Abstract
Restriction fragment length polymorphism (RFLP) analysis is one of the oldest, most convenient and least expensive methods of genotyping, but is limited by the availability of restriction endonuclease sites. Here we present a novel method of employing CRISPR/Cas-derived RNA-guided engineered nucleases (RGENs) in RFLP analysis. We prepare RGENs by complexing recombinant Cas9 protein derived from Streptococcus pyogenes with in vitro transcribed guide RNAs that are complementary to the DNA sequences of interest. Then, we genotype recurrent mutations found in cancer and small insertions or deletions (indels) induced in cultured cells and animals by RGENs and other engineered nucleases such as transcription activator-like effector nucleases (TALENs). Unlike T7 endonuclease I or Surveyor assays that are widely used for genotyping engineered nuclease-induced mutations, RGEN-mediated RFLP analysis can detect homozygous mutant clones that contain identical biallelic indel sequences and is not limited by sequence polymorphisms near the nuclease target sites.
Similar content being viewed by others
Introduction
RGENs are derived from the adaptive immune system in bacteria and archaea known as clustered, regularly interspaced, short palindromic repeats (CRISPR)/CRISPR-associated protein (Cas)1; they now belong to a family of genome-editing engineered nucleases that induce site-specific DNA cleavages in cells and organisms, whose repair via endogenous DNA double-strand break (DSB) repair systems gives rise to targeted mutagenesis2,3 and chromosomal rearrangements4,5. Unlike zinc-finger nucleases (ZFNs) or transcription activator-like effector (TALE) nucleases (TALENs), whose DNA sequence specificities are determined by protein moieties, RGENs recognize and cleave DNA in a targeted manner using two separate components, that is, a small RNA component termed CRISPR RNA (crRNA) or single-chain guide RNA (sgRNA) that hybridizes with a 20-base pair (bp) long target DNA sequence and the Cas9 protein, derived from Streptococcus pyogenes, which recognizes the NGG (or, to a lesser extent, NAG6) trinucleotide sequence known as the protospacer-adjacent motif (PAM). Thus, an RGEN site can be shown as 5′-X20NGG-3′, where X20 corresponds to the crRNA sequence and N is any base. RGENs consist of Cas9, invariable transactivating crRNA (tracrRNA) and target-specific crRNA (or sgRNA that consists of essential portions of tracrRNA and crRNA), and are readily reprogrammed by replacing crRNA (or sgRNA)7. We and others have recently used RGENs for genome engineering in bacteria8, mammalian cells9,10,11,12, animals13,14,15,16,17,18,19 and plants20,21,22.
Engineered nuclease-induced mutations are detected by various methods, which include mismatch-sensitive Surveyor or T7 endonuclease I (T7E1) assays23, RFLP analysis24, fluorescent PCR25, DNA melting analysis26 and Sanger and deep sequencing. The T7E1 and Surveyor assays are widely used but often underestimate mutation frequencies because the assays detect heteroduplexes (formed by the hybridization of mutant and wild-type sequences or two different mutant sequences); they fail to detect homoduplexes formed by the hybridization of two identical mutant sequences. Thus, these assays cannot distinguish homozygous bialleic mutant clones from wild-type cells nor heterozygous biallelic mutants from heterozygous monoalleic mutants (Fig. 1a,b). In addition, sequence polymorphisms near the nuclease target site can produce confounding results because the enzymes can cleave heteroduplexes formed by hybridization of these different wild-type alleles. RFLP analysis is free of these limitations and therefore is a method of choice. Indeed, RFLP analysis was one of the first methods used to detect engineered nuclease-mediated mutations24. Unfortunately, however, it is limited by the availability of appropriate restriction sites.
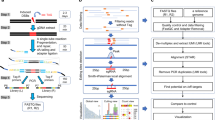
(a) A conceptual diagram of the T7E1 and RGEN-RFLP assays. Comparison of assay cleavage reactions in four possible scenarios after engineered nuclease treatment in a diploid cell: (WT) wild type, (Mono) a monoallelic mutation, (Hetero) different biallelic mutations and (Homo) identical biallelic mutations. Black lines represent PCR products derived from each allele; yellow and red boxes indicate insertion/deletion mutations generated by non-homologous end-joining. (b) Expected results of T7E1 and RGEN digestion resolved by electrophoresis. (c) In vitro cleavage assay of a linearized plasmid containing the C4BPB target site bearing indels. DNA sequences of individual plasmid substrates (bottom panel). The PAM sequence is underlined. Inserted bases are shown in red. Arrows (upper panel) indicate expected positions of DNA bands cleaved by the wild-type-specific RGEN after electrophoresis.
Here we present a novel method of detecting and quantifying engineered nucleases-induced mutations in cultured cells and organisms by employing RGENs in RFLP analysis. Unlike T7E1 or Surveyor assays, RGEN-mediated RFLP analysis can distinguish homozygous bialleic mutant clones from wild-type cells and is not limited by sequence polymorphisms near the nuclease target sites. In addition, we show that RGEN-RFLP analysis can be used to genotype naturally-occurring indels and cancer-associated mutations in human cell lines.
Results
RGEN-mediated RFLP analysis
We reasoned that RGENs can be employed in RFLP analysis, bypassing the use of conventional restriction enzymes. New RGENs with desired DNA specificities can be readily created by replacing crRNA; no de novo purification of custom proteins is required once recombinant Cas9 protein is available. Engineered nucleases, including RGENs, induce small insertions or deletions (indels) at target sites when the DSBs caused by the nucleases are repaired by error-prone non-homologous end-joining. RGENs that are designed to recognize the target sequences cleave wild-type sequences efficiently but cannot cleave mutant sequences with indels (Fig. 1a,b).
We first tested whether RGENs can differentially cleave plasmids that contain wild type or modified C4BPB target sequences that harbour 1- to 3-base indels at the cleavage site. None of the six plasmids with these indels were cleaved by a C4BPB-specific RGEN9 composed of target-specific crRNA, tracrRNA and recombinant Cas9 protein (Fig. 1c). In contrast, the plasmid with the intact target sequence was cleaved efficiently by this RGEN.
Genotyping indels induced by RGENs via RGEN-RFLP analysis
To test the feasibility of using RGEN-mediated RFLP (RGEN-RFLP in short) analysis for the detection of RGEN-induced mutations, we utilized K562 human cancer cell clones that had been modified at the C4BPB locus using the C4BPB-targeting RGEN. Among five C4BPB mutant clones we analysed, four clones had both wild-type and mutant alleles (heterozygous +/−) and one clone (#5) had only mutant alleles (compound heterozygous) (Fig. 2a). All of the mutations that occurred in these clones resulted in the loss of the RGEN target site. Thus, when the C4BPB mutant clones were subjected to RGEN genotyping, PCR amplicons of heterozygous +/− clones that contained both wild-type and mutant alleles were partially digested, and those of compound heterozygous −/− clones that did not contain the wild-type allele were not digested at all (Fig. 2a). In line with the plasmid digestion results, even a single-base insertion at the target site blocked the digestion of amplified mutant alleles by the C4BPB RGEN, showing the high specificity of RGEN genotyping. In contrast, the PCR products amplified from wild-type genomic DNA were digested completely by the RGEN. We subjected the PCR amplicons to the mismatch-sensitive T7E1 assay in parallel. Notably, the T7E1 assay failed to distinguish the compound heterozygous clone (#5) from the four +/− clones.
(a) Analysis of C4BPB-disrupted clones using RGEN-RFLP and T7E1 assays. Arrows indicate expected positions of DNA bands cleaved by RGEN or T7E1. The RGEN target site is shown in blue. Red arrows indicate the RGEN cleavage site. The PAM sequence is underlined. The number of inserted or deleted bases is shown. (b) Analysis of mouse founders that carried RGEN-induced homozygous biallelic mutations at the Foxn1 gene using RGEN-RFLP and T7E1 assays.
We also used RGEN-RFLP to analyse three mutant mice that had been created by injecting a Foxn1-targeting RGEN into one-cell stage embryos19. These founder mice carried identical biallelic mutations at the target site (Fig. 2b). As expected, the T7E1 assay failed to distinguish these homozygous mutant clones from the wild-type control, because annealing of the same mutant sequences forms homoduplexes. In contrast, the RGEN used for the creation of these mice successfully distinguished them from the wild-type control: The RGEN cleaved the PCR products amplified from wild-type genomic DNA completely but did not cleave those amplified from the mutants at all. Thus, RGEN-RFLP analysis has a critical advantage over the conventional mismatch-sensitive nuclease assay in the analysis of clones containing mutations induced by engineered nucleases.
We also investigated whether RGEN-RFLP analysis is a quantitative method. Genomic DNA samples isolated from the C4BPB null clone and the wild-type cells were mixed at various ratios and used for PCR amplifications. The PCR products were subjected to RGEN genotyping and the T7E1 assay in parallel (Fig. 3). As expected, DNA cleavage by the RGEN was proportional to the wild type to mutant ratio. In contrast, results of the T7E1 assay correlated poorly with mutation frequencies inferred from the ratios and were inaccurate, especially at high mutant %, a situation in which complementary mutant sequences can hybridize with each other to form homoduplexes.
Genomic DNA samples from wild type and C4BPB-disrupted K562 cells (clone #5 in Fig. 2a) were mixed in various ratios and subjected to RGEN-RFLP and T7E1 assays. Data points are represented as mean±s.e.m. of two independent experiments.
Next, we designed and tested a new RGEN that targets a highly polymorphic locus, HLA-B, which encodes Human Leukocyte Antigen B (also known as MHC class I protein). HeLa cells were transfected with RGEN plasmids, and the genomic DNA was subjected to T7E1 and RGEN-RFLP analyses in parallel (Fig. 4a). T7E1 produced false-positive bands that resulted from sequence polymorphisms near the target site (Fig. 4b). As expected, however, the same RGEN used for gene disruption cleaved PCR products from wild-type cells completely but those from RGEN-transfected cells partially, indicating the presence of RGEN-induced indels at the target site. This result shows that RGEN-RFLP analysis has a clear advantage over the T7E1 assay, especially when it is not known whether target genes have polymorphisms or variations in cells of interest.
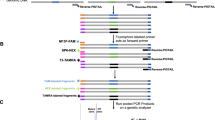
(a) HeLa cells were transfected with a HLA-B-specific RGEN. RGEN-induced mutations were analysed using RGEN-RFLP and T7E1 assays in parallel. (b) The DNA sequence, which surrounds the RGEN target site, is that of a PCR amplicon from HeLa cells. Polymorphic positions are shown in red. The RGEN target site is shown in blue. The PAM sequence is underlined. Arrows indicate PCR primer sequences.
RGEN-mediated genotyping of ZFN and TALEN-induced mutations
We then applied RGEN genotyping to the analysis of mutant mouse founders that had been established using a Pibf1-specific TALEN27 (Supplementary Fig. 1). We designed and used two RGENs specific to the TALEN target site (Fig. 5a). One RGEN recognized the NGG PAM sequence and the other RGEN recognized the NAG PAM sequence. These two RGENs successfully detected various mutations, which ranged from 1–27-bp deletions. Unlike the T7E1 assay, RGEN genotyping enabled differential detection of heterozygous +/− founders (#4 and 5) from compound heterozygous −/− founders (#1, 3, 6, 8 and 11) (Fig. 5a). In addition, we used RGENs to detect mutations induced in human cells by a CCR5-specific ZFN, representing yet another class of engineered nucleases (Fig. 5b). These results show that RGENs can detect mutations induced by nucleases other than RGENs themselves. In fact, we expect that RGENs can be designed to detect mutations induced by most, if not all, engineered nucleases. The only limitation in the design of an RGEN genotyping assay is the requirement for the GG or AG (CC or CT on the complementary strand) dinucleotide in the PAM sequence recognized by the Cas9 protein, which occurs once per 4 bp on average. Indels induced anywhere within the seed region of several bases in crRNA and the PAM nucleotides are expected to disrupt RGEN-catalyzed DNA cleavage. Indeed, we identified at least one RGEN site in most (98%) of the ZFN and TALEN sites we had reported previously4,23,28,29 (Supplementary Table 1). In contrast, only 36 sites (26%) have appropriate sites for RFLP analysis using conventional restriction enzymes.
(a) Genotyping of mouse founders generated by a PIBF-targeting TALEN. The left and right half-sites of the PIBF TALEN are shown in red and blue, respectively. The PAM sequence recognized by Cas9 is underlined. (b) RGEN-mediated genotyping of ZFN-induced mutations. The left and right half-sites of the CCR5-ZFN are shown in red and blue, respectively. Black lines indicate RGEN target sites. Arrows indicate DNA bands cleaved by RGEN or T7E1.
RGEN-RFLP analysis of naturally occurring variations
RGEN-RFLP analysis has applications beyond genotyping of engineered nuclease-induced mutations. We sought to use RGEN genotyping to detect recurrent mutations found in cancer and naturally occurring polymorphisms. We chose the human colorectal cancer cell line, HCT116, which carries a gain-of-function 3-bp deletion in the oncogenic CTNNB1 gene encoding beta-catenin30. PCR products amplified from HCT116 genomic DNA were cleaved partially by both wild-type-specific and mutant-specific RGENs, in line with the heterozygous genotype in HCT116 cells (Fig. 6a). In sharp contrast, PCR products amplified from DNA from HeLa cells harbouring only wild-type alleles were digested completely by the wild-type-specific RGEN and were not cleaved at all by the mutation-specific RGEN.
(a) A recurrent mutation (c.133–135 deletion of TCT) in the human CTNNB1 gene in HCT116 cells was detected by RGENs. (b) Genotyping of the KRAS substitution mutation (c.34G>A) in the A549 cancer cell line with RGENs that contain mismatched guide RNA. Mismatched nucleotides are shown in red. HeLa cells were used as a negative control. Arrows indicate DNA bands cleaved by RGENs. DNA sequences confirmed by Sanger sequencing are shown.
We also noted that HEK293 cells harbour the 32-bp deletion (del32) in the CCR5 gene, which encodes an essential co-receptor of HIV infection: Homozygous del32 CCR5 carriers are immune to HIV infection31. We designed one RGEN specific to the del32 allele and the other to the wild-type allele. As expected, the wild-type-specific RGEN cleaved the PCR products obtained from K562, SKBR3 or HeLa cells (used as wild-type controls) completely but those from HEK293 cells partially (Supplementary Fig. 2a), confirming the presence of the uncleavable del32 allele in HEK293 cells. Unexpectedly, however, the del32-specific RGEN cleaved the PCR products from wild-type cells as efficiently as those from HEK293 cells. Interestingly, this RGEN had an off-target site with a single-base mismatch immediately downstream of the on-target site (Supplementary Fig. 2). These results suggest that RGENs can be used to detect naturally occurring indels but cannot distinguish sequences with single-nucleotide polymorphisms or point mutations due to their off-target effects.
Genotyping of single-nucleotide variations via RGEN-RFLP
To genotype oncogenic single-nucleotide variations32 using RGENs, we attenuated RGEN activity by employing a single-base mismatched guide RNA instead of a perfectly matched RNA. RGENs that contained the perfectly matched guide RNA specific to the wild-type sequence or mutant sequence cleaved both sequences (Supplementary Figs 3 and 4). In contrast, RGENs that contained a single-base mismatched guide RNA distinguished the two sequences, enabling genotyping of three recurrent oncogenic point mutations in the KRAS, PIK3CA and IDH1 genes in human cancer cell lines (Fig. 6b and Supplementary Fig. 5a,b). In addition, we were able to detect point mutations in the BRAF and NRAS genes using RGENs that recognize the NAG PAM sequence (Supplementary Fig. 5c,d). We believe that we can use RGEN-RFLP to genotype almost any, if not all, mutations or polymorphisms in the human and other genomes.
Discussion
Here we showed that RGEN-RFLP analysis can be used for genotyping both engineered nuclease-induced mutations and naturally occurring variations in cultured cells and animals. Unlike the T7E1 assay, RGEN-RFLP analysis is not limited by sequence polymorphisms near the nuclease target sites and distinguishes homozygous biallelic mutant clones from wild-type clones and heterozygous monoallelic mutant (+/−) clones from heterozygous biallelic mutant (−/−) clones. These features can make RGEN-RFLP a method of choice for detecting indels induced by RGENs and other nucleases. We observed, however, that RGENs often cannot distinguish off-target DNA sequences with a single-nucleotide mismatch. This off-target DNA cleavage by RGENs does not limit the utility of RGEN-RFLP for measuring the frequency of nuclease-induced mutations because error-prone non-homologous end-joining repair of DSBs rarely produces single-nucleotide substitutions. We employed attenuated guide RNAs that contain a single-base mismatch rather than perfectly matched guide RNAs to distinguish the wild-type sequences from the mutant sequences with substitutions. This result is consistent with our recent report that RGENs transfected into cultured human cells discriminate on-target sites from off-target sites that differ by two bases efficiently but not those that differ by a single base33. We also found that, at least partially due to off-target DNA cleavage, RGEN concentrations need to be optimized for each RFLP analysis (Supplementary Table 2).
Next-generation sequencing platforms are now widely used for genotyping in basic and biomedical research, including genotyping of mutations induced by engineered nucleases, and will soon be used broadly for diagnostics. RFLP analysis, however, may remain a method of choice in many settings, because it is a fast and inexpensive method with no requirement for special equipment. RGEN-RFLP analysis will broaden the utility of this convenient and robust method in a wide range of applications, enabling genotyping of both naturally occurring and nuclease-induced mutations.
Methods
RGEN components
crRNA and tracrRNA were prepared by in vitro transcription using MEGAshortcript T7 kit (Ambion) according to the manufacturer’s instruction. Transcribed RNAs were resolved on an 8% denaturing urea-PAGE gel. The gel slice containing RNA was cut out and transferred to elution buffer. RNA was recovered in nuclease-free water followed by phenol:chloroform extraction, chloroform extraction and ethanol precipitation. Purified RNA was quantified by spectrometry. Templates for crRNA were prepared by annealing an oligonucleotide whose sequence is shown as 5′-GAAATTAATACGACTCACTATAGGX20GTTTTAGAGCTATGCTGTTTTG-3′, in which X20 is the target sequence, and its complementary oligonucleotide. The template for tracrRNA was synthesized by extension of forward and reverse oligonucleotides (5′-GAAATTAATACGACTCACTATAGGAACCATTC AAAACAGCATAGCAAGTTAAAATAAGGCTAGTCCG-3′ and 5′-AAAAAAAGCACCGACT CGGTGCCACTTTTTCAAGTTGATAACGGACTAGCCTTATTTTAACTTGCTATG-3′) using Phusion polymerase (New England Biolabs).
Recombinant Cas9 protein
Recombinant Cas9 protein was purchased from ToolGen, Inc. or purified from E. coli. The Cas9 DNA construct that encodes S. pyogenes Cas9 fused to the His6-tag at the C terminus was inserted in the pET-28a expression vector. The recombinant Cas9 protein was expressed in E. coli strain BL21(DE3) cultured in LB medium at 25 °C for 4 h after induction with 1 mM IPTG. Cells were harvested and resuspended in buffer containing 20 mM Tris (PH 8.0), 500 mM NaCl, 5 mM imidazole and 1 mM PMSF. Cells were frozen in liquid nitrogen, thawed at 4 °C and sonicated. After centrifugation, Cas9 protein in the lysate was bound to Ni-NTA agarose resin (Qiagen), washed with buffer containing 20 mM Tris (pH 8.0), 500 mM NaCl and 20 mM imidazole, and eluted with buffer containing 20 mM Tris (pH 8.0), 500 mM NaCl and 250 mM imidazole. Purified Cas9 protein was dialyzed against 20 mM HEPES (pH 7.5), 150 mM KCl, 1 mM DTT and 10% glycerol and analysed by SDS–PAGE.
T7 endonuclease I assay
The T7E1 assay was performed as described previously23. In brief, PCR products amplified using genomic DNA were denatured at 95 °C, reannealed at 16 °C and incubated with 5 units of T7 Endonuclease I (New England BioLabs) for 20 min at 37 °C. The reaction products were resolved using 2–2.5% agarose gel electrophoresis. The DNA sequences of PCR primers are shown in Supplementary Table 3. Full gel images are shown in Supplementary Figs 6–10.
RGEN-RFLP assay
PCR products (100–150 ng) were incubated for 60 min at 37 °C with optimized concentrations (Supplmentary Table 2) of Cas9 protein, tracrRNA, crRNA in 10 μl NEB buffer 3 (1 × ). After the cleavage reaction, RNase A (4 μg) was added, and the reaction mixture was incubated for 30 min at 37 °C to remove RNA. Reactions were stopped with 6 × stop solution buffer containing 30% glycerol, 1.2% SDS and 100 mM EDTA. Products were resolved with 1–2.5% agarose gel electrophoresis and visualized with EtBr staining. Full gel images are shown in Supplementary Figs 6–10.
Plasmid cleavage assay
Restriction enzyme-treated linearized plasmid (100 ng) was incubated for 60 min at 37 °C with Cas9 protein (0.1 μg), tracrRNA (60 ng) and crRNA (25 ng) in 10 μl NEB 3 buffer (1 × ). Reactions were stopped with 6 × stop solution containing 30% glycerol, 1.2% SDS and100 mM EDTA. Products were resolved with 1% agarose gel electrophoresis and visualized with EtBr staining.
Additional information
How to cite this article: Kim, J. M. et al. Genotyping with CRISPR-Cas-derived RNA-guided endonucleases. Nat. Commun. 5:3157 doi: 10.1038/ncomms4157 (2014).
Change history
06 February 2015
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Wiedenheft, B., Sternberg, S. H. & Doudna, J. A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 482, 331–338 (2012).
Bibikova, M., Beumer, K., Trautman, J. K. & Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 300, 764 (2003).
Porteus, M. H. & Baltimore, D. Chimeric nucleases stimulate gene targeting in human cells. Science 300, 763 (2003).
Lee, H. J., Kim, E. & Kim, J. S. Targeted chromosomal deletions in human cells using zinc finger nucleases. Genome Res. 20, 81–89 (2010).
Lee, H. J., Kweon, J., Kim, E., Kim, S. & Kim, J. S. Targeted chromosomal duplications and inversions in the human genome using zinc finger nucleases. Genome Res. 22, 539–548 (2012).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, L. A. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotechnol. 31, 233–239 (2013).
Cho, S. W., Kim, S., Kim, J. M. & Kim, J. S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 230–232 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013).
Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229 (2013).
Gratz, S. J. et al. Genome engineering of Drosophila with the CRISPR RNA-guided Cas9 nuclease. Genetics 194, 1029–1035 (2013).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-Mediated genome engineering. Cell 153, 910–918 (2013).
Chang, N. et al. Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res. 23, 465–472.
Shen, B. et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res. 23, 720–723.
Cho, S. W., Lee, J., Carroll, D. & Kim, J. S. Heritable gene knockout in Caenorhabditis elegans by direct injection of Cas9-sgRNA ribonucleoproteins. Genetics 195, 1177–1180 (2013).
Sung, Y. H. et al. Highly efficient gene knockout in mice and zebrafish with RNA-guied endonucleases. Genome Res doi: 10.1101/gr.163394.113 (2013).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 31, 686–688 (2013).
Li, J. F. et al. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat. Biotechnol. 31, 688–691 (2013).
Nekrasov, V., Staskawicz, B., Weigel, D., Jones, J. D. & Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 691–693 (2013).
Kim, H. J., Lee, H. J., Kim, H., Cho, S. W. & Kim, J. S. Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Res. 19, 1279–1288 (2009).
Urnov, F. D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005).
Kim, H., Um, E., Cho, S. R., Jung, C. & Kim, J. S. Surrogate reporters for enrichment of cells with nuclease-induced mutations. Nat. Methods 8, 941–943 (2011).
Dahlem, T. J. et al. Simple methods for generating and detecting locus-specific mutations induced with TALENs in the zebrafish genome. PLoS Genet. 8, e1002861 (2012).
Sung, Y. H. et al. Knockout mice created by TALEN-mediated gene targeting. Nat. Biotechnol. 31, 23–24 (2013).
Kim, S., Lee, M. J., Kim, H., Kang, M. & Kim, J. S. Preassembled zinc-finger arrays for rapid construction of ZFNs. Nat. Methods 8, 7 (2011).
Kim, Y. et al. A library of TAL effector nucleases spanning the human genome. Nat. Biotechnol. 31, 251–258 (2013).
Chan, T. A., Wang, Z., Dang, L. H., Vogelstein, B. & Kinzler, K. W. Targeted inactivation of CTNNB1 reveals unexpected effects of beta-catenin mutation. Proc. Natl Acad. Sci. USA 99, 8265–8270 (2002).
Samson, M. et al. Resistance to HIV-1 infection in caucasian individuals bearing mutant alleles of the CCR-5 chemokine receptor gene. Nature 382, 722–725 (1996).
Forbes, S. A. et al. COSMIC (the Catalogue of Somatic Mutations in Cancer): a resource to investigate acquired mutations in human cancer. Nucleic Acids Res. 38, D652–D657 (2010).
Cho, S. W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res doi: 10.1101/gr.162339.113 (2013).
Acknowledgements
J.-S.K. was supported by the National Research Foundation of Korea (2013000718).
Author information
Authors and Affiliations
Contributions
J.-S.K. and S.K. conceived the project. J.-S.K. and J.M.K. wrote the manuscript. J.M.K. and D.K. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
J.M.K. and J.-S.K. are authors on a patent entitled ‘Composition for cleaving a target DNA comprising a guide RNA specific for the target DNA and Cas protein’ (patent application number PCR/KR2013/009488). The remaining authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-10 and Supplementary Tables 1-3 (PDF 1354 kb)
Rights and permissions
About this article
Cite this article
Kim, J., Kim, D., Kim, S. et al. Genotyping with CRISPR-Cas-derived RNA-guided endonucleases. Nat Commun 5, 3157 (2014). https://doi.org/10.1038/ncomms4157
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms4157
This article is cited by
-
Cryptochrome 1 mediates light-dependent inclination magnetosensing in monarch butterflies
Nature Communications (2021)
-
CPEB3-mediated MTDH mRNA translational suppression restrains hepatocellular carcinoma progression
Cell Death & Disease (2020)
-
Physiological impact and comparison of mutant screening methods in piwil2 KO founder Nile tilapia produced by CRISPR/Cas9 system
Scientific Reports (2020)
-
Ryk modulates the niche activity of mesenchymal stromal cells by fine-tuning canonical Wnt signaling
Experimental & Molecular Medicine (2020)
-
A qPCR method for genome editing efficiency determination and single-cell clone screening in human cells
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.