Abstract
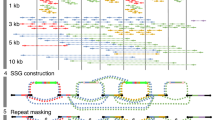
High-throughput sequencing technologies promise to transform the fields of genetics and comparative biology by delivering tens of thousands of genomes in the near future. Although it is feasible to construct de novo genome assemblies in a few months, there has been relatively little attention to what is lost by sole application of short sequence reads. We compared the recent de novo assemblies using the short oligonucleotide analysis package (SOAP), generated from the genomes of a Han Chinese individual and a Yoruban individual, to experimentally validated genomic features. We found that de novo assemblies were 16.2% shorter than the reference genome and that 420.2 megabase pairs of common repeats and 99.1% of validated duplicated sequences were missing from the genome. Consequently, over 2,377 coding exons were completely missing. We conclude that high-quality sequencing approaches must be considered in conjunction with high-throughput sequencing for comparative genomics analyses and studies of genome evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Huang, S. et al. The genome of the cucumber, Cucumis sativus L. Nat. Genet. 41, 1275–1281 (2009).
Li, R. et al. The sequence and de novo assembly of the giant panda genome. Nature 463, 311–317 (2010).
Genome 10K Community of Scientists. Genome 10K: a proposal to obtain whole-genome sequence for 10,000 vertebrate species. J. Hered. 100, 659–674 (2009).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Wheeler, D.A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Myers, E.W. et al. A whole-genome assembly of Drosophila. Science 287, 2196–2204 (2000).
Pevzner, P.A., Tang, H. & Waterman, M.S. An Eulerian path approach to DNA fragment assembly. Proc. Natl. Acad. Sci. USA 98, 9748–9753 (2001).
Chaisson, M.J., Brinza, D. & Pevzner, P.A. De novo fragment assembly with short mate-paired reads: Does the read length matter? Genome Res. 19, 336–346 (2009).
Simpson, J.T. et al. ABySS: a parallel assembler for short read sequence data. Genome Res. 19, 1117–1123 (2009).
Schuster, S.C. et al. Complete Khoisan and Bantu genomes from southern Africa. Nature 463, 943–947 (2010).
Green, P. Whole-genome disassembly. Proc. Natl. Acad. Sci. USA 99, 4143–4144 (2002).
Schatz, M.C., Delcher, A.L. & Salzberg, S.L. Assembly of large genomes using second-generation sequencing. Genome Res. 20, 1165–1173 (2010).
Meader, S., Hillier, L.W., Locke, D., Ponting, C.P. & Lunter, G. Genome assembly quality: assessment and improvement using the neutral indel model. Genome Res. 20, 675–684 (2010).
Zhang, Z., Schwartz, S., Wagner, L. & Miller, W. A greedy algorithm for aligning DNA sequences. J. Comput. Biol. 7, 203–214 (2000).
Li, R. et al. Building the sequence map of the human pan-genome. Nat. Biotechnol. 28, 57–63 (2010).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
Mills, R.E., Bennett, E.A., Iskow, R.C. & Devine, S.E. Which transposable elements are active in the human genome? Trends Genet. 23, 183–191 (2007).
Bailey, J.A., Yavor, A.M., Massa, H.F., Trask, B.J. & Eichler, E.E. Segmental duplications: organization and impact within the current human genome project assembly. Genome Res. 11, 1005–1017 (2001).
She, X. et al. Shotgun sequence assembly and recent segmental duplications within the human genome. Nature 431, 927–930 (2004).
Alkan, C. et al. Personalized copy number and segmental duplication maps using next-generation sequencing. Nat. Genet. 41, 1061–1067 (2009).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Doggett, N.A. et al. A 360-kb interchromosomal duplication of the human HYDIN locus. Genomics 88, 762–771 (2006).
Worley, K.C. & Gibbs, R.A. Genetics: decoding a national treasure. Nature 463, 303–304 (2010).
Kidd, J.M. et al. Characterization of missing human genome sequences and copy-number polymorphic insertions. Nat. Methods 7, 365–371 (2010).
Acknowledgements
We thank E. Karakoc and P. Sudmant for helpful discussions, T. Marques-Bonet and J.M. Kidd for providing the nonredundant gene table, and T. Brown for proofreading the manuscript. This work was partly supported by US National Institutes of Health grant HG002385 to E.E.E. E.E.E. receives funds as an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.A. and E.E.E. conceived the study and wrote the manuscript. C.A. and S.S. analyzed the data.
Corresponding author
Ethics declarations
Competing interests
E.E.E. is a scientific advisory board member of Pacific Biosciences.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–2, Supplementary Table 2, Supplementary Note (PDF 402 kb)
Supplementary Table 1
Contamination found in reported human new sequence insertions from the genomes of two individuals. (XLS 406 kb)
Supplementary Table 3
Analysis of nonredundant autosomal genes in the YH genome assembly. (XLS 5110 kb)
Supplementary Table 4
Analysis of nonredundant autosomal coding exons in the YH genome. NOTE: This is a tab-delimited text file with 171,751 rows of data. Confirm that all data will load into your application before proceeding. (TXT 12294 kb)
Supplementary Table 5
Assigned positions of duplicated sequences (YH) to the NCBI build 36 assembly. (XLS 2229 kb)
Rights and permissions
About this article
Cite this article
Alkan, C., Sajjadian, S. & Eichler, E. Limitations of next-generation genome sequence assembly. Nat Methods 8, 61–65 (2011). https://doi.org/10.1038/nmeth.1527
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1527