of principle and first cases using preimplantation genetic haplotyping — a paradigm shift for embryo diagnosis. Reprod. Biomed. Online 13, 110–119 (2006).
Hellani, A., Coskun, S., Tbakhi, A. & Al-Hassan, S. Clinical application of multiple displacement amplification in preimplantation genetic diagnosis. Reprod. Biomed. Online 10, 376–380 (2005).
Burlet, P. et al. Multiple displacement amplification improves PGD for fragile X syndrome. Mol. Hum. Reprod. 12, 647–652 (2006).
Jiang, Z., Zhang, X., Deka, R. & Jin, L. Genome amplification of single sperm using multiple displacement amplification. Nucleic Acids Res. 33, e91 (2005).
Ballantyne, K.N., van Oorschot, R.A., Mitchell, R.J. & Koukoulas, I. Molecular crowding increases the amplification success of multiple displacement amplification and short tandem repeat genotyping. Anal. Biochem. 355, 298–303 (2006).
LeCaignec, C. et al. Single-cell chromosomal imbalances detection by array CGH. Nucleic Acids Res. 34, e68 (2006).
Paez, J.G. et al. Genome coverage and sequence fidelity of phi29 polymerase-based multiple strand displacement whole genome amplification. Nucleic Acids Res. 32, e71 (2004).
In the version of this article initially published online, the third and fourth panels of Figure 1 (p. 1965) were incorrectly drawn. Figure 1 has been corrected in all versions of the article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Spits, C., Le Caignec, C., De Rycke, M. et al. Whole-genome multiple displacement amplification from single cells. Nat Protoc 1, 1965–1970 (2006). https://doi.org/10.1038/nprot.2006.326
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.326
This article is cited by
-
Development of preimplantation genetic testing for monogenic reference materials using next-generation sequencing
BMC Medical Genomics (2024)
-
Children born after assisted reproduction more commonly carry a mitochondrial genotype associating with low birthweight
Nature Communications (2024)
-
Identification of carrier status of Xp22.31 microdeletions associated with X-linked ichthyosis at the single-cell level using haplotype linkage analysis by karyomapping
Journal of Assisted Reproduction and Genetics (2023)
-
Limitations of gene editing assessments in human preimplantation embryos
Nature Communications (2023)
-
Isolation and sequencing of a single copy of an introgressed chromosome from a complex genome for gene and SNP identification
Theoretical and Applied Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.