Abstract
Gliomas arising in the brainstem and thalamus are devastating tumors that are difficult to surgically resect. To determine the genetic and epigenetic landscape of these tumors, we performed exomic sequencing of 14 brainstem gliomas (BSGs) and 12 thalamic gliomas. We also performed targeted mutational analysis of an additional 24 such tumors and genome-wide methylation profiling of 45 gliomas. This study led to the discovery of tumor-specific mutations in PPM1D, encoding wild-type p53–induced protein phosphatase 1D (WIP1), in 37.5% of the BSGs that harbored hallmark H3F3A mutations encoding p.Lys27Met substitutions. PPM1D mutations were mutually exclusive with TP53 mutations in BSG and attenuated p53 activation in vitro. PPM1D mutations were truncating alterations in exon 6 that enhanced the ability of PPM1D to suppress the activation of the DNA damage response checkpoint protein CHK2. These results define PPM1D as a frequent target of somatic mutation and as a potential therapeutic target in brainstem gliomas.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Schwartzentruber, J. et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482, 226–231 (2012).
Khuong-Quang, D.A. et al. K27M mutation in histone H3.3 defines clinically and biologically distinct subgroups of pediatric diffuse intrinsic pontine gliomas. Acta Neuropathol. 124, 439–447 (2012).
Wu, G. et al. Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat. Genet. 44, 251–253 (2012).
Puget, S. et al. Mesenchymal transition and PDGFRA amplification/mutation are key distinct oncogenic events in pediatric diffuse intrinsic pontine gliomas. PLoS ONE 7, e30313 (2012).
Jones, D.T. et al. Recurrent somatic alterations of FGFR1 and NTRK2 in pilocytic astrocytoma. Nat. Genet. 45, 927–932 (2013).
Bulavin, D.V. et al. Amplification of PPM1D in human tumors abrogates p53 tumor-suppressor activity. Nat. Genet. 31, 210–215 (2002).
Ruark, E. et al. Mosaic PPM1D mutations are associated with predisposition to breast and ovarian cancer. Nature 493, 406–410 (2013).
Kleiblova, P. et al. Gain-of-function mutations of PPM1D/Wip1 impair the p53-dependent G1 checkpoint. J. Cell Biol. 201, 511–521 (2013).
Dudgeon, C. et al. Genetic variants and mutations of PPM1D control the response to DNA damage. Cell Cycle 12, 2656–2664 (2013).
Yan, H. et al. IDH1 and IDH2 mutations in gliomas. N. Engl. J. Med. 360, 765–773 (2009).
Turcan, S. et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483, 479–483 (2012).
Duncan, C.G. et al. A heterozygous IDH1R132H/WT mutation induces genome-wide alterations in DNA methylation. Genome Res. 22, 2339–2355 (2012).
Sturm, D. et al. Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22, 425–437 (2012).
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478 (2012).
Mootha, V.K. et al. PGC-1α–responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Oliva-Trastoy, M. et al. The Wip1 phosphatase (PPM1D) antagonizes activation of the Chk2 tumour suppressor kinase. Oncogene 26, 1449–1458 (2007).
Lu, X., Nannenga, B. & Donehower, L.A. PPM1D dephosphorylates Chk1 and p53 and abrogates cell cycle checkpoints. Genes Dev. 19, 1162–1174 (2005).
Yoda, A. et al. Intrinsic kinase activity and SQ/TQ domain of Chk2 kinase as well as N-terminal domain of Wip1 phosphatase are required for regulation of Chk2 by Wip1. J. Biol. Chem. 281, 24847–24862 (2006).
Moon, S.H., Nguyen, T.A., Darlington, Y., Lu, X. & Donehower, L.A. Dephosphorylation of γ-H2AX by WIP1: an important homeostatic regulatory event in DNA repair and cell cycle control. Cell Cycle 9, 2092–2096 (2010).
Yagi, H. et al. A small molecule inhibitor of p53-inducible protein phosphatase PPM1D. Bioorg. Med. Chem. Lett. 22, 729–732 (2012).
Gilmartin, A.G. et al. Allosteric Wip1 phosphatase inhibition through flap-subdomain interaction. Nat. Chem. Biol. 10, 181–187 (2014).
Jin, G. et al. Disruption of wild type IDH1 suppresses D-2-hydroxyglutarate production in IDH1-mutated gliomas. Cancer Res. 73, 496–501 (2013).
Verhaak, R.G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Parsons, D.W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812 (2008).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Monti, S., Tamayo, P., Mesirov, J. & Golub, T. Consensus clustering: a resampling-based method for class discovery and visualization of gene expression microarray data. Mach. Learn. 52, 91–118 (2003).
R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2012).
Irizarry, R.A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003).
Dabney, A.R. Classification of microarrays to nearest centroids. Bioinformatics 21, 4148–4154 (2005).
Kohli, M., Rago, C., Lengauer, C., Kinzler, K.W. & Vogelstein, B. Facile methods for generating human somatic cell gene knockouts using recombinant adeno-associated viruses. Nucleic Acids Res. 32, e3 (2004).
Acknowledgements
This study was partly supported by National Cancer Institute grant R01CA140316 (H.Y.), the American Cancer Society (RSG-10-126-01-CCE) (H.Y.), a Pediatric Brain Tumor Foundation Institute grant (D.D.B.), a Voices Against Brain Cancer Foundation grant (D.D.B.), a James S. McDonnell Foundation grant (H.Y.), The V Foundation (H.Y.), an Accelerate Brain Cancer Cure Foundation grant (H.Y.), a Pediatric Brain Tumor Foundation Institute at Duke Early Career Award (Z.J.R.) and Natural Science Foundation of China grants 30772237 and 81241078 (L.Z.).
Author information
Authors and Affiliations
Contributions
L.Z., H.Y., Z.J.R. and L.H.C. designed research. L.H.C., Z.J.R., J.B.W. and H.Y. wrote the manuscript. L.Z., L.H.C., H.W., W.Z., J.F., S.J., R.Y., P.J.K., J.Z., Z. Wu, S.H., Y.W. and J.B.W. performed the experiments. Z. Wang and H.Z. contributed to the biological validation of the functions of mutant PPM1D. H.S.F. and A.H.F. contributed to the inspection of clinical data. L.H.C., S.Y., S.J., S.W., G.L., R.E.M., Y.H., Z.J.R., D.D.B. and H.Y. analyzed the data.
Corresponding authors
Ethics declarations
Competing interests
Under agreements between Duke University, Agios Pharmaceuticals, Blueprint Medicines and Personal Genome Diagnostics, Inc., H.Y. and D.D.B. are entitled to a share of the royalties received by Duke University on the sales of products related to genes described in this manuscript. S.W. and H.Y. are co-founders and own stocks of Beijing Pangenomics Technology, Co. Ltd.
Integrated supplementary information
Supplementary Figure 1 Kaplan-Meier survival curves of patients with brainstem tumors.
(a) Comparison of patients with PPM1D mutation versus wild-type PPM1D. (b) Comparison of patients with PPM1D mutation versus TP53 mutation. (c) Comparison of patients with PPM1D/H3F3A mutations versus TP53/H3F3A mutations.
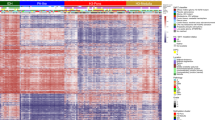
Supplementary Figure 2 Consensus clustering by methylation β value and Lorenz curve.
Our three groups could be mapped almost perfectly to IDH1 and H3F3A mutation status. We also noted that samples with hemizygous IDH1 mutation could be differentiated from samples with heterozygous IDH1 mutation.
Supplementary Figure 3 Principal-component analysis for our 45 samples.
Tumor samples with H3F3A mutation (HIST), IDH1 mutation (IDH) and neither mutation (WT) could be differentiated by DNA methylation levels.
Supplementary Figure 4 Box plot of DNA methylation levels for sample averages.
The group with IDH1 mutation shows a hypermethylated pattern, whereas the group with H3F3A mutation shows a hypomethylated pattern.
Supplementary Figure 5 Density plot of probes for DNA methylation levels (β values) in different groups.
The group with IDH1 mutation (red) shows a hypermethylated pattern, and the group with H3F3A (K27) mutation (orange) shows a hypomethylated pattern.
Supplementary Figure 6 Electropherograms of HCT116 parental, repaired 1 and repaired 2 lines.
Electropherograms demonstrate repair of the heterozygous frameshift mutation to a wild-type PPM1D sequence in these lines.
Supplementary Figure 7 Colony formation assay for the HCT116 parental, repaired 1 and repaired 2 lines.
Colony formation was assessed in the absence of ionizing radiation and under 2 Gy or 4 Gy of ionizing radiation. Representative stained plates with colony formation are shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1, 2, 5–7 and 9 (PDF 1543 kb)
Supplementary Tables 3, 4 and 8
Supplementary Tables 3, 4 and 8 (XLSX 108 kb)
Rights and permissions
About this article
Cite this article
Zhang, L., Chen, L., Wan, H. et al. Exome sequencing identifies somatic gain-of-function PPM1D mutations in brainstem gliomas. Nat Genet 46, 726–730 (2014). https://doi.org/10.1038/ng.2995
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2995
This article is cited by
-
PPM1D suppresses p53-dependent transactivation and cell death by inhibiting the Integrated Stress Response
Nature Communications (2022)
-
PPM1D mutations are oncogenic drivers of de novo diffuse midline glioma formation
Nature Communications (2022)
-
TP53 wild-type/PPM1D mutant diffuse intrinsic pontine gliomas are sensitive to a MDM2 antagonist
Acta Neuropathologica Communications (2021)
-
Inhibition of the DNA damage response phosphatase PPM1D reprograms neutrophils to enhance anti-tumor immune responses
Nature Communications (2021)
-
The transcriptional landscape of Shh medulloblastoma
Nature Communications (2021)