Abstract
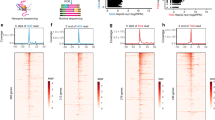
Pharmacological perturbation is a powerful tool for understanding mRNA synthesis, but identification of the specific steps of this multi-step process that are targeted by small molecules remains challenging. Here we applied strand-specific total RNA sequencing (RNA-seq) to identify and distinguish specific pharmacological effects on transcription and pre-mRNA processing in human cells. We found unexpectedly that the natural product isoginkgetin, previously described as a splicing inhibitor, inhibits transcription elongation. Compared to well-characterized elongation inhibitors that target CDK9, isoginkgetin caused RNA polymerase accumulation within a broader promoter-proximal band, indicating that elongation inhibition by isoginkgetin occurs after release from promoter-proximal pause. RNA-seq distinguished isoginkgetin and CDK9 inhibitors from topoisomerase I inhibition, which alters elongation across gene bodies. We were able to detect these and other specific defects in mRNA synthesis at low sequencing depth using simple metagene-based metrics. These metrics now enable total-RNA-seq-based screening for high-throughput identification of pharmacological effects on individual stages of mRNA synthesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Referenced accessions
Gene Expression Omnibus
References
Bensaude, O. Inhibiting eukaryotic transcription: which compound to choose? How to evaluate its activity? Transcription 2, 103–108 (2011).
Chodosh, L.A., Fire, A., Samuels, M. & Sharp, P.A. 5,6-Dichloro-1-beta-D-ribofuranosylbenzimidazole inhibits transcription elongation by RNA polymerase II in vitro. J. Biol. Chem. 264, 2250–2257 (1989).
Marshall, N.F. & Price, D.H. Control of formation of two distinct classes of RNA polymerase II elongation complexes. Mol. Cell. Biol. 12, 2078–2090 (1992).
Wada, T. et al. DSIF, a novel transcription elongation factor that regulates RNA polymerase II processivity, is composed of human Spt4 and Spt5 homologs. Genes Dev. 12, 343–356 (1998).
Yamaguchi, Y. et al. NELF, a multisubunit complex containing RD, cooperates with DSIF to repress RNA polymerase II elongation. Cell 97, 41–51 (1999).
Chao, S.H. et al. Flavopiridol inhibits P-TEFb and blocks HIV-1 replication. J. Biol. Chem. 275, 28345–28348 (2000).
Chao, S.H. & Price, D.H. Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo. J. Biol. Chem. 276, 31793–31799 (2001).
Pirngruber, J. et al. CDK9 directs H2B monoubiquitination and controls replication-dependent histone mRNA 3′-end processing. EMBO Rep. 10, 894–900 (2009).
Rahl, P.B. et al. c-Myc regulates transcriptional pause release. Cell 141, 432–445 (2010).
Laitem, C. et al. CDK9 inhibitors define elongation checkpoints at both ends of RNA polymerase II-transcribed genes. Nat. Struct. Mol. Biol. 22, 396–403 (2015).
Jonkers, I. & Lis, J.T. Getting up to speed with transcription elongation by RNA polymerase II. Nat. Rev. Mol. Cell Biol. 16, 167–177 (2015).
Kwak, H. & Lis, J.T. Control of transcriptional elongation. Annu. Rev. Genet. 47, 483–508 (2013).
Adelman, K. & Lis, J.T. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nat. Rev. Genet. 13, 720–731 (2012).
Bösken, C.A. et al. The structure and substrate specificity of human Cdk12/Cyclin K. Nat. Commun. 5, 3505 (2014).
Eick, D. & Geyer, M. The RNA polymerase II carboxy-terminal domain (CTD) code. Chem. Rev. 113, 8456–8490 (2013).
Moore, M.J. & Proudfoot, N.J. Pre-mRNA processing reaches back to transcription and ahead to translation. Cell 136, 688–700 (2009).
Pawellek, A. et al. Identification of small-molecule inhibitors of pre-mRNA splicing. J. Biol. Chem. 289, 34683–34698 (2014).
Disney, M.D. Short-circuiting RNA splicing. Nat. Chem. Biol. 4, 723–724 (2008).
O'Brien, K., Matlin, A.J., Lowell, A.M. & Moore, M.J. The biflavonoid isoginkgetin is a general inhibitor of pre-mRNA splicing. J. Biol. Chem. 283, 33147–33154 (2008).
Gray, J.M. et al. SnapShot-seq: a method for extracting genome-wide, in vivo mRNA dynamics from a single total RNA sample. PLoS One 9, e89673 (2014).
Tseng, C.-K.K. et al. Human telomerase RNA processing and quality control. Cell Rep. 13, 2232–2243 (2015).
Huranová, M. et al. The differential interaction of snRNPs with pre-mRNA reveals splicing kinetics in living cells. J. Cell Biol. 191, 75–86 (2010).
Kaida, D. et al. U1 snRNP protects pre-mRNAs from premature cleavage and polyadenylation. Nature 468, 664–668 (2010).
Almada, A.E., Wu, X., Kriz, A.J., Burge, C.B. & Sharp, P.A. Promoter directionality is controlled by U1 snRNP and polyadenylation signals. Nature 499, 360–363 (2013).
Ntini, E. et al. Polyadenylation site-induced decay of upstream transcripts enforces promoter directionality. Nat. Struct. Mol. Biol. 20, 923–928 (2013).
Ji, X. et al. SR proteins collaborate with 7SK and promoter-associated nascent RNA to release paused polymerase. Cell 153, 855–868 (2013).
Lin, S., Coutinho-Mansfield, G., Wang, D., Pandit, S. & Fu, X.-D. The splicing factor SC35 has an active role in transcriptional elongation. Nat. Struct. Mol. Biol. 15, 819–826 (2008).
Misteli, T. & Spector, D.L. RNA polymerase II targets pre-mRNA splicing factors to transcription sites in vivo. Mol. Cell 3, 697–705 (1999).
Gaidatzis, D., Burger, L., Florescu, M. & Stadler, M.B. Analysis of intronic and exonic reads in RNA-seq data characterizes transcriptional and post-transcriptional regulation. Nat. Biotechnol. 33, 722–729 (2015).
Khodor, Y.L. et al. Nascent-seq indicates widespread cotranscriptional pre-mRNA splicing in Drosophila. Genes Dev. 25, 2502–2512 (2011).
Preker, P. et al. RNA exosome depletion reveals transcription upstream of active human promoters. Science 322, 1851–1854 (2008).
Jonkers, I., Kwak, H. & Lis, J.T. Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. eLife 3, e02407 (2014).
Albert, B.J. et al. Meayamycin inhibits pre-messenger RNA splicing and exhibits picomolar activity against multidrug-resistant cells. Mol. Cancer Ther. 8, 2308–2318 (2009).
Gao, Y., Vogt, A., Forsyth, C.J. & Koide, K. Comparison of splicing factor 3b inhibitors in human cells. ChemBioChem 14, 49–52 (2013).
Lutz, C.S. et al. Interaction between the U1 snRNP-A protein and the 160-kD subunit of cleavage-polyadenylation specificity factor increases polyadenylation efficiency in vitro. Genes Dev. 10, 325–337 (1996).
Mayer, A. et al. Native elongating transcript sequencing reveals human transcriptional activity at nucleotide resolution. Cell 161, 541–554 (2015).
Capranico, G. et al. The effects of camptothecin on RNA polymerase II transcription: roles of DNA topoisomerase I. Biochimie 89, 482–489 (2007).
Ljungman, M. & Hanawalt, P.C. The anti-cancer drug camptothecin inhibits elongation but stimulates initiation of RNA polymerase II transcription. Carcinogenesis 17, 31–35 (1996).
Darzacq, X. et al. In vivo dynamics of RNA polymerase II transcription. Nat. Struct. Mol. Biol. 14, 796–806 (2007).
Singh, J. & Padgett, R.A. Rates of in situ transcription and splicing in large human genes. Nat. Struct. Mol. Biol. 16, 1128–1133 (2009).
Veloso, A. et al. Genome-wide transcriptional effects of the anti-cancer agent camptothecin. PLoS One 8, e78190 (2013).
Stoilov, P., Lin, C.-H., Damoiseaux, R., Nikolic, J. & Black, D.L. A high-throughput screening strategy identifies cardiotonic steroids as alternative splicing modulators. Proc. Natl. Acad. Sci. USA 105, 11218–11223 (2008).
Younis, I. et al. Rapid-response splicing reporter screens identify differential regulators of constitutive and alternative splicing. Mol. Cell. Biol. 30, 1718–1728 (2010).
Fong, N. et al. Effects of transcription elongation rate and Xrn2 exonuclease activity on RNA polymerase II termination suggest widespread kinetic competition. Mol. Cell 60, 256–267 (2015).
Sansó, M. et al. P-TEFb regulation of transcription termination factor Xrn2 revealed by a chemical genetic screen for Cdk9 substrates. Genes Dev. 30, 117–131 (2016).
Shishkin, A.A. et al. Simultaneous generation of many RNA-seq libraries in a single reaction. Nat. Methods 12, 323–325 (2015).
Core, L.J., Waterfall, J.J. & Lis, J.T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Rabani, M. et al. Metabolic labeling of RNA uncovers principles of RNA production and degradation dynamics in mammalian cells. Nat. Biotechnol. 29, 436–442 (2011).
Schwalb, B. et al. TT-seq maps the human transient transcriptome. Science 352, 1225–1228 (2016).
Parkhomchuk, D. et al. Transcriptome analysis by strand-specific sequencing of complementary DNA. Nucleic Acids Res. 37, e123 (2009).
Mayer, A. & Churchman, L.S. Genome-wide profiling of RNA polymerase transcription at nucleotide resolution in human cells with native elongating transcript sequencing. Nat. Protoc. 11, 813–833 (2016).
Wuarin, J. & Schibler, U. Physical isolation of nascent RNA chains transcribed by RNA polymerase II: evidence for cotranscriptional splicing. Mol. Cell. Biol. 14, 7219–7225 (1994).
Churchman, L.S. & Weissman, J.S. Nascent transcript sequencing visualizes transcription at nucleotide resolution. Nature 469, 368–373 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Acknowledgements
We thank M. Hemberg and D. Harmin for advice on bioinformatic analyses. This work was funded by R01 MH101528-01. M.S. is also supported by National Science Foundation grant 1349248. S.A.B. is supported by the Harvard Medical School Center of Excellence in Systems Pharmacology NIH grant P50 GM107618 and the Giovanni Armenise-Harvard Foundation. H.M.L. and L.S.C's contributions were funded by NHGRI: R01 HG007173. We thank K. Koide (University of Pittsburgh) for sharing meayamycin.
Author information
Authors and Affiliations
Contributions
M.S. and J.M.G. conceived the study. S.A.B. performed all of the experiments except those mentioned below. A.S. performed the western blot in Figure 4a and data analysis of meayamycin experiments. H.M.L. performed the NET-seq protocol. H.M.L. and L.S.C. helped analyze the NET-Seq data. J.M.G., S.A.B., and M.S. performed bioinformatics analysis. J.M.G., S.A.B., A.S., and M.S. interpreted the results. J.M.G., S.A.B., and M.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Table 1 and Supplementary Figures 1–5 (PDF 9252 kb)
Rights and permissions
About this article
Cite this article
Boswell, S., Snavely, A., Landry, H. et al. Total RNA-seq to identify pharmacological effects on specific stages of mRNA synthesis. Nat Chem Biol 13, 501–507 (2017). https://doi.org/10.1038/nchembio.2317
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2317
This article is cited by
-
GeneWalk identifies relevant gene functions for a biological context using network representation learning
Genome Biology (2021)
-
Target identification of hepatic fibrosis using Pien Tze Huang based on mRNA and lncRNA
Scientific Reports (2021)
-
Splicing modulators: on the way from nature to clinic
The Journal of Antibiotics (2021)
-
Hinokiflavone and Related C–O–C-Type Biflavonoids as Anti-cancer Compounds: Properties and Mechanism of Action
Natural Products and Bioprospecting (2021)