Abstract
Prior biological knowledge and phenotype information may help to identify disease genes from human whole-genome and whole-exome sequencing studies. We developed Phenolyzer (http://phenolyzer.usc.edu), a tool that uses prior information to implicate genes involved in diseases. Phenolyzer exhibits superior performance over competing methods for prioritizing Mendelian and complex disease genes, based on disease or phenotype terms entered as free text.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Lyon, G.J. & Wang, K. Genome Med. 4, 58 (2012).
Wang, K., Li, M. & Hakonarson, H. Nucleic Acids Res. 38, e164 (2010).
Cingolani, P. et al. Fly (Austin) 6, 80–92 (2012).
McLaren, W. et al. Bioinformatics 26, 2069–2070 (2010).
Jäger, M. et al. Hum. Mutat. 35, 548–555 (2014).
Habegger, L. et al. Bioinformatics 28, 2267–2269 (2012).
Bamshad, M.J. et al. Nat. Rev. Genet. 12, 745–755 (2011).
Aerts, S. et al. Nat. Biotechnol. 24, 537–544 (2006).
Schlicker, A., Lengauer, T. & Albrecht, M. Bioinformatics 26, i561–i567 (2010).
Singleton, M.V. et al. Am. J. Hum. Genet. 94, 599–610 (2014).
Javed, A., Agrawal, S. & Ng, P.C. Nat. Methods 11, 935–937 (2014).
Safran, M. et al. Database 2010, baq020 (2010).
Makita, Y. et al. Nucleic Acids Res. 41, W109–W114 (2013).
Yue, P., Melamud, E. & Moult, J. BMC Bioinformatics 7, 166 (2006).
Köhler, S. et al. Am. J. Hum. Genet. 85, 457–464 (2009).
Robinson, P.N. et al. Genome Res. 24, 340–348 (2014).
Chial, H. Nat. Educ. 1, 192 (2008).
Saunders, C.J. et al. Sci. Translat. Med. 4, 154ra135 (2012).
Forbes, S.A. et al. Nucleic Acids Res. 39, D945–D950 (2011).
De Rubeis, S. et al. Nature 515, 209–215 (2014).
Zhang, R. et al. Database 2014, bau090 (2014).
van der Harst, P. et al. Nature 492, 369–375 (2012).
Chang, X. & Wang, K. J. Med. Genet. 49, 433–436 (2012).
Amberger, J., Bocchini, C. & Hamosh, A. Hum. Mutat. 32, 564–567 (2011).
Rath, A. et al. Hum. Mutat. 33, 803–808 (2012).
Landrum, M.J. et al. Nucleic Acids Res. 42, D980–D985 (2014).
Pagon, R.A. et al. GeneReviewshttp://www.ncbi.nlm.nih.gov/books/NBK1116/?partid=1250 (1993).
Hindorff, L.A., Junkins, H.A., Mehta, J. & Manolio, T. A Catalog of Published Genome-Wide Association Studies. https://www.genome.gov/26525384 (National Human Genome Research Institute, 2011).
Peri, S. et al. Nucleic Acids Res. 32, D497–D501 (2004).
Geer, L.Y. et al. Nucleic Acids Res. 38, D492–D496 (2010).
Seal, R.L., Gordon, S.M., Lush, M.J., Wright, M.W. & Bruford, E.A. Nucleic Acids Res. 39, D514–D519 (2011).
Bovolenta, L.A., Acencio, M.L. & Lemke, N. BMC Genomics 13, 405 (2012).
Davis, A.P., Wiegers, T.C., Rosenstein, M.C. & Mattingly, C.J. Database 2012, bar065 (2012).
Schriml, LM. et al. Nucleic Acids Res. 40, D940–D946 (2012).
Robinson, P.N. & Mundlos, S. Clin. Genet. 77, 525–534 (2010).
Burren, O.S. et al. Nucleic Acids Res. 39, D997–D1001 (2011).
Lim, J.E. et al. BMC Med. Inform. Decis. Mak. 10, 76 (2010).
Elding, H., Lau, W., Swallow, D.M. & Maniatis, N. Am. J. Hum. Genet. 92, 107–113 (2013).
Liu, H. et al. Nucleic Acids Res. 39, D991–D996 (2011).
Acknowledgements
This work was supported by US National Institutes of Health grant R01-HG006465 to K.W. We thank members of the Wang laboratory for testing the Phenolyzer website.
Author information
Authors and Affiliations
Contributions
H.Y. compiled the data, performed the computational experiments, developed software tools and drafted the manuscript. P.N.R. advised on phenotype data analysis and interpretation. K.W. designed the study, supervised its execution and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
K.W. is a board member and stock holder of Tute Genomics, a bioinformatics software company.
Integrated supplementary information
Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'.
The largest word represents the user’s input term, ‘Autism’. The pink round rectangles represent disease names corresponding to each term. The balls represent genes. The darker the color, the more the node contributes. The longer the round rectangles, the more a disease name contributes. The larger the ball, the more significantly a gene is related with the term. A reported gene is blue and a predicted gene is yellow. Four types of edges correspond to four different types of gene-gene relationships as illustrated in the legend. The figure can be zoomed in by mouse operations in the web server to facilitate closer examination of the edge types.
Supplementary Figure 2 The wordcloud for all the interpreted names related to ‘Cancer’.
This is the wordcloud generated by Phenolyzer corresponding to the input term ‘cancer’. Some of the most frequently occurring words include ‘cancer’, ‘neoplasm’, ‘cell’, ‘carcinoma’, ‘tumor’ and ‘malignant’.
Supplementary Figure 3 A snapshot of the output from the Phenolyzer web server.
(a) ‘Summary’ includes the link to wordcloud, as well as the input settings and the links to output files for download. (b) ‘Barplot’ is a bar plot on at most 500 highest-ranked genes with normalized scores. (c) ‘Network’ is the interactive gene-disease-term network, with controlling buttons below it. (d) ‘Details’ shows how the score for each gene is calculated, including the links to each publication or database website.
Supplementary Figure 4 Comparison between Phenolyzer and other tools to find disease genes for cancer, rheumatoid arthritis, autism and anemia.
The AUC and ROC curve plot showing the performance comparison between Phenolyzer and other tools, on four gene sets of different complex diseases. (a, c, e and g) For each software, the AUC is calculated as the area under the ROC curve. (b, d, f and h) The ROC curve is plotted as True Positive Rate versus False Positive Rate. ‘Phenolyzer Phenotype’ is the Phenolyzer results with phenotype terms as input (the same input as Phenomizer). ‘Phenolyzer Logistic’ is Phenolyzer with weights trained with Logistic Regression model, compared with ‘Phenolyzer no training’. ‘Phenolyzer Seed’ is Phenolyzer’s seed gene result without the seed gene growth step, thus only representing the genes found in Phenolyzer’s disease-gene mapping knowledgebase.
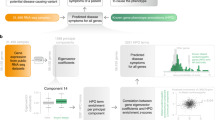
Supplementary Figure 5 Evaluation of Phenolyzer by phenotype terms as input.
Phenolyzer with phenotype terms (rather than disease names) as input is able to prioritize most genes as ‘Top 1’ for the 14 monogenic diseases, which is similar as Phenomizer.
Supplementary Figure 6 Phenolyzer’s results on four case studies.
(a and b) The candidate gene lists generated from two studies on ‘Craniopharyngiomas’ and ‘SHORT syndrome’ were used as input into Phenolyer. The network plot shows that BRAF and PIK3R1 are the genes with the highest scores corresponding to each disease separately. (c) For the CNV study of ‘Osteoporosis’, the generated significant CNV regions were used as input, and the Phenolyzer network successfully identified the correct gene, UGT2B17. (d) Combined with wANNOVAR, we first filtered the variants into a small list, then included all the genes in the variant list as the input into Phenolyzer. The correct gene PKLR was identified as the top gene for ‘hemolytic anemia’.
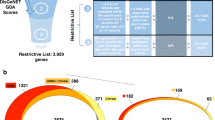
Supplementary Figure 7 Illustration of the disease or phenotype term interpretation process.
The term is first processed through a word match to several different data source, DO (Disease Ontology), CTD Medic disease ontology vocabulary, HPO (Human Phenotype Ontology), OMIM synonym, OMIM descriptors, and Phenolyzer’s compiled disease vocabulary. After the first match, the disease names are directly returned for Phenolyzer’s compiled disease names and OMIM synonyms. For DO and CTD, an ontology search will retrieve all the descendent disease names and synonyms. For OMIM descriptors, they are mapped into OMIM diseases with a conditional probability as reliability. For HPO, an ontology search first finds all the descendent phenotypes, then the phenotypes are mapped into diseases with reliabilities.
Supplementary Figure 8 Learning curve through the training with Gradient Descent Algorithm.
The gradient descent algorithm iteratively reduces the cost and changes weight vector w into the direction opposite to the gradient of the cost function. After 10,000 iterations, with learning rate at 1, the slope of the curve is close to 0 and demonstrates that the number of learning steps is sufficient.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1 and 2, and Supplementary Note 1 (PDF 2236 kb)
Supplementary Data 1
The result dataset of causal genes for the 14 monogenic diseases. (XLSX 18 kb)
Supplementary Data 2
The result dataset of the 590 monogenic disease genes. (XLSX 28 kb)
Supplementary Data 3
The result dataset of candidate genes for the four complex diseases (XLSX 9004 kb)
Supplementary Data 4
The result dataset of novel discovered genes from four high-profile human genetics journals. (XLSX 45 kb)
Supplementary Data 5
The original prioritized gene lists for ‘craniopharyngiomas’ (XLSX 18 kb)
Supplementary Data 6
The original prioritized gene lists for ‘SHORT syndrome’ (XLSX 10 kb)
Supplementary Data 7
The original prioritized gene lists for ‘osteoporosis’ (XLSX 46 kb)
Supplementary Data 8
The full disease names and HPO identifiers for the term ‘autism’ (XLSX 19 kb)
Supplementary Software
Phenolyzer Software (ZIP 211361 kb)
Rights and permissions
About this article
Cite this article
Yang, H., Robinson, P. & Wang, K. Phenolyzer: phenotype-based prioritization of candidate genes for human diseases. Nat Methods 12, 841–843 (2015). https://doi.org/10.1038/nmeth.3484
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3484
This article is cited by
-
Using multi-scale genomics to associate poorly annotated genes with rare diseases
Genome Medicine (2024)
-
Analysis of the relationship of refractory gout between potential biomarkers and diet structure and lifestyle based on 1H-NMR
Journal of Orthopaedic Surgery and Research (2024)
-
Pan-caner analysis identifies PSMA7 as a targets for amplification at 20q13.33 in tumorigenesis
Scientific Reports (2024)
-
Novel lissencephaly-associated NDEL1 variant reveals distinct roles of NDE1 and NDEL1 in nucleokinesis and human cortical malformations
Acta Neuropathologica (2024)
-
Pharmacovariome scanning using whole pharmacogene resequencing coupled with deep computational analysis and machine learning for clinical pharmacogenomics
Human Genomics (2023)