Abstract
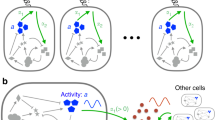
Biology emerges from interactions between molecules, which are challenging to elucidate with current techniques. An orthogonal approach is to probe for 'response signatures' that identify specific circuit motifs. For example, bistability, hysteresis, or irreversibility are used to detect positive feedback loops. For adapting systems, such signatures are not known. Only two circuit motifs generate adaptation: negative feedback loops (NFLs) and incoherent feed-forward loops (IFFLs). On the basis of computational testing and mathematical proofs, we propose differential signatures: in response to oscillatory stimulation, NFLs but not IFFLs show refractory–period stabilization (robustness to changes in stimulus duration) or period skipping. Applying this approach to yeast, we identified the circuit dominating cell cycle timing. In Caenorhabditis elegans AWA neurons, which are crucial for chemotaxis, we uncovered a Ca2+ NFL leading to adaptation that would be difficult to find by other means. These response signatures allow direct access to the outlines of the wiring diagrams of adapting systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Pomerening, J.R., Sontag, E.D. & Ferrell, J.E. Jr. Building a cell cycle oscillator: hysteresis and bistability in the activation of Cdc2. Nat. Cell Biol. 5, 346–351 (2003).
Xiong, W. & Ferrell, J.E. Jr. A positive-feedback-based bistable 'memory module' that governs a cell fate decision. Nature 426, 460–465 (2003).
Charvin, G., Oikonomou, C., Siggia, E.D. & Cross, F.R. Origin of irreversibility of cell cycle start in budding yeast. PLoS Biol. 8, e1000284 (2010).
Alberts, B. et al. Molecular Biology of the Cell (Garland Science, 2015).
Behar, M., Hao, N., Dohlman, H.G. & Elston, T.C. Mathematical and computational analysis of adaptation via feedback inhibition in signal transduction pathways. Biophys. J. 93, 806–821 (2007).
Ma, W., Trusina, A., El-Samad, H., Lim, W.A. & Tang, C. Defining network topologies that can achieve biochemical adaptation. Cell 138, 760–773 (2009).
Ascensao, J.A. et al. Non-monotonic response to monotonic stimulus: Regulation of glyoxylate shunt gene-expression dynamics in Mycobacterium tuberculosis. PLOS Comput. Biol. 12, e1004741 (2016).
Alon, U. An Introduction to Systems Biology (Chapman and Hall/CRC, 2007).
Friedlander, T. & Brenner, N. Adaptive response by state-dependent inactivation. Proc. Natl. Acad. Sci. USA 106, 22558–22563 (2009).
Ferrell, J.E. Jr. Perfect and near-perfect adaptation in cell signaling. Cell Syst. 2, 62–67 (2016).
Lim, S. et al. Negative feedback governs gonadotrope frequency-decoding of gonadotropin releasing hormone pulse-frequency. PLoS One 4, e7244 (2009).
Tsaneva-Atanasova, K., Mina, P., Caunt, C.J., Armstrong, S.P. & McArdle, C.A. Decoding GnRH neurohormone pulse frequency by convergent signalling modules. J. R. Soc. Interface 9, 170–182 (2012).
Tyson, J.J., Chen, K.C. & Novak, B. Sniffers, buzzers, toggles and blinkers: dynamics of regulatory and signaling pathways in the cell. Curr. Opin. Cell Biol. 15, 221–231 (2003).
Castillo-Hair, S.M., Igoshin, O.A. & Tabor, J.J. How to train your microbe: methods for dynamically characterizing gene networks. Curr. Opin. Microbiol. 24, 113–123 (2015).
Segall, J.E., Block, S.M. & Berg, H.C. Temporal comparisons in bacterial chemotaxis. Proc. Natl. Acad. Sci. USA 83, 8987–8991 (1986).
Geva-Zatorsky, N., Dekel, E., Batchelor, E., Lahav, G. & Alon, U. Fourier analysis and systems identification of the p53 feedback loop. Proc. Natl. Acad. Sci. USA 107, 13550–13555 (2010).
Mitchell, A., Wei, P. & Lim, W.A. Oscillatory stress stimulation uncovers an Achilles' heel of the yeast MAPK signaling network. Science 350, 1379–1383 (2015).
Heltberg, M., Kellogg, R.A., Krishna, S., Tay, S. & Jensen, M.H. Noise induces hopping between NF-κB entrainment modes. Cell Syst. 3, 532–539.e3 (2016).
Block, S.M., Segall, J.E. & Berg, H.C. Adaptation kinetics in bacterial chemotaxis. J. Bacteriol. 154, 312–323 (1983).
Tu, Y., Shimizu, T.S. & Berg, H.C. Modeling the chemotactic response of Escherichia coli to time-varying stimuli. Proc. Natl. Acad. Sci. USA 105, 14855–14860 (2008).
Muzzey, D., Gómez-Uribe, C.A., Mettetal, J.T. & van Oudenaarden, A. A systems-level analysis of perfect adaptation in yeast osmoregulation. Cell 138, 160–171 (2009).
Wang, C.J., Bergmann, A., Lin, B., Kim, K. & Levchenko, A. Diverse sensitivity thresholds in dynamic signaling responses by social amoebae. Sci. Signal. 5, ra17 (2012).
Takeda, K. et al. Incoherent feedforward control governs adaptation of activated ras in a eukaryotic chemotaxis pathway. Sci. Signal. 5, ra2 (2012).
Larsch, J. et al. A circuit for gradient climbing in C. elegans chemotaxis. Cell Rep. 12, 1748–1760 (2015).
Shoval, O. et al. Fold-change detection and scalar symmetry of sensory input fields. Proc. Natl. Acad. Sci. USA 107, 15995–16000 (2010).
Adler, M., Szekely, P., Mayo, A. & Alon, U. Optimal regulatory circuit topologies for fold-change detection. Cell Syst. 4, 171–181.e8 (2017).
Morgan, D.O. The Cell Cycle: Principles of Control (New Science Press, 2007).
Simon, I. et al. Serial regulation of transcriptional regulators in the yeast cell cycle. Cell 106, 697–708 (2001).
Lee, T.I. et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298, 799–804 (2002).
Orlando, D.A. et al. Global control of cell-cycle transcription by coupled CDK and network oscillators. Nature 453, 944–947 (2008).
Simmons Kovacs, L.A. et al. Cyclin-dependent kinases are regulators and effectors of oscillations driven by a transcription factor network. Mol. Cell 45, 669–679 (2012).
Rahi, S.J., Pecani, K., Ondracka, A., Oikonomou, C. & Cross, F.R. The CDK-APC/C oscillator predominantly entrains periodic cell-cycle transcription. Cell 165, 475–487 (2016).
Charvin, G., Cross, F.R. & Siggia, E.D. Forced periodic expression of G1 cyclins phase-locks the budding yeast cell cycle. Proc. Natl. Acad. Sci. USA 106, 6632–6637 (2009).
Wäsch, R. & Cross, F.R. APC-dependent proteolysis of the mitotic cyclin Clb2 is essential for mitotic exit. Nature 418, 556–562 (2002).
Drapkin, B.J., Lu, Y., Procko, A.L., Timney, B.L. & Cross, F.R. Analysis of the mitotic exit control system using locked levels of stable mitotic cyclin. Mol. Syst. Biol. 5, 328 (2009).
Hilliard, M.A. et al. In vivo imaging of C. elegans ASH neurons: cellular response and adaptation to chemical repellents. EMBO J. 24, 63–72 (2005).
Rengarajan, S. & Hallem, E.A. Olfactory circuits and behaviors of nematodes. Curr. Opin. Neurobiol. 41, 136–148 (2016).
Larsch, J., Ventimiglia, D., Bargmann, C.I. & Albrecht, D.R. High-throughput imaging of neuronal activity in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 110, E4266–E4273 (2013).
Colbert, H.A. & Bargmann, C.I. Odorant-specific adaptation pathways generate olfactory plasticity in C. elegans. Neuron 14, 803–812 (1995).
Kato, S., Xu, Y., Cho, C.E., Abbott, L.F. & Bargmann, C.I. Temporal responses of C. elegans chemosensory neurons are preserved in behavioral dynamics. Neuron 81, 616–628 (2014).
Zwaal, R.R. et al. The sarco-endoplasmic reticulum Ca2+ ATPase is required for development and muscle function in Caenorhabditis elegans. J. Biol. Chem. 276, 43557–43563 (2001).
Shoval, O., Alon, U. & Sontag, E. Symmetry invariance for adapting biological systems. SIAM J. Appl. Dyn. Syst. 10, 857–886 (2011).
Ferrell, J.E. Jr. Self-perpetuating states in signal transduction: positive feedback, double-negative feedback and bistability. Curr. Opin. Cell Biol. 14, 140–148 (2002).
Acknowledgements
We thank C.I. Bargmann for mentorship, support, and comments on the manuscript. We thank E. Siggia for fruitful discussions. The work was supported by US National Institutes of Health grant 5RO1-GM078153-07 (F.R.C.), NRSA Training Grant CA009673-36A1 (S.J.R.), a Merck Postdoctoral Fellowship at The Rockefeller University (S.J.R.), and the Simons Foundation (S.J.R.). J.L. was supported by a fellowship from the Boehringer Ingelheim Fonds. E.D.S. was partially supported by the US Office of Naval Research (ONR N00014-13-1-0074) and the US Air Force Office of Scientific Research (AFOSR FA9550-14-1-0060).
Author information
Authors and Affiliations
Contributions
Conceptualization and writing: S.J.R., J.L., K.P., A.Y.K., N.M., K.T.-A., E.S., and F.R.C. Experiments and data analysis: S.J.R., J.L., and K.P. Mathematical proofs: S.J.R. and E.D.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Examples showing that IFFLs and NFLs can show qualitatively similar responses to linearly ramping, exponentially ramping, or on-off stimuli; fits to the same experimental data; model circuits under continuous stimulation; and examples of period skipping for NFLs.
A-H: Examples showing that various response characteristics from the literature can appear in both IFFLs and NFLs. Adapted from refs. (8, 19, 20, 21, 22, 23) θ(t) is a step function, which is zero for t < 0 and equal to one for t > 0. The linear ramp tθ(t) is equal to t for t > 0 and equal to zero for t < 0. The exponential ramp (ext - 1)×θ(t) is equal to (ext - 1) for t > 0 and equal to zero for t < 0. A, B: Adaptation to step inputs and linear ramps; C, D: adaptation to step inputs but failure to adapt to linear ramps; E, F: adaptation to step inputs but failure to adapt to exponential ramps; G, H: overshoot below steady-state when the input is turned off (red dashed line, switch S = 1 to S = 0 at t = 0). I, J: Fits to the same experimental data: Black: The 15’th response pulses at odor pulse duration d = 10’’ and period T = 42’’ were taken from our C. elegans recordings, averaged over all worms, and then normalized to the mean of the 9’th and 10’th response pulses. Red: We guessed parameters for each model and then fine-tuned them to the data by a steepest-descent algorithm minimizing the root mean square deviation (RMSD). K, L: Examples showing the behavior of the circuits in Fig. 1 E, F (same as Fig. 1 I, J) under continuous stimulation. With long stimulus (S) pulses (blue), the response R is effectively shut off from the stimulus S when I exceeds I0 in the model circuits in Fig. 1 E, F (same as Fig. 1 I, J). In the NFL circuit, I can drop below I0 again while the stimulus is on, allowing the stimulus S to cause another spike in R, and repeat. However, softening the step inhibition function (here: θ(I0 - I)) easily abolishes sustained oscillations (not shown). For the model calculations in Fig. 1 J, we plot Tmax(d) for pulse durations d before there is a second response spike elicited by the same stimulus pulse. M, N: Examples of period skipping with 4T-periodic (M, one NFL) or more complicated (N, interlocking NFLs) responses to T-periodic stimuli. Peaks are marked by black circles. Periodic stimuli are not shown. K-N: Levels of Stimulus, Inhibitor, and Output (color-coded) indicated in arbitrary units as a function of time.
Supplementary Figure 2 Systematic enumeration of 3+1 NFL and 3+1 IFFL circuit topologies for computational analysis.
Each circuit topology is associated with a set of equations that are indicated. These models were further diversified by replacing I(t), R(t), X(t), Z(t) by other functional forms, e.g., I(t) → I(t)/(1 + I(t)/0.1), as specified. To keep the number of combinations computationally tractable, the number of replacements had to be limited as specified. The numbers next to the circuit names indicate the number of resulting combinations of parameters and functional forms that were evaluated.
Supplementary Figure 3 Systematic enumeration of 4+1 IFFL circuit topologies with an intervening upstream node.
Also see Supplementary Figure 2.
Supplementary Figure 4 Systematic enumeration of 4+1 IFFL circuit topologies with a node between the stimulus S and the response node R.
Also see Supplementary Figure 2.
Supplementary Figure 5 Systematic enumeration of 4+1 IFFL circuit topologies with an adapting node Z, which is depressed by the stimulus and which in turn depresses the response node R.
Also see Supplementary Figure 2.
Supplementary Figure 6 Systematic enumeration of 4+1 IFFL circuit topologies with a node between the stimulus S and the inhibitory node I (or the activating node X).
Also see Supplementary Figure 2.
Supplementary Figure 7 Examples of Tmax(d) plots from the computational search.
Examples of Tmax(d) plots from the computational search summarized in Table 1. A-C: λ is arbitrary. A: An IFFL model with simple (non-cooperative) Michaelis-Menten inhibition. B: An NFL model with simple (non-cooperative) Michaelis-Menten inhibition showing that Tmax(d) can even have a negative slope. C: An NFL model in which the inhibitor I degrades the response R.
Supplementary Figure 8 Detailed analysis of NFL circuits.
Detailed analysis of NFL circuits in Fig. 1 F, J showing that Tmax(d) (grey lines) and pulse periods and widths leading to period skipping (magenta-green-blue triangles below magenta lines) are close for a variety of model parameters. Thus, a search for Tmax may be expected to lead to observing period skipping as well, if it occurs. Grey lines: Tmax(d) plotted with the same shades of grey as in Fig. 1 J. Magenta lines: In the triangles between the green, blue, and magenta lines (below the magenta lines), periodic solutions with the same period T as the stimulus period are unstable and period skipping is observed. Blue dashed lines: The regions of interest for our analysis are to the left of the blue dashed lines in each panel; to the right of the blue dashed lines, each stimulus pulse is so long that the circuits respond at least twice to each stimulus pulse (R goes up at least twice), because each stimulus pulse extends beyond the time when the circuit recovers from adaptation, i.e., when I drops below I0 again, and can be activated again. This may be a feature of these models that is not observed in many biological NFLs and thus we refrain from analyzing the models in this regime. Green dashed lines: To the left of the green dashed lines, the pulse durations d are too short (alternatively, the pulse periods T too long) for enough inhibitor I to accumulate during each periodic stimulus pulse to block the stimulus pulses at all and thus for the responses to show any adaptation; θ(I0 - I) is always equal to 1. To the right of the green dashed lines, the pulse durations d are long enough (alternatively, the pulse periods T short enough) for enough inhibitor to accumulate during each stimulus pulse to block the stimulus pulses at some point after the onset of each stimulus pulse. For any fixed pulse period T, making the pulse durations d longer than specified at the green boundary has the same effect as stimulus pulses with pulse durations at the green boundary because the inhibitor cuts the stimulus off (θ(I0 - I) = 0) for pulse durations to the right of the boundary line.
Supplementary Figure 9 Disentangling multiple circuit motifs.
A: Tmax(d) for a pathway combining an IFFL circuit (through I1) with slower kinetics and an NFL circuit (through I2) with faster kinetics, acting in parallel. After the onset of on-off pulsing, the NFL circuit influences the output quickly whereas the IFFL circuit affects the output more slowly. So, we measured the running average of the output (= O(d, T)) at different time points 50, 150,... after the onset of the periodic on-off stimulus. For the NFL-only circuit (I2 set to 0, dashed lines), which is presented for comparison, Tmax(d) is strongly stabilized and in fact decreases at intermediate d. In the full circuit, as O(d, T) is defined at later time points, Tmax stabilization slowly disappears (black→blue→green→cyan). (dI1/dt = S - λ1I1, dI2/dt = R - λ2I2, dR/dt = S/(1 + (I2/I0)n) - (1 + κI1)R, O(t) = R3(t), λ1 = 1/200, λ2 = 2/10, κ = 1/100, n = 1, I0 = 1/10).
Supplementary Figure 10 Processing of C. elegans experimental data and additional pharmacological experiments.
For details, see Methods. In all experiments, the main odor pulses (here: d = 20’’, T = 42’’) were preceded by 10 preparatory odor pulses of duration 10’’ and period 60’’. A: One imaged frame (cropped, of about 20000 total frames per experiment, recorded at 10 Hz) showing GCaMP fluorescence from the AWA neuron pair of a single worm. FR,i(t) is the average intensity over the area indicated by a green square, centered on the AWA neurons, and FBG,i(t) is the median intensity over the region indicated by a red ring. Scale bar: 100μm. B: FnoBG,i(t) = FR,i(t) - FBG,i(t) plotted for different worms from the same experiment. The red trace is filtered out because the baseline moves too much (6.5%) before and after the preparatory odor pulse 10. The orange trace is discarded because the noise-to-signal ratio (0.11-0.15) is too high. See Methods for more details. C: A piecewise linear function FBL,i(t) (blue) representing the time-dependent baseline fluorescence is fit through FnoBG,i(t) before each response pulse. D: Plot of Fi(t) = FnoBG,i(t)/FBL,i(t)-1, which is corrected for the baseline fluorescence FBL,i(t). E: Fi(t) is plotted for the preparatory pulses and its running average over a time window of size T is plotted for the main pulses. The mean of Fi(t) over the preparatory pulses 9 and 10 (= Fnorm,i, area indicated in red divided by 120’’) is shown as a horizontal red bar. F: Fi(t) is normalized by Fnorm,i; then, the normalized running average is fit to a straight line between 100’’-800’’ after the beginning of the main odor pulses. G: The value of the linear fit at 100’’ after the beginning of the main odor pulses represents the output data point Oi(d, T) for worm i at pulse duration d and period T. H, I: Fig. 3 I underlain for comparison. H: Same as Fig. 3 E except thapsigargin applied longer (for additional three odor pulse periods). Mean over 13 worms. I: Thapsigargin applied but odor stimulus pulses turned off. Thapsigargin presentation in the absence of odor only caused minor and brief increases in Ca2+, potentially, by disrupting baseline Ca2+ maintenance. Mean over 14 worms.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Table 1 and Supplementary Notes 1–3.
Supplementary Software
MATLAB code for computational exploration of model circuits.
Rights and permissions
About this article
Cite this article
Rahi, S., Larsch, J., Pecani, K. et al. Oscillatory stimuli differentiate adapting circuit topologies. Nat Methods 14, 1010–1016 (2017). https://doi.org/10.1038/nmeth.4408
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4408
This article is cited by
-
Automated neuron tracking inside moving and deforming C. elegans using deep learning and targeted augmentation
Nature Methods (2024)
-
Multidimensional characterization of inducible promoters and a highly light-sensitive LOV-transcription factor
Nature Communications (2023)
-
Automatic synchronisation of the cell cycle in budding yeast through closed-loop feedback control
Nature Communications (2021)
-
Cyclin/Forkhead-mediated coordination of cyclin waves: an autonomous oscillator rationalizing the quantitative model of Cdk control for budding yeast
npj Systems Biology and Applications (2021)
-
A convolutional neural network segments yeast microscopy images with high accuracy
Nature Communications (2020)