Abstract
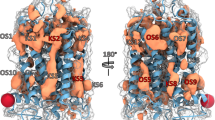
G protein–coupled receptors (GPCRs) are intensely studied as drug targets and for their role in signaling. With the determination of the first crystal structures, interest in structure-based ligand discovery increased. Unfortunately, for most GPCRs no experimental structures are available. The determination of the D3 receptor structure and the challenge to the community to predict it enabled a fully prospective comparison of ligand discovery from a modeled structure versus that of the subsequently released crystal structure. Over 3.3 million molecules were docked against a homology model, and 26 of the highest ranking were tested for binding. Six had affinities ranging from 0.2 to 3.1 μM. Subsequently, the crystal structure was released and the docking screen repeated. Of the 25 compounds selected, five had affinities ranging from 0.3 to 3.0 μM. One of the new ligands from the homology model screen was optimized for affinity to 81 nM. The feasibility of docking screens against modeled GPCRs more generally is considered.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Overington, J.P., Al-Lazikani, B. & Hopkins, A.L. How many drug targets are there? Nat. Rev. Drug Discov. 5, 993–996 (2006).
Cherezov, V. et al. High-resolution crystal structure of an engineered human β2-adrenergic G protein–coupled receptor. Science 318, 1258–1265 (2007).
Warne, T. et al. Structure of a beta1-adrenergic G-protein-coupled receptor. Nature 454, 486–491 (2008).
Jaakola, V.P. et al. The 2.6 angstrom crystal structure of a human A2A adenosine receptor bound to an antagonist. Science 322, 1211–1217 (2008).
Carlsson, J. et al. Structure-based discovery of A2A adenosine receptor ligands. J. Med. Chem. 53, 3748–3755 (2010).
Katritch, V. et al. Structure-based discovery of novel chemotypes for adenosine A2A receptor antagonists. J. Med. Chem. 53, 1799–1809 (2010).
Kolb, P. et al. Structure-based discovery of β2-adrenergic receptor ligands. Proc. Natl. Acad. Sci. USA 106, 6843–6848 (2009).
Vassilatis, D.K. et al. The G protein-coupled receptor repertoires of human and mouse. Proc. Natl. Acad. Sci. USA 100, 4903–4908 (2003).
Donnelly, D. & Findlay, J.B.C. Seven-helix receptors: structure and modelling. Curr. Opin. Struct. Biol. 4, 582–589 (1994).
Kurczab, R., Nowak, M., Chilmonczyk, Z., Sylte, I. & Bojarski, A.J. The development and validation of a novel virtual screening cascade protocol to identify potential serotonin 5–HT7R antagonists. Bioorg. Med. Chem. Lett. 20, 2465–2468 (2010).
Salo, O.M.H. et al. Virtual screening of novel CB2 ligands using a comparative model of the human cannabinoid CB2 receptor. J. Med. Chem. 48, 7166–7171 (2005).
Bissantz, C., Schalon, C., Guba, W. & Stahl, M. Focused library design in GPCR projects on the example of 5–HT2c agonists: Comparison of structure-based virtual screening with ligand-based search methods. Proteins 61, 938–952 (2005).
Kratochwil, N.A. et al. An automated system for the analysis of G protein-coupled receptor transmembrane binding pockets: alignment, receptor-based pharmacophores, and their application. J. Chem. Inf. Model. 45, 1324–1336 (2005).
Shi, L. & Javitch, J.A. The binding site of aminergic G protein-coupled receptors: the transmembrane segments and second extracellular loop. Annu. Rev. Pharmacol. Toxicol. 42, 437–467 (2002).
Shi, L. & Javitch, J.A. The second extracellular loop of the dopamine D2 receptor lines the binding-site crevice. Proc. Natl. Acad. Sci. USA 101, 440–445 (2004).
de Graaf, C., Foata, N., Engkvist, O. & Rognan, D. Molecular modeling of the second extracellular loop of G-protein coupled receptors and its implication on structure-based virtual screening. Proteins 71, 599–620 (2008).
de Graaf, C., Rognan, D. & Customizing, G. Protein-coupled receptor models for structure-based virtual screening. Curr. Pharm. Des. 15, 4026–4048 (2009).
Michino, M. et al. Community-wide assessment of GPCR structure modelling and ligand docking: GPCR Dock 2008. Nat. Rev. Drug Discov. 8, 455–463 (2009).
Tikhonova, I.G. et al. Discovery of novel agonists and antagonists of the free fatty acid receptor 1 (FFAR1) using virtual screening. J. Med. Chem. 51, 625–633 (2008).
Varady, J. et al. Molecular modeling of the three-dimensional structure of dopamine 3 (D3) subtype receptor: discovery of novel and potent D3 ligands through a hybrid pharmacophore- and structure-based database searching approach. J. Med. Chem. 46, 4377–4392 (2003).
Becker, O.M. et al. G protein-coupled receptors: In silico drug discovery in 3D. Proc. Natl. Acad. Sci. USA 101, 11304–11309 (2004).
Engel, S. et al. A virtual screen for diverse ligands: discovery of selective G protein-coupled receptor antagonists. J. Am. Chem. Soc. 130, 5115–5123 (2008).
Evers, A. & Klabunde, T. Structure-based drug discovery using GPCR homology modeling: successful virtual screening for antagonists of the α1A adrenergic receptor. J. Med. Chem. 48, 1088–1097 (2005).
Evers, A. & Klebe, G. Successful virtual screening for a submicromolar antagonist of the neurokinin-1 receptor based on a ligand-supported homology model. J. Med. Chem. 47, 5381–5392 (2004).
Kellenberger, E. et al. Identification of nonpeptide CCR5 receptor agonists by structure-based virtual screening. J. Med. Chem. 50, 1294–1303 (2007).
Cavasotto, C.N. et al. Discovery of novel chemotypes to a G-protein-coupled receptor through ligand-steered homology modeling and structure-based virtual screening. J. Med. Chem. 51, 581–588 (2008).
Kiss, R. et al. Discovery of novel human histamine H4 receptor ligands by large-scale structure-based virtual screening. J. Med. Chem. 51, 3145–3153 (2008).
Kufareva, I. et al. Status of GPCR modeling and docking as reflected by community-wide GPCR Dock 2010 assessment. Structure 19, 1108–1126 (2011).
Chien, E.Y.T. et al. Structure of the human dopamine D3 receptor in complex with a D2/D3 selective antagonist. Science 330, 1091–1095 (2010).
Evers, A., Gohlke, H. & Klebe, G. Ligand-supported homology modelling of protein binding-sites using lnowledge-based potentials. J. Mol. Biol. 334, 327–345 (2003).
Katritch, V., Rueda, M., Lam, P.C.-H., Yeager, M. & Abagyan, R. GPCR 3D homology models for ligand screening: lessons learned from blind predictions of adenosine A2a receptor complex. Proteins 78, 197–211 (2010).
Yang, Q. & Sharp, K.A. Building alternate protein structures using the elastic network model. Proteins 74, 682–700 (2009).
Eswar, N. et al. Comparative protein structure modeling with Modeller. Curr. Protoc. Bioinformatics 5, 1–30 (2006).
Shen, M.-Y. & Sali, A. Statistical potential for assessment and prediction of protein structures. Protein Sci. 15, 2507–2524 (2006).
Huang, N., Shoichet, B.K. & Irwin, J.J. Benchmarking sets for molecular docking. J. Med. Chem. 49, 6789–6801 (2006).
Ballesteros, J.A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995).
Kuhn, B., Mohr, P. & Stahl, M. Intramolecular hydrogen bonding in medicinal chemistry. J. Med. Chem. 53, 2601–2611 (2010).
Irwin, J.J. et al. Automated docking screens: a feasibility study. J. Med. Chem. 52, 5712–5720 (2009).
Mysinger, M.M. & Shoichet, B.K. Rapid context-dependent ligand desolvation in molecular docking. J. Chem. Inf. Model. 50, 1561–1573 (2010).
Irwin, J.J. & Shoichet, B.K. ZINC—A free database of commercially available compounds for virtual screening. J. Chem. Inf. Model. 45, 177–182 (2005).
Newman, A.H. et al. N-(4-(4-(2,3-Dichloro- or 2-methoxyphenyl)piperazin-1-yl)butyl)heterobiarylcarboxamides with functionalized linking chains as high affinity and enantioselective D3 receptor antagonists. J. Med. Chem. 52, 2559–2570 (2009).
Bucci, M., Goodman, C. & Sheppard, T.L. A decade of chemical biology. Nat. Chem. Biol. 6, 847–854 (2010).
Kobilka, B.K. & Deupi, X. Conformational complexity of G-protein-coupled receptors. Trends Pharmacol. Sci. 28, 397–406 (2007).
Hajduk, P.J. & Greer, J. A decade of fragment-based drug design: strategic advances and lessons learned. Nat. Rev. Drug Discov. 6, 211–219 (2007).
Gloriam, D.E., Foord, S.M., Blaney, F.E. & Garland, S.L. Definition of the G protein-coupled receptor transmembrane bundle binding pocket and calculation of receptor similarities for drug design. J. Med. Chem. 52, 4429–4442 (2009).
Pei, J., Kim, B.H. & Grishin, N.V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008).
Allen, F.H. The Cambridge Structural Database: a quarter of a million crystal structures and rising. Acta Crystallogr. B 58, 380–388 (2002).
Ferreira, R.S. et al. Complementarity between a docking and a high-throughput screen in discovering new cruzain inhibitors. J. Med. Chem. 53, 4891–4905 (2010).
Jensen, N.H. et al. N-Desalkylquetiapine, a potent norepinephrine reuptake inhibitor and partial 5-HT1A agonist, as a putative mediator of quetiapine′s antidepressant activity. Neuropsychopharmacology 33, 2303–2312 (2008).
Barnea, G. et al. The genetic design of signaling cascades to record receptor activation. Proc. Natl. Acad. Sci. USA 105, 64–69 (2008).
Acknowledgements
We thank Q. Yang and K. Sharp for help with 3K-ENM, M. Burlingame with help with compound LC/MS and G. Barnea (Brown University) for supplying beginning Tango constructs and a plasmid encoding the human V2 Tango receptor. We thank OpenEye Scientific Software for the use of OEChem and Omega. Supported by US National Institutes of Health grants GM59957 and GM71630 (to B.K.S.), U54GM093342 (to A. Sali and B.K.S.), GM71790 and R01GM54762 (to A. Sali), the National Institutes of Mental Health Psychoactive Drug Screening Program (to B.L.R.), postdoctoral fellowships from the Knut and Alice Wallenberg Foundation (to J.C.) and National Research Service Award-Kirschstein fellowships F32GM096544 (to R.G.C.) and F32GM088991 (to A. Schlessinger).
Author information
Authors and Affiliations
Contributions
Docking and homology modeling was conducted by J.C. and R.G.C., the latter with assistance from H.F., A. Schlessinger and A. Sali, J.J.I. and B.K.S. assisted with compound selection and strategy, J.J.I. lent expertise with the DOCK Blaster toolchain and fixed problems with ZINC. B.L.R. and V.S. were responsible for the pharmacological guidance as to the appropriate specificity tests, while V.S. conducted all the experiments. All authors contributed to the writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 1772 kb)
Supplementary Data Set 1
SupplementaryDataset1.xlsx (XLSX 146 kb)
Supplementary Data Set 2
model_receptor.txt (TXT 137 kb)
Supplementary Data Set 3
model_ligands.txt (TXT 20 kb)
Supplementary Data Set 4
crystal_receptor.txt (TXT 138 kb)
Supplementary Data Set 5
crystal_ligands.txt (TXT 16 kb)
Rights and permissions
About this article
Cite this article
Carlsson, J., Coleman, R., Setola, V. et al. Ligand discovery from a dopamine D3 receptor homology model and crystal structure. Nat Chem Biol 7, 769–778 (2011). https://doi.org/10.1038/nchembio.662
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.662
This article is cited by
-
A practical guide to large-scale docking
Nature Protocols (2021)
-
Ultra-large library docking for discovering new chemotypes
Nature (2019)
-
A perspective on multi‐target drug discovery and design for complex diseases
Clinical and Translational Medicine (2018)
-
Fungal G-protein-coupled receptors: mediators of pathogenesis and targets for disease control
Nature Microbiology (2018)
-
A new era of rationally designed antipsychotics
Nature (2018)