Abstract
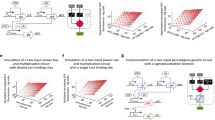
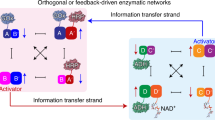
Biological organisms use complex molecular networks to navigate their environment and regulate their internal state. The development of synthetic systems with similar capabilities could lead to applications such as smart therapeutics or fabrication methods based on self-organization. To achieve this, molecular control circuits need to be engineered to perform integrated sensing, computation and actuation. Here we report a DNA-based technology for implementing the computational core of such controllers. We use the formalism of chemical reaction networks as a 'programming language' and our DNA architecture can, in principle, implement any behaviour that can be mathematically expressed as such. Unlike logic circuits, our formulation naturally allows complex signal processing of intrinsically analogue biological and chemical inputs. Controller components can be derived from biologically synthesized (plasmid) DNA, which reduces errors associated with chemically synthesized DNA. We implement several building-block reaction types and then combine them into a network that realizes, at the molecular level, an algorithm used in distributed control systems for achieving consensus between multiple agents.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Drexler, K. E. Molecular engineering: an approach to the development of general capabilities for molecular manipulation. Proc. Natl Acad. Sci. USA 78, 5275–5278 (1981).
Koo, O. M., Rubinstein, I. & Onyuksel, H. Role of nanotechnology in targeted drug delivery and imaging: a concise review. Nanomedicine: NBM 1, 193–212 (2005).
Hess, H. Engineering applications of biomolecular motors. Annu. Rev. Biomed. Eng. 13, 429–450 (2011).
Seeman, N. C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nature Chem. 3, 103–113 (2011).
Dirks, R. M. & Pierce, N. A. Triggered amplification by hybridization chain reaction. Proc. Natl Acad. Sci. USA 101, 15275–15278 (2004).
Seelig, G., Yurke, B. & Winfree, E. Catalyzed relaxation of a metastable DNA fuel. J. Am. Chem. Soc. 128, 12211–12220 (2006).
Turberfield, A. J. et al. DNA fuel for free-running nanomachines. Phys. Rev. Lett. 90, 118102 (2003).
Yin, P., Choi, H. M. T., Calvert, C. R. & Pierce, N. A. Programming biomolecular self-assembly pathways. Nature 451, 318–322 (2008).
Zhang, D. Y., Turberfield, A. J., Yurke, B. & Winfree, E. Engineering entropy-driven reactions and networks catalyzed by DNA. Science 318, 1121–1125 (2007).
Levy, M. & Ellington, A. D. Exponential growth by cross-catalytic cleavage of deoxyribozymogens. Proc. Natl Acad. Sci. USA 100, 6416–6421 (2003).
Cardelli, L. Two-domain DNA strand displacement. Math. Struct. Comput. Sci. 23, 247–271 (2013).
Oishi, K. & Klavins, E. Biomolecular implementation of linear I/O systems. IET Syst. Biol. 5, 252–260 (2011).
Phillips, A. & Cardelli, L. A programming language for composable DNA circuits. J. R. Soc. Interface 6, S419–S436 (2009).
Qian, L., Soloveichik, D. & Winfree, E. Efficient Turing-universal computation with DNA polymers. DNA Comput. Mol. Program. 6518, 123–140 (2011).
Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011).
Qian, L., Winfree, E. & Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 475, 368–372 (2011).
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Soloveichik, D., Seelig, G. & Winfree, E. DNA as a universal substrate for chemical kinetics. Proc. Natl Acad. Sci. USA 107, 5393–5398 (2010).
Stojanovic, M. N. & Stefanovic, D. A deoxyribozyme-based molecular automaton. Nature Biotechnol. 21, 1069–1074 (2003).
Benenson, Y. et al. Programmable and autonomous computing machine made of biomolecules. Nature 414, 430–434 (2001).
Kim, J. & Winfree, E. Synthetic in vitro transcriptional oscillators. Mol. Syst. Biol. 7, 465 (2011).
Montagne, K., Plasson, R., Sakai, Y., Fujii, T. & Rondelez, Y. Programming an in vitro DNA oscillator using a molecular networking strategy. Mol. Syst. Biol. 7, 466 (2011).
Willner, I., Shlyahovsky, B., Zayats, M. & Willner, B. DNAzymes for sensing, nanobiotechnology and logic gate applications. Chem. Soc. Rev. 37, 1153–1165 (2008).
Ran, T., Kaplan, S. & Shapiro, E. Molecular implementation of simple logic programs. Nature Nanotech. 4, 642–648 (2009).
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2010).
Omabegho, T., Sha, R. & Seeman, N. C. A bipedal DNA Brownian motor with coordinated legs. Science 324, 67–71 (2009).
Wickham, S. F. J. et al. Direct observation of stepwise movement of a synthetic molecular transporter. Nature Nanotech. 6, 166–169 (2011).
Muscat, R. A., Bath, J. & Turberfield, A. J. A programmable molecular robot. Nano Lett. 11, 982–987 (2011).
Yurke, B., Turberfield, A. J., Mills, A. P., Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012).
Hemphill, J. & Deiters, A. DNA Computation in mammalian cells: microRNA logic operations. J. Am. Chem. Soc. 135, 10512–10518 (2013).
Arkin, A. & Ross, J. Computational functions in biochemical reaction networks. Biophys. J. 67, 560–578 (1994).
Epstein, I. R. & Pojman, J. A. An Introduction to Nonlinear Chemical Dynamics: Oscillations, Waves, Patterns, and Chaos (Oxford Univ. Press, 1998).
Magnasco, M. O. Chemical kinetics is Turing universal. Phys. Rev. Lett. 78, 1190–1193 (1997).
Senum, P. & Riedel, M. Rate-independent constructs for chemical computation. PLoS ONE 6, e21414 (2011).
Soloveichik, D., Cook, M., Winfree, E. & Bruck, J. Computation with finite stochastic chemical reaction networks. Nat. Comput. 7, 615–633 (2008).
Tyson, J. J., Chen, K. C. & Novak, B. Sniffers, buzzers, toggles and blinkers: dynamics of regulatory and signaling pathways in the cell. Curr. Opin. Cell Biol. 15, 221–231 (2003).
Peterson, J. L. Petri Net Theory and the Modeling of Systems 290 (Prentice-Hall, 1981).
Zhang, D. Y. & Winfree, E. Robustness and modularity properties of a non-covalent DNA catalytic reaction. Nucleic Acids Res. 38, 4182–4197 (2010).
Lin, C. et al. In vivo cloning of artificial DNA nanostructures. Proc. Natl Acad. Sci. USA 105, 17626–17631 (2008).
Ducani, C., Kaul, C., Moche, M., Shih, W. M. & Högberg, B. Enzymatic production of 'monoclonal stoichiometric' single-stranded DNA oligonucleotides. Nature Methods 10, 647–652 (2013).
Chen, X., Briggs, N., McLain, J. R. & Ellington, A. D. Stacking nonenzymatic circuits for high signal gain. Proc. Natl Acad. Sci. USA 110, 5386–5391 (2013).
Angluin, D., Aspnes, J. & Eisenstat, D. A simple population protocol for fast robust approximate majority. Distrib. Comput. 21, 87–102 (2008).
Zhang, D. Y. & Winfree, E. Control of DNA strand displacement kinetics using toehold exchange. J. Am. Chem. Soc. 131, 17303–17314 (2009).
Lakin, M. R., Youssef, S., Cardelli, L. & Phillips, A. Abstractions for DNA circuit design. J. R. Soc. Interface 9, 470–486 (2012).
Cardelli, L. & Csikász-Nagy, A. The cell cycle switch computes approximate majority. Sci. Rep. 2, 656 (2012).
Zhang, D. & Seelig, G. in DNA Computing and Molecular Programming (eds Sakakibara, Y. & Mi, Y.) Vol. 6518, 176–186 (Lecture Notes in Computer Science, Springer, 2011).
Acknowledgements
The authors thank E. Winfree, E. Klavins and D.Y. Zhang for discussions and comments on the manuscript. This work was supported by the National Science Foundation (grant NSF-CCF 1117143 to G.S. and D.S.). G.S. was supported by a Burroughs Wellcome Career Award at the Scientific Interface. D.S. was supported by an NIGMS Systems Biology Center grant (P50 GM081879).
Author information
Authors and Affiliations
Contributions
All authors designed the experiments and co-wrote the paper. Y-J.C. performed the wetlab experiments. N.D. and A.P. performed computational experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Information (PDF 4054 kb)
Rights and permissions
About this article
Cite this article
Chen, YJ., Dalchau, N., Srinivas, N. et al. Programmable chemical controllers made from DNA. Nature Nanotech 8, 755–762 (2013). https://doi.org/10.1038/nnano.2013.189
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2013.189
This article is cited by
-
Real-time signal processing via chemical reactions for a microfluidic molecular communication system
Nature Communications (2023)
-
Generation of DNA oligomers with similar chemical kinetics via in-silico optimization
Communications Chemistry (2023)
-
A nanopore interface for higher bandwidth DNA computing
Nature Communications (2022)
-
Molecular device design based on chemical reaction networks: state feedback controller, static pre-filter, addition gate control system and full-dimensional state observer
Journal of Mathematical Chemistry (2022)