Abstract
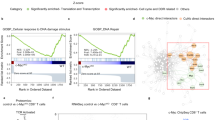
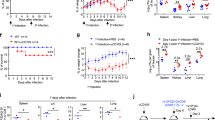
We found upregulation of expression of the microRNA miR-155 in primary effector and effector memory CD8+ T cells, but low miR-155 expression in naive and central memory cells. Antiviral CD8+ T cell responses and viral clearance were impaired in miR-155-deficient mice, and this defect was intrinsic to CD8+ T cells, as miR-155-deficient CD8+ T cells mounted greatly diminished primary and memory responses. Conversely, miR-155 overexpression augmented antiviral CD8+ T cell responses in vivo. Gene-expression profiling showed that miR-155-deficient CD8+ T cells had enhanced type I interferon signaling and were more susceptible to interferon's antiproliferative effect. Inhibition of the type I interferon–associated transcription factors STAT1 or IRF7 resulted in enhanced responses of miR-155-deficient CD8+ T cells in vivo. We have thus identified a previously unknown role for miR-155 in regulating responsiveness to interferon and CD8+ T cell responses to pathogens in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Lodish, H.F., Zhou, B., Liu, G. & Chen, C.Z. Micromanagement of the immune system by microRNAs. Nat. Rev. Immunol. 8, 120–130 (2008).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Miranda, K.C. et al. A pattern-based method for the identification of microRNA binding sites and their corresponding heteroduplexes. Cell 126, 1203–1217 (2006).
Friedman, R.C., Farh, K.K., Burge, C.B. & Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Lee, Y., Jeon, K., Lee, J.T., Kim, S. & Kim, V.N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, 4663–4670 (2002).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Lund, E., Guttinger, S., Calado, A., Dahlberg, J.E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Muljo, S.A. et al. Aberrant T cell differentiation in the absence of Dicer. J. Exp. Med. 202, 261–269 (2005).
Zhang, N. & Bevan, M.J. Dicer controls CD8+ T-cell activation, migration, and survival. Proc. Natl. Acad. Sci. USA 107, 21629–21634 (2010).
Clurman, B.E. & Hayward, W.S. Multiple proto-oncogene activations in avian leukosis virus-induced lymphomas: evidence for stage-specific events. Mol. Cell Biol. 9, 2657–2664 (1989).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611 (2007).
Vigorito, E. et al. microRNA-155 regulates the generation of immunoglobulin class-switched plasma cells. Immunity 27, 847–859 (2007).
Thai, T.H. et al. Regulation of the germinal center response by microRNA-155. Science 316, 604–608 (2007).
Costinean, S. et al. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in E(mu)-miR155 transgenic mice. Proc. Natl. Acad. Sci. USA 103, 7024–7029 (2006).
Tili, E. et al. Modulation of miR-155 and miR-125b levels following lipopolysaccharide/TNF-α stimulation and their possible roles in regulating the response to endotoxin shock. J. Immunol. 179, 5082–5089 (2007).
O'Connell, R.M. et al. MicroRNA-155 promotes autoimmune inflammation by enhancing inflammatory T cell development. Immunity 33, 607–619 (2010).
Kohlhaas, S. et al. Cutting edge: The Foxp3 target miR-155 contributes to the development of regulatory T cells. J. Immunol. 182, 2578–2582 (2009).
Lu, L.F. et al. Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity 30, 80–91 (2009).
Tsai, C.Y., Allie, S.R., Zhang, W. & Usherwood, E.J. MicroRNA miR-155 affects antiviral effector and effector memory CD8 T cell differentiation. J. Virol. 87, 2348–2351 (2013).
Lind, E.F., Elford, A.R. & Ohashi, P.S. Micro-RNA 155 Is required for optimal CD8+ T cell responses to acute viral and intracellular bacterial challenges. J. Immunol. 190, 1210–1216 (2013).
Kolumam, G.A., Thomas, S., Thompson, L.J., Sprent, J. & Murali-Krishna, K. Type I interferons act directly on CD8 T cells to allow clonal expansion and memory formation in response to viral infection. J. Exp. Med. 202, 637–650 (2005).
Curtsinger, J.M. et al. IFNs provide a third signal to CD8 T cells to stimulate clonal expansion and differentiation. J. Immunol. 174, 4465–4469 (2005).
Gil, M.P., Salomon, R., Louten, J. & Biron, C.A. Modulation of STAT1 protein levels: a mechanism shaping CD8 T-cell responses in vivo. Blood 107, 987–993 (2006).
Marshall, H.D., Urban, S.L. & Welsh, R.M. Virus-induced transient immune suppression and the inhibition of T cell proliferation by type I interferon. J. Virol. 85, 5929–5939 (2011).
McNally, J.M. et al. Attrition of bystander CD8 T cells during virus-induced T-cell and interferon responses. J. Virol. 75, 5965–5976 (2001).
Ebert, P.J., Jiang, S., Xie, J., Li, Q.J. & Davis, M.M. An endogenous positively selecting peptide enhances mature T cell responses and becomes an autoantigen in the absence of microRNA miR-181a. Nat. Immunol. 10, 1162–1169 (2009).
Li, Q.J. et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 129, 147–161 (2007).
Trotta, R. et al. miR-155 regulates IFN-γ production in natural killer cells. Blood 119, 3478–3485 (2012).
Agarwal, P. et al. Gene regulation and chromatin remodeling by IL-12 and type I IFN in programming for CD8 T cell effector function and memory. J. Immunol. 183, 1695–1704 (2009).
Sana, T.R., Janatpour, M.J., Sathe, M., McEvoy, L.M. & McClanahan, T.K. Microarray analysis of primary endothelial cells challenged with different inflammatory and immune cytokines. Cytokine 29, 256–269 (2005).
Tsuchihashi, S., Zhai, Y., Fondevila, C., Busuttil, R.W. & Kupiec-Weglinski, J.W. HO-1 upregulation suppresses type 1 IFN pathway in hepatic ischemia/reperfusion injury. Transplant. Proc. 37, 1677–1678 (2005).
Kashiwada, M. et al. Downstream of tyrosine kinases-1 and Src homology 2-containing inositol 5′-phosphatase are required for regulation of CD4+CD25+ T cell development. J. Immunol. 176, 3958–3965 (2006).
O'Connell, R.M., Chaudhuri, A.A., Rao, D.S. & Baltimore, D. Inositol phosphatase SHIP1 is a primary target of miR-155. Proc. Natl. Acad. Sci. USA 106, 7113–7118 (2009).
Cocolakis, E. et al. Smad signaling antagonizes STAT5-mediated gene transcription and mammary epithelial cell differentiation. J. Biol. Chem. 283, 1293–1307 (2008).
Erickson, S. et al. Interferon-α inhibits Stat5 DNA-binding in IL-2 stimulated primary T-lymphocytes. Eur. J. Biochem. 269, 29–37 (2002).
Haasch, D. et al. T cell activation induces a noncoding RNA transcript sensitive to inhibition by immunosuppressant drugs and encoded by the proto-oncogene, BIC. Cell Immunol. 217, 78–86 (2002).
Dondi, E., Rogge, L., Lutfalla, G., Uze, G. & Pellegrini, S. Down-modulation of responses to type I IFN upon T cell activation. J. Immunol. 170, 749–756 (2003).
Erickson, S. et al. Interferon-α inhibits proliferation in human T lymphocytes by abrogation of interleukin 2-induced changes in cell cycle-regulatory proteins. Cell Growth Differ. 10, 575–582 (1999).
Gimeno, R., Lee, C.K., Schindler, C. & Levy, D.E. Stat1 and Stat2 but not Stat3 arbitrate contradictory growth signals elicited by α/β interferon in T lymphocytes. Mol. Cell Biol. 25, 5456–5465 (2005).
Tanabe, Y. et al. Cutting edge: role of STAT1, STAT3, and STAT5 in IFN-αβ responses in T lymphocytes. J. Immunol. 174, 609–613 (2005).
Linsley, P.S. et al. Transcripts targeted by the microRNA-16 family cooperatively regulate cell cycle progression. Mol. Cell Biol. 27, 2240–2252 (2007).
Tsang, J., Zhu, J. & van Oudenaarden, A. MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Mol. Cell 26, 753–767 (2007).
Wherry, E.J. et al. Lineage relationship and protective immunity of memory CD8 T cell subsets. Nat. Immunol. 4, 225–234 (2003).
Usherwood, E.J. et al. Immunological control of murine gammaherpesvirus infection is independent of perforin. J. Gen. Virol. 78, 2025–2030 (1997).
Borowski, A.B. et al. Memory CD8+ T cells require CD28 costimulation. J. Immunol. 179, 6494–6503 (2007).
Dolfi, D.V. et al. Dendritic cells and CD28 costimulation are required to sustain virus-specific CD8+ T cell responses during the effector phase in vivo. J. Immunol. 186, 4599–4608 (2011).
Shen, L., Evel-Kabler, K., Strube, R. & Chen, S.Y. Silencing of SOCS1 enhances antigen presentation by dendritic cells and antigen-specific anti-tumor immunity. Nat. Biotechnol. 22, 1546–1553 (2004).
Abate, A., Zhao, H., Wong, R.J. & Stevenson, D.K. The role of Bach1 in the induction of heme oxygenase by tin mesoporphyrin. Biochem. Biophys. Res. Commun. 354, 757–763 (2007).
Lu, Y. et al. Loss of SOCS3 gene expression converts STAT3 function from anti-apoptotic to pro-apoptotic. J. Biol. Chem. 281, 36683–36690 (2006).
Lin, R., Mamane, Y. & Hiscott, J. Multiple regulatory domains control IRF-7 activity in response to virus infection. J. Biol. Chem. 275, 34320–34327 (2000).
Irizarry, R.A. et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 31, e15 (2003).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank W. Gerhard (Wistar Institute) for influenza virus strain A/Puerto Rico/8/34; D. Topham (University of Rochester) for influenza virus strain A/WSN/33 expressing OVA(257–264); H. Shen (University of Pennsylvania) for OVA-expressing L. monocytogenes; E. Vigorito (Babraham Institute) for control and miR-155-expressing MigR1 vectors; G. Takaesu (Keio University) for STAT1YF-IRES-GFP-pMX (DN-STAT1) and control retroviruses; B. tenOever (Mount Sinai School of Medicine) for DN-IRF7 plasmid; P. Marack (University of Colorado Health Sciences Center) for the MSCV-IRES-Thy-1.1 vector; and the Penn Molecular Profiling facility at the University of Pennsylvania for microarray assays. Some of this work was presented at the 96th Annual Meeting of the American Association of Immunologists in 2009. Supported by the US National Institutes of Health (U19 AI83022 and U19 AI82630 to E.J.W.; and R01 AI66215 and R01 AI46719), the Department of Microbiology and Immunology (P.D.K.) and the Biotechnology and Biological Sciences Research Council and Medical Research Council (M.T.)
Author information
Authors and Affiliations
Contributions
D.T.G., infection with influenza virus, adoptive transfer, in vitro proliferation, immunoblot analysis, siRNA transfection, retroviral transduction and RT-PCR; E.S., infection with L. monocytogenes, adoptive transfer, in vitro proliferation, siRNA transfection and RT-PCR; J.L.H., A.C.B., J.A.F. and J.N., infection with influenza virus, flow cytometry, BrdU assays, RT-PCR and mouse breeding; T.A.D., E.S. and E.J.W., microarray data analysis; Y.M.M., adoptive transfer and data analysis; E.S., D.T.G., A.C.B., E.J.W., M.T. and P.D.K., study design, data analysis and manuscript authorship; and all authors, discussion of results and comments on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Table 1 (PDF 599 kb)
Rights and permissions
About this article
Cite this article
Gracias, D., Stelekati, E., Hope, J. et al. The microRNA miR-155 controls CD8+ T cell responses by regulating interferon signaling. Nat Immunol 14, 593–602 (2013). https://doi.org/10.1038/ni.2576
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2576
This article is cited by
-
Non-coding RNAs in disease: from mechanisms to therapeutics
Nature Reviews Genetics (2024)
-
The dietary sweetener sucralose is a negative modulator of T cell-mediated responses
Nature (2023)
-
Roles of the miR-155 in Neuroinflammation and Neurological Disorders: A Potent Biological and Therapeutic Target
Cellular and Molecular Neurobiology (2023)
-
Systemic Listeria monocytogenes infection in aged mice induces long-term neuroinflammation: the role of miR-155
Immunity & Ageing (2022)
-
Role of miR-155 in inflammatory autoimmune diseases: a comprehensive review
Inflammation Research (2022)