Abstract
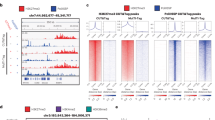
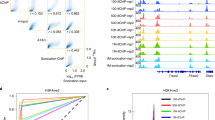
Chromatin immunoprecipitation assays have contributed greatly to our understanding of the role of histone modifications in gene regulation. However, they do not permit analysis with single-cell resolution, thus confounding analyses of heterogeneous cell populations. Here we present a method that permits visualization of histone modifications of single genomic loci with single-cell resolution in formaldehyde-fixed paraffin-embedded tissue sections based on combined use of in situ hybridization and proximity ligation assays. We show that dimethylation of lysine 4 of histone H3 (H3K4me2) at the MYH11 locus is restricted to the smooth muscle cell (SMC) lineage in human and mouse tissue sections and that the mark persists even in phenotypically modulated SMC in atherosclerotic lesions that show no detectable expression of SMC marker genes. This methodology has promise for broad applications in the study of epigenetic mechanisms in complex multicellular tissues in development and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Law, J.A. & Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Azuara, V. et al. Chromatin signatures of pluripotent cell lines. Nat. Cell Biol. 8, 532–538 (2006).
Bernstein, B.E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Cedar, H. & Bergman, Y. Epigenetics of haematopoietic cell development. Nat. Rev. Immunol. 11, 478–488 (2011).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Litt, M.D., Simpson, M., Gaszner, M., Allis, C.D. & Felsenfeld, G. Correlation between histone lysine methylation and developmental changes at the chicken β-globin locus. Science 293, 2453–2455 (2001).
Shechter, D. et al. Analysis of histones in Xenopus laevis. I. A distinct index of enriched variants and modifications exists in each cell type and is remodeled during developmental transitions. J. Biol. Chem. 284, 1064–1074 (2009).
Manabe, I. & Owens, G.K. Recruitment of serum response factor and hyperacetylation of histones at smooth muscle-specific regulatory regions during differentiation of a novel P19-derived in vitro smooth muscle differentiation system. Circ. Res. 88, 1127–1134 (2001).
McDonald, O.G., Wamhoff, B.R., Hoofnagle, M.H. & Owens, G.K. Control of SRF binding to CArG box chromatin regulates smooth muscle gene expression in vivo. J. Clin. Invest. 116, 36–48 (2006).
Miano, J.M., Cserjesi, P., Ligon, K.L., Periasamy, M. & Olson, E.N. Smooth muscle myosin heavy chain exclusively marks the smooth muscle lineage during mouse embryogenesis. Circ. Res. 75, 803–812 (1994).
Salmon, M., Gomez, D., Greene, E., Shankman, L. & Owens, G.K. Cooperative binding of KLF4, pELK-1, and HDAC2 to a G/C repressor element in the SM22α promoter mediates transcriptional silencing during SMC phenotypic switching in vivo. Circ. Res. 111, 685–696 (2012).
Owens, G.K., Kumar, M.S. & Wamhoff, B.R. Molecular regulation of vascular smooth muscle cell differentiation in development and disease. Physiol. Rev. 84, 767–801 (2004).
Alexander, M.R. & Owens, G.K. Epigenetic control of smooth muscle cell differentiation and phenotypic switching in vascular development and disease. Annu. Rev. Physiol. 74, 13–40 (2012).
Dahl, J.A. & Collas, P. A rapid micro chromatin immunoprecipitation assay (ChIP). Nat. Protoc. 3, 1032–1045 (2008).
Roh, T.Y., Ngau, W.C., Cui, K., Landsman, D. & Zhao, K. High-resolution genome-wide mapping of histone modifications. Nat. Biotechnol. 22, 1013–1016 (2004).
Söderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Lievens, S. & Tavernier, J. Single protein complex visualization: seeing is believing. Nat. Methods 3, 971–972 (2006).
Rantala, J.K. et al. SHARPIN is an endogenous inhibitor of ß1-integrin activation. Nat. Cell Biol. 13, 1315–1324 (2011).
Brobeil, A. et al. PTPIP51 is phosphorylated by Lyn and c-Src kinases lacking dephosphorylation by PTP1B in acute myeloid leukemia. Leuk. Res. 35, 1367–1375 (2011).
Wirth, A. et al. G12-G13–LARG–mediated signaling in vascular smooth muscle is required for salt-induced hypertension. Nat. Med. 14, 64–68 (2008).
Lampugnani, M.G. et al. A novel endothelial-specific membrane protein is a marker of cell-cell contacts. J. Cell Biol. 118, 1511–1522 (1992).
Li, L., Miano, J.M., Mercer, B. & Olson, E.N. Expression of the SM22α promoter in transgenic mice provides evidence for distinct transcriptional regulatory programs in vascular and visceral smooth muscle cells. J. Cell Biol. 132, 849–859 (1996).
Regan, C.P., Adam, P.J., Madsen, C.S. & Owens, G.K. Molecular mechanisms of decreased smooth muscle differentiation marker expression after vascular injury. J. Clin. Invest. 106, 1139–1147 (2000).
Carmeliet, P. Mechanisms of angiogenesis and arteriogenesis. Nat. Med. 6, 389–395 (2000).
Hanahan, D. Signaling vascular morphogenesis and maintenance. Science 277, 48–50 (1997).
Gomez, D. & Owens, G.K. Smooth muscle cell phenotypic switching in atherosclerosis. Cardiovasc. Res. 95, 156–164 (2012).
Weibrecht, I. et al. Visualising individual sequence-specific protein-DNA interactions in situ. N. Biotechnol. 29, 589–598 (2012).
Gustafsdottir, S.M. et al. In vitro analysis of DNA-protein interactions by proximity ligation. Proc. Natl. Acad. Sci. USA 104, 3067–3072 (2007).
You, J.S. & Jones, P.A. Cancer genetics and epigenetics: two sides of the same coin? Cancer Cell 22, 9–20 (2012).
Cui, K. et al. Chromatin signatures in multipotent human hematopoietic stem cells indicate the fate of bivalent genes during differentiation. Cell Stem Cell 4, 80–93 (2009).
Nguyen, A.T., Taranova, O., He, J. & Zhang, Y. DOT1L, the H3K79 methyltransferase, is required for MLL-AF9-mediated leukemogenesis. Blood 117, 6912–6922 (2011).
Ng, R.K. & Gurdon, J.B. Epigenetic inheritance of cell differentiation status. Cell Cycle 7, 1173–1177 (2008).
Ng, R.K. & Gurdon, J.B. Epigenetic memory of an active gene state depends on histone H3.3 incorporation into chromatin in the absence of transcription. Nat. Cell Biol. 10, 102–109 (2008).
Christova, R. & Oelgeschlager, T. Association of human TFIID-promoter complexes with silenced mitotic chromatin in vivo. Nat. Cell Biol. 4, 79–82 (2002).
Caplice, N.M. et al. Smooth muscle cells in human coronary atherosclerosis can originate from cells administered at marrow transplantation. Proc. Natl. Acad. Sci. USA 100, 4754–4759 (2003).
Acknowledgements
We thank M.E. McCanna and R.S. Tripathi for their knowledge and technical expertise, J.W. Mandell (University of Virginia) for providing human brain sections and S. Offermanns (Max Planck Institute) for Myh11-CreERT2 mice. This work was supported by US National Institutes of Health grants R01 HL57353, R01 HL098538 and R01 HL087867 (to G.K.O.). D.G. is supported by the American Heart Association Postdoctoral Fellowship 11POST7760009. L.S.S. is funded by a predoctoral American Heart Association Fellowship 11PRE17008. A.T.N. is funded by a postdoctoral American Heart Association Fellowship 12POST11630032.
Author information
Authors and Affiliations
Contributions
G.K.O. supervised this study; D.G. and G.K.O. conceived of the ISH-PLA strategies, designed studies and wrote the paper; D.G. generated labeled DNA probes, performed immunostaining and all ISH-PLA experiments and analyzed data; D.G. performed in vitro experiments, ChIP and quantitative PCR; L.S.S. generated Myh11-CreERT2 ROSA STOP-flox EYFP+/+ mice; L.S.S. and A.T.N. performed immunostaining on mouse sections; and D.G., L.S.S. and A.T.N. performed image acquisition and analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Table 1 (PDF 2255 kb)
Rights and permissions
About this article
Cite this article
Gomez, D., Shankman, L., Nguyen, A. et al. Detection of histone modifications at specific gene loci in single cells in histological sections. Nat Methods 10, 171–177 (2013). https://doi.org/10.1038/nmeth.2332
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2332
This article is cited by
-
Epigenetic regulation in metabolic diseases: mechanisms and advances in clinical study
Signal Transduction and Targeted Therapy (2023)
-
Cellular microarrays for assessing single-cell phenotypic changes in vascular cell populations
Biomedical Microdevices (2023)
-
How vascular smooth muscle cell phenotype switching contributes to vascular disease
Cell Communication and Signaling (2022)
-
Lipid phosphate phosphatase 3 in smooth muscle cells regulates angiotensin II-induced abdominal aortic aneurysm formation
Scientific Reports (2022)
-
An update on the phenotypic switching of vascular smooth muscle cells in the pathogenesis of atherosclerosis
Cellular and Molecular Life Sciences (2022)