Abstract
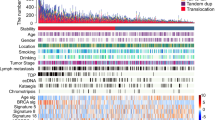
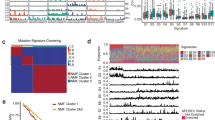
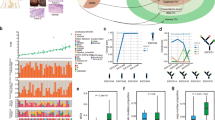
Esophageal squamous cell carcinoma (ESCC) is among the most common malignancies, but little is known about its spatial intratumoral heterogeneity (ITH) and temporal clonal evolutionary processes. To address this, we performed multiregion whole-exome sequencing on 51 tumor regions from 13 ESCC cases and multiregion global methylation profiling for 3 of these 13 cases. We found an average of 35.8% heterogeneous somatic mutations with strong evidence of ITH. Half of the driver mutations located on the branches of tumor phylogenetic trees targeted oncogenes, including PIK3CA, NFE2L2 and MTOR, among others. By contrast, the majority of truncal and clonal driver mutations occurred in tumor-suppressor genes, including TP53, KMT2D and ZNF750, among others. Interestingly, phyloepigenetic trees robustly recapitulated the topological structures of the phylogenetic trees, indicating a possible relationship between genetic and epigenetic alterations. Our integrated investigations of spatial ITH and clonal evolution provide an important molecular foundation for enhanced understanding of tumorigenesis and progression in ESCC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Torre, L.A. et al. Global cancer statistics, 2012. CA Cancer J. Clin. 65, 87–108 (2015).
Ferlay, J. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136, E359–E386 (2015).
Enzinger, P.C. & Mayer, R.J. Esophageal cancer. N. Engl. J. Med. 349, 2241–2252 (2003).
Agrawal, N. et al. Comparative genomic analysis of esophageal adenocarcinoma and squamous cell carcinoma. Cancer Discov. 2, 899–905 (2012).
Song, Y. et al. Identification of genomic alterations in oesophageal squamous cell cancer. Nature 509, 91–95 (2014).
Lin, D.C. et al. Genomic and molecular characterization of esophageal squamous cell carcinoma. Nat. Genet. 46, 467–473 (2014).
Gao, Y.B. et al. Genetic landscape of esophageal squamous cell carcinoma. Nat. Genet. 46, 1097–1102 (2014).
Zhang, L. et al. Genomic analyses reveal mutational signatures and frequently altered genes in esophageal squamous cell carcinoma. Am. J. Hum. Genet. 96, 597–611 (2015).
Cheng, C. et al. Whole-genome sequencing reveals diverse models of structural variations in esophageal squamous cell carcinoma. Am. J. Hum. Genet. 98, 256–274 (2016).
McGranahan, N. & Swanton, C. Biological and therapeutic impact of intratumor heterogeneity in cancer evolution. Cancer Cell 27, 15–26 (2015).
Futreal, P.A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Lawrence, M.S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci. Transl. Med. 7, 283ra54 (2015).
Durinck, S. et al. Temporal dissection of tumorigenesis in primary cancers. Cancer Discov. 1, 137–143 (2011).
Shah, S.P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012).
Lohr, J.G. et al. Widespread genetic heterogeneity in multiple myeloma: implications for targeted therapy. Cancer Cell 25, 91–101 (2014).
Landau, D.A. et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 152, 714–726 (2013).
Shi, Z.Z. et al. Consistent and differential genetic aberrations between esophageal dysplasia and squamous cell carcinoma detected by array comparative genomic hybridization. Clin. Cancer Res. 19, 5867–5878 (2013).
Shang, L. et al. ANO1 protein as a potential biomarker for esophageal cancer prognosis and precancerous lesion development prediction. Oncotarget 7, 24374–24382 (2016).
Britschgi, A. et al. Calcium-activated chloride channel ANO1 promotes breast cancer progression by activating EGFR and CAMK signaling. Proc. Natl. Acad. Sci. USA 110, E1026–E1034 (2013).
Luo, M.L. et al. Amplification and overexpression of CTTN (EMS1) contribute to the metastasis of esophageal squamous cell carcinoma by promoting cell migration and anoikis resistance. Cancer Res. 66, 11690–11699 (2006).
Lu, P. et al. Genome-wide gene expression profile analyses identify CTTN as a potential prognostic marker in esophageal cancer. PLoS One 9, e88918 (2014).
de Bruin, E.C. et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
Gerlinger, M. et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat. Genet. 46, 225–233 (2014).
Murugaesu, N. et al. Tracking the genomic evolution of esophageal adenocarcinoma through neoadjuvant chemotherapy. Cancer Discov. 5, 821–831 (2015).
Rosenthal, R., McGranahan, N., Herrero, J., Taylor, B.S. & Swanton, C. DeconstructSigs: delineating mutational processes in single tumors distinguishes DNA repair deficiencies and patterns of carcinoma evolution. Genome Biol. 17, 31 (2016).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Toh, Y., Egashira, A. & Yamamoto, M. Epigenetic alterations and their clinical implications in esophageal squamous cell carcinoma. Gen. Thorac. Cardiovasc. Surg. 61, 262–269 (2013).
Agarwal, R. et al. Epigenomic program of Barrett's-associated neoplastic progression reveals possible involvement of insulin signaling pathways. Endocr. Relat. Cancer 19, L5–L9 (2012).
Alvarez, H. et al. Widespread hypomethylation occurs early and synergizes with gene amplification during esophageal carcinogenesis. PLoS Genet. 7, e1001356 (2011).
Brocks, D. et al. Intratumor DNA methylation heterogeneity reflects clonal evolution in aggressive prostate cancer. Cell Rep. 8, 798–806 (2014).
Mazor, T. et al. DNA methylation and somatic mutations converge on the cell cycle and define similar evolutionary histories in brain tumors. Cancer Cell 28, 307–317 (2015).
Robinson, D.F. & Foulds, L.R. Comparison of phylogenetic trees. Math. Biosci. 53, 131–147 (1981).
Oricchio, E. et al. The Eph-receptor A7 is a soluble tumor suppressor for follicular lymphoma. Cell 147, 554–564 (2011).
López-Nieva, P. et al. EPHA7, a new target gene for 6q deletion in T-cell lymphoblastic lymphomas. Carcinogenesis 33, 452–458 (2012).
Yu, J. et al. Methylation of protocadherin 10, a novel tumor suppressor, is associated with poor prognosis in patients with gastric cancer. Gastroenterology 136, 640–651.e1 (2009).
Zhao, Y. et al. A novel Wnt regulatory axis in endometrioid endometrial cancer. Cancer Res. 74, 5103–5117 (2014).
Saulnier, A. et al. Inactivation of the putative suppressor gene DOK1 by promoter hypermethylation in primary human cancers. Int. J. Cancer 130, 2484–2494 (2012).
Mercier, P.L. et al. Characterization of DOK1, a candidate tumor suppressor gene, in epithelial ovarian cancer. Mol. Oncol. 5, 438–453 (2011).
Bergman, Y. & Cedar, H. DNA methylation dynamics in health and disease. Nat. Struct. Mol. Biol. 20, 274–281 (2013).
Quante, T. & Bird, A. Do short, frequent DNA sequence motifs mould the epigenome? Nat. Rev. Mol. Cell Biol. 17, 257–262 (2016).
Baylin, S.B. & Jones, P.A. A decade of exploring the cancer epigenome —biological and translational implications. Nat. Rev. Cancer 11, 726–734 (2011).
Lay, F.D. et al. The role of DNA methylation in directing the functional organization of the cancer epigenome. Genome Res. 25, 467–477 (2015).
Irizarry, R.A. et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat. Genet. 41, 178–186 (2009).
Doi, A. et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat. Genet. 41, 1350–1353 (2009).
Gharwan, H. & Groninger, H. Kinase inhibitors and monoclonal antibodies in oncology: clinical implications. Nat. Rev. Clin. Oncol. 13, 209–227 (2016).
Gerlinger, M. et al. Cancer: evolution within a lifetime. Annu. Rev. Genet. 48, 215–236 (2014).
Cao, W. et al. Multiple region whole-exome sequencing reveals dramatically evolving intratumor genomic heterogeneity in esophageal squamous cell carcinoma. Oncogenesis 4, e175 (2015).
Yap, T.A., Gerlinger, M., Futreal, P.A., Pusztai, L. & Swanton, C. Intratumor heterogeneity: seeing the wood for the trees. Sci. Transl. Med. 4, 127ps10 (2012).
Wang, J. et al. Downregulation of EphA7 by hypermethylation in colorectal cancer. Oncogene 24, 5637–5647 (2005).
Kiehl, S. et al. ABCB4 is frequently epigenetically silenced in human cancers and inhibits tumor growth. Sci. Rep. 4, 6899 (2014).
Jao, T.M. et al. Protocadherin 10 suppresses tumorigenesis and metastasis in colorectal cancer and its genetic loss predicts adverse prognosis. Int. J. Cancer 135, 2593–2603 (2014).
Narayan, G. et al. PCDH10 promoter hypermethylation is frequent in most histologic subtypes of mature lymphoid malignancies and occurs early in lymphomagenesis. Genes Chromosom. Cancer 52, 1030–1041 (2013).
Chen, W. et al. Cancer statistics in China, 2015. CA Cancer J. Clin. 66, 115–132 (2016).
Faust, G.G. & Hall, I.M. SAMBLASTER: fast duplicate marking and structural variant read extraction. Bioinformatics 30, 2503–2505 (2014).
Van der Auwera, G.A. et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
Koboldt, D.C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Stachler, M.D. et al. Paired exome analysis of Barrett's esophagus and adenocarcinoma. Nat. Genet. 47, 1047–1055 (2015).
Olshen, A.B., Venkatraman, E.S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Carter, S.L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Kumar, P., Henikoff, S. & Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Adzhubei, I., Jordan, D.M. & Sunyaev, S.R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. Chapter 7, Unit 7.20 (2013).
Triche, T.J. Jr., Weisenberger, D.J., Van Den Berg, D., Laird, P.W. & Siegmund, K.D. Low-level processing of Illumina Infinium DNA Methylation BeadArrays. Nucleic Acids Res. 41, e90 (2013).
Desper, R. & Gascuel, O. Fast and accurate phylogeny reconstruction algorithms based on the minimum-evolution principle. J. Comput. Biol. 9, 687–705 (2002).
Takahashi, T. et al. Estimation of the fraction of cancer cells in a tumor DNA sample using DNA methylation. PLoS One 8, e82302 (2013).
Zack, T.I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Pagès, F. et al. In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. J. Clin. Oncol. 27, 5944–5951 (2009).
de Miranda, N.F. et al. Infiltration of Lynch colorectal cancers by activated immune cells associates with early staging of the primary tumor and absence of lymph node metastases. Clin. Cancer Res. 18, 1237–1245 (2012).
Punt, S. et al. Whole-transcriptome analysis of flow-sorted cervical cancer samples reveals that B cell expressed TCL1A is correlated with improved survival. Oncotarget 6, 38681–38694 (2015).
Gorter, A., Prins, F., van Diepen, M., Punt, S. & van der Burg, S.H. The tumor area occupied by Tbet+ cells in deeply invading cervical cancer predicts clinical outcome. J. Transl. Med. 13, 295 (2015).
Reinius, L.E. et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PLoS One 7, e41361 (2012).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Ceccarelli, M. et al. Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164, 550–563 (2016).
Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Wu, H., Caffo, B., Jaffee, H.A., Irizarry, R.A. & Feinberg, A.P. Redefining CpG islands using hidden Markov models. Biostatistics 11, 499–514 (2010).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Song, Q. et al. A reference methylome database and analysis pipeline to facilitate integrative and comparative epigenomics. PLoS One 8, e81148 (2013).
Acknowledgements
We thank H. Shen and D. Weisenberger as well as A.D. Jeyasekharan for their kind help on analysis and discussion. This work was funded by the Singapore Ministry of Health's National Medical Research Council (NMRC) through its Singapore Translational Research (STaR) Investigator Award to H.P.K., an NMRC Individual Research Grant (NMRC/1311/2011) and the NMRC Centre Grant awarded to the National University Cancer Institute of Singapore, the National Research Foundation Singapore and the Singapore Ministry of Education under its Research Centres of Excellence initiatives to H.P.K. D.-C.L. was supported by the American Society of Hematology Fellow Scholar Award, the National Natural Science Foundation of China (81672786) and National Center for Advancing Translational Sciences UCLA CTSI Grant UL1TR000124. M.-R.W. was supported by the National Natural Science Foundation of China (81330052, 81520108023 and 81321091). Y.Z. was supported by the Beijing Natural Science Foundation (7151008). This study was partially supported by a generous donation from the Melamed family and NIH/NCI grant 1U01CA184826 as well as institutional support from the Samuel Oschin Comprehensive Cancer Institute to B.P.B. and H.Q.D.
Author information
Authors and Affiliations
Contributions
M.-R.W., D.-C.L., B.P.B. and H.P.K. conceived and designed the experiments. J.-J.H., D.-C.L., H.Q.D., W.-Q.W., B.P.B., M.-R.W. and H.P.K. wrote the manuscript. J.-J.H., D.-C.L., Y.J., C.C., C.-C.L., X.X. and Y.C. performed the experiments. J.-J.H., H.Q.D., A.M., B.P.B. and Z.-Z.S. performed statistical analysis. J.-J.H., D.-C.L., H.Q.D., Y.-Y.J., B.P.B. and H.P.K. analyzed the data. X.X. contributed reagents. W.-Q.W. contributed materials. J.-W.W. and J.-J.H. read slides with hematoxylin and eosin staining. D.-C.L., Y.Z., Q.-M.Z. and H.P.K. jointly supervised research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Fitting somatic variants to evolutionary models based on phylogenetic trees.
Mutations that were mapped to the shared and private branches of trees and were 100% compatible with tree structure are shown on the branch where they occurred; those incompatible with tree structure are not shown. As trunk variants were defined by their presence in all regions, they were by definition consistent and are not shown because of space limitations. The total number of mutations in each case and the percentage that were compatible with the evolutionary tree model are provided above each tree (in parentheses).
Supplementary Figure 2 Correlation of early and late somatic mutations with potential targeting approaches.
Candidate potential targeting approaches, which have been developed or are under evaluation, were selected on the basis of The Cancer Genome Analysis (TCGA; http://www.broadinstitute.org/cancer/cga/target).
Supplementary Figure 3 ITH of CNAs in ESCC.
Heat maps displaying recurrent putative driver CNAs for each tumor region (driver CNAs were taken from our previous results; Nat. Genet. 46, 467–473, 2014). Chromosome segments with a log2 ratio between 0.5 and 1.0 were classified as gains, greater than 1.0 were classified as amplifications, less than –0.5 were classified as losses and less than –1.0 were classified as deletions.
Supplementary Figure 4 The number of clonal and subclonal mutations in ESCC cases.
Clonal/subclonal status is described in the Online Methods.
Supplementary Figure 6 Reconstruction of phyloepigenetic trees with four different probe selection cutoffs (0.2, 0.3, 0.4 and 0.5).
The probe selection cutoff defines the minimum delta methylation level (β value) used for variable selection (‘shared’ versus ‘private’ status) to choose the methylation probes to be used in construction of trees. Each row shows one case, and each column shows the resulting tree for each cutoff. The cutoff used in the results section (Fig. 4a) was 0.3. The number of probes selected (n) and Robinson-Foulds (RF) distance to the original tree (cutoff = 0.3) are provided for each tree (except for the one with cutoff = 0.3, which is the reference). ESCC05 did not have enough probes to construct a tree at cutoff = 0.5.
Supplementary Figure 7 Immune cells make up most of the non-cancer cell fraction in ESCC samples.
Representative IHC photos of the four spatially distinct regions for case ESCC05, showing that immune cells (LCA/CD45 positive staining; highlighted within the yellow squares) make up most of the non-cancer cells in this tumor. Cancer cells are shown in the red squares). Scale bar, 200 μm.
Supplementary Figure 8 Reconstruction of the phyloepigenetic trees with mitigation for the effects of immune cell content.
We recalculated each phyloepigenetic tree using one of two methods to mitigate the effects of immune cells within the samples. For the first method, immune cell adjusted, we estimated the fraction of leukocytes in each sample using profiles of immune-specific methylation probes, as described previously (Nat. Biotechnol. 30, 413–421, 2012; Nat. Genet. 45, 1134–1140, 2013). We then used the methylation profile of pure leukocytes, along with the estimated leukocyte/cancer cell mixture, to infer the cancer cell methylation value for every probe (probe numbers are not identical because of different probes being below the detection limit in different experiments). For the second method, dichotomized, we used only probes that had a fully unmethylated state in pure leukocytes and dichotomized/binarized each probe on the basis of the minimum methylation level in tumor samples (Online Methods). For each of the immune cell adjusted and dichotomized trees, the RF distance shows similarity to the reference version.
Supplementary Figure 9 Unsupervised hierarchical clustering of methylation values at shared probes.
‘Shared’ probes were those selected as having similar values across different regions of the same tumor (the number of shared probes is given above each heat map as n). The shared probes were compared to normal adjacent tissue and divided into those that had consistently higher methylation values (hypermethylated) or lower methylation values (hypomethylated) in the tumor samples. The probes in each heat map were clustered using hierarchical clustering with the Euclidean distance metric.
Supplementary Figure 10 Analysis of tumor cell content in ESCC samples.
Representative hematoxylin and eosin photographs of case ESCC02. Scale bar, 2 mm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1, 5 and 7. (PDF 2039 kb)
Supplementary Table 2
Detailed information of all somatic mutations in 51 tumor regions from 13 patients with ESCC. (XLSX 1129 kb)
Supplementary Table 3
Validation by PCR and Sanger sequencing. (XLSX 13 kb)
Supplementary Table 4
Copy number of each chromosomal segment in 51 tumor regions from 13 patients with ESCC. (XLSX 994 kb)
Supplementary Table 6
Mutations incompatible with the phylogenetic tree. (XLSX 17 kb)
Rights and permissions
About this article
Cite this article
Hao, JJ., Lin, DC., Dinh, H. et al. Spatial intratumoral heterogeneity and temporal clonal evolution in esophageal squamous cell carcinoma. Nat Genet 48, 1500–1507 (2016). https://doi.org/10.1038/ng.3683
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3683
This article is cited by
-
Understanding PI3K/Akt/mTOR signaling in squamous cell carcinoma: mutated PIK3CA as an example
Molecular Biomedicine (2024)
-
Tracking the evolution of esophageal squamous cell carcinoma under dynamic immune selection by multi-omics sequencing
Nature Communications (2023)
-
Large-scale genomic analyses reveal alterations and mechanisms underlying clonal evolution and immune evasion in esophageal cancer
Nature Communications (2023)
-
CRISPR-based DNA methylation editing of NNT rescues the cisplatin resistance of lung cancer cells by reducing autophagy
Archives of Toxicology (2023)
-
RIG-I promotes cell proliferation in esophageal squamous cell carcinoma by facilitating p21 degradation
Medical Oncology (2023)