Abstract
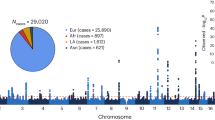
Schizophrenia is a severe mental disorder affecting ∼1% of the world population, with heritability of up to 80%. To identify new common genetic risk factors, we performed a genome-wide association study (GWAS) in the Han Chinese population. The discovery sample set consisted of 3,750 individuals with schizophrenia and 6,468 healthy controls (1,578 cases and 1,592 controls from northern Han Chinese, 1,238 cases and 2,856 controls from central Han Chinese, and 934 cases and 2,020 controls from the southern Han Chinese). We further analyzed the strongest association signals in an additional independent cohort of 4,383 cases and 4,539 controls from the Han Chinese population. Meta-analysis identified common SNPs that associated with schizophrenia with genome-wide significance on 8p12 (rs16887244, P = 1.27 × 10−10) and 1q24.2 (rs10489202, P = 9.50 × 10−9). Our findings provide new insights into the pathogenesis of schizophrenia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Craddock, N., O'donovan, M.C. & Owen, M.J. The genetics of schizophrenia and bipolar disorder: dissecting psychosis. J. Med. Genet. 42, 193–204 (2005).
McCarthy, S.E. et al. Microduplications of 16p11. 2 are associated with schizophrenia. Nat. Genet. 41, 1223–1227 (2009).
O'Donovan, M.C. et al. Identification of loci associated with schizophrenia by genome-wide association and follow-up. Nat. Genet. 40, 1053–1055 (2008).
Purcell, S.M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Shi, J. et al. Common variants on chromosome 6p22. 1 are associated with schizophrenia. Nature 460, 753–757 (2009).
Stefansson, H. et al. Common variants conferring risk of schizophrenia. Nature 460, 744–747 (2009).
Vacic, V. et al. Duplications of the neuropeptide receptor gene VIPR2 confer significant risk for schizophrenia. Nature 471, 499–503 (2011).
Xu, B. et al. Strong association of de novo copy number mutations with sporadic schizophrenia. Nat. Genet. 40, 880–885 (2008).
Stranger, B.E. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007).
Stranger, B.E. et al. Population genomics of human gene expression. Nat. Genet. 39, 1217–1224 (2007).
Gibbs, J.R. et al. Abundant quantitative trait loci exist for DNA methylation and gene expression in human brain. PLoS Genet. 6, e1000952 (2010).
Gaughran, F., Payne, J., Sedgwick, P.M., Cotter, D. & Berry, M. Hippocampal FGF-2 and FGFR1 mRNA expression in major depression, schizophrenia and bipolar disorder. Brain Res. Bull. 70, 221–227 (2006).
Klejbor, I. et al. Fibroblast growth factor receptor signaling affects development and function of dopamine neurons–inhibition results in a schizophrenia-like syndrome in transgenic mice. J. Neurochem. 97, 1243–1258 (2006).
Terwisscha van Scheltinga, A.F., Bakker, S.C. & Kahn, R.S. Fibroblast growth factors in schizophrenia. Schizophr. Bull. 36, 1157–1166 (2010).
O'Donovan, M.C. et al. Analysis of 10 independent samples provides evidence for association between schizophrenia and a SNP flanking fibroblast growth factor receptor 2. Mol. Psychiatry 14, 30–36 (2009).
Tkachev, D. et al. Oligodendrocyte dysfunction in schizophrenia and bipolar disorder. Lancet 362, 798–805 (2003).
He, G. et al. MPZL1/PZR, a novel candidate predisposing schizophrenia in Han Chinese. Mol. Psychiatry 11, 748–751 (2006).
Chen, J. et al. Genetic structure of the Han Chinese population revealed by genome-wide SNP variation. Am. J. Hum. Genet. 85, 775–785 (2009).
Li, T. et al. Common variants in major histocompatibility complex region and TCF4 gene are significantly associated with schizophrenia in Han Chinese. Biol. Psychiatry 68, 671–673 (2010).
Hindorff, L.A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 106, 9362–9367 (2009).
Korn, J.M. et al. Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat. Genet. 40, 1253–1260 (2008).
Chen, Z.J. et al. Genome-wide association study identifies susceptibility loci for polycystic ovary syndrome on chromosome 2p16.3, 2p21 and 9q33.3. Nat. Genet. 43, 55–59 (2011).
Thomas, G. et al. Capillary and microelectrophoretic separations of ligase detection reaction products produced from low-abundant point mutations in genomic DNA. Electrophoresis 25, 1668–1677 (2004).
Yi, P. et al. PCR/LDR/capillary electrophoresis for detection of single-nucleotide differences between fetal and maternal DNA in maternal plasma. Prenat. Diagn. 29, 217–222 (2009).
Patterson, N., Price, A.L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Barrett, J.C., Fry, B., Maller, J. & Daly, M.J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Shi, Y.Y. & He, L. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res. 15, 97–98 (2005).
Mantel, N. & Haenszel, W. Statistical aspects of the analysis of data from retrospective studies. J. Natl. Cancer Inst. 22, 719–748 (1959).
Petukhova, L. et al. Genome-wide association study in alopecia areata implicates both innate and adaptive immunity. Nature 466, 113–117 (2010).
Menashe, I., Rosenberg, P.S. & Chen, B.E. PGA: power calculator for case-control genetic association analyses. BMC Genet. 9, 36 (2008).
Acknowledgements
We are deeply grateful to all the participants as well as to the doctors working on this project. The authors also thank the editors and anonymous reviewers for their valuable comments on the manuscript. This work was supported by grants from the 973 Program (2010CB529600, 2009AA022701, 2006AA02A407), the Natural Science Foundation of China (81130022, 81121001, 31000553), the Foundation for the Author of National Excellent Doctoral Dissertation of China (201026), the Program for New Century Excellent Talents in University (NCET-09-0550), the Shanghai Municipal Health Bureau program (2008095), the Shanghai Changning Health Bureau program (2008406002), the Shanghai Municipal Commission of Science and Technology Program (09DJ1400601), the National Key Technology R&D Program (2006BAI05A09) and the Shanghai Leading Academic Discipline Project (B205).
Author information
Authors and Affiliations
Contributions
Y.S. and L.H. conceived of and designed the study. Y.S. supervised all the experiments and data analysis. Y.S. and Z.L. conducted data analyses and drafted the manuscript. Y.S., Z.L., F.Z., D.S.C., S.S., D.R. and L.H. revised the manuscript. Y.S., G.F., Q.X., J.C., Y.X., D.L., P.W., P.Y., B. Liu, W.S., G.Z. and W.J. recruited samples. T. Wang, J.J., T.L., J.S., J.C., Q.W., W.L., L.Z., H.Z., B. Li, C.W., S.Q. and G.H. performed or contributed to the experiments. S.S., S.C., T.W., E.S., S.T., A.P., M.M.N., M.R., R.A.O., D.A.C., D.R., D.S.C., H.S. and K.S. provided the SGENE-plus data. All authors critically reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–12 and Supplementary Note (PDF 463 kb)
Rights and permissions
About this article
Cite this article
Shi, Y., Li, Z., Xu, Q. et al. Common variants on 8p12 and 1q24.2 confer risk of schizophrenia. Nat Genet 43, 1224–1227 (2011). https://doi.org/10.1038/ng.980
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.980
This article is cited by
-
Genome-wide Mendelian randomization identifies actionable novel drug targets for psychiatric disorders
Neuropsychopharmacology (2023)
-
Involvement of the long intergenic non-coding RNA LINC00461 in schizophrenia
BMC Psychiatry (2022)
-
The schizophrenia-associated missense variant rs13107325 regulates dendritic spine density
Translational Psychiatry (2022)
-
Fibroblast Growth Factor Signalling in the Diseased Nervous System
Molecular Neurobiology (2021)
-
Overdispersed gene expression in schizophrenia
npj Schizophrenia (2020)