Abstract
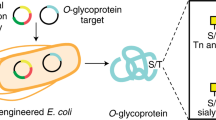
The Campylobacter jejuni protein glycosylation locus (pgl) encodes machinery for asparagine-linked (N-linked) glycosylation and serves as the archetype for bacterial N-linked glycosylation. This machinery has been functionally transferred into Escherichia coli, enabling convenient mechanistic dissection of the N-linked glycosylation process in this genetically tractable host. Here we sought to identify sequence determinants in the oligosaccharyltransferase PglB that restrict its specificity to only those glycan acceptor sites containing a negatively charged residue at the −2 position relative to asparagine. This involved creation of a genetic assay, glycosylation of secreted N-linked acceptor proteins (glycoSNAP), that facilitates high-throughput screening of glycophenotypes in E. coli. Using this assay, we isolated several C. jejuni PglB variants that could glycosylate an array of noncanonical acceptor sequences, including one in a eukaryotic N-glycoprotein. These results underscore the utility of glycoSNAP for shedding light on poorly understood aspects of N-linked glycosylation and for engineering designer N-linked glycosylation biocatalysts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Apweiler, R., Hermjakob, H. & Sharon, N. On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta 1473, 4–8 (1999).
Zielinska, D.F., Gnad, F., Wisniewski, J.R. & Mann, M. Precision mapping of an in vivo N-glycoproteome reveals rigid topological and sequence constraints. Cell 141, 897–907 (2010).
Helenius, A. & Aebi, M. Intracellular functions of N-linked glycans. Science 291, 2364–2369 (2001).
Helenius, A. & Aebi, M. Roles of N-linked glycans in the endoplasmic reticulum. Annu. Rev. Biochem. 73, 1019–1049 (2004).
Varki, A. Biological roles of oligosaccharides: all of the theories are correct. Glycobiology 3, 97–130 (1993).
Mitra, N., Sinha, S., Ramya, T.N. & Surolia, A. N-linked oligosaccharides as outfitters for glycoprotein folding, form and function. Trends Biochem. Sci. 31, 156–163 (2006).
Aebi, M., Bernasconi, R., Clerc, S. & Molinari, M. N-glycan structures: recognition and processing in the ER. Trends Biochem. Sci. 35, 74–82 (2010).
Abu-Qarn, M., Eichler, J. & Sharon, N. Not just for Eukarya anymore: protein glycosylation in Bacteria and Archaea. Curr. Opin. Struct. Biol. 18, 544–550 (2008).
Schwarz, F. & Aebi, M. Mechanisms and principles of N-linked protein glycosylation. Curr. Opin. Struct. Biol. 21, 576–582 (2011).
Szymanski, C.M. & Wren, B.W. Protein glycosylation in bacterial mucosal pathogens. Nat. Rev. Microbiol. 3, 225–237 (2005).
Larkin, A., Chang, M.M., Whitworth, G.E. & Imperiali, B. Biochemical evidence for an alternate pathway in N-linked glycoprotein biosynthesis. Nat. Chem. Biol. 9, 367–373 (2013).
Zufferey, R. et al. STT3, a highly conserved protein required for yeast oligosaccharyl transferase activity in vivo. EMBO J. 14, 4949–4960 (1995).
Lizak, C., Gerber, S., Numao, S., Aebi, M. & Locher, K.P. X-ray structure of a bacterial oligosaccharyltransferase. Nature 474, 350–355 (2011).
Matsumoto, S. et al. Crystal structures of an archaeal oligosaccharyltransferase provide insights into the catalytic cycle of N-linked protein glycosylation. Proc. Natl. Acad. Sci. USA 110, 17868–17873 (2013).
Kowarik, M. et al. Definition of the bacterial N-glycosylation site consensus sequence. EMBO J. 25, 1957–1966 (2006).
Pandhal, J. et al. Inverse metabolic engineering to improve Escherichia coli as an N-glycosylation host. Biotechnol. Bioeng. 110, 2482–2493 (2013).
Ihssen, J. et al. Structural insights from random mutagenesis of Campylobacter jejuni oligosaccharyltransferase PglB. BMC Biotechnol. 12, 67 (2012).
Çelik, E., Fisher, A.C., Guarino, C., Mansell, T.J. & DeLisa, M.P. A filamentous phage display system for N-linked glycoproteins. Protein Sci. 19, 2006–2013 (2010).
Dürr, C., Nothaft, H., Lizak, C., Glockshuber, R. & Aebi, M. The Escherichia coli glycophage display system. Glycobiology 20, 1366–1372 (2010).
Valderrama-Rincon, J.D. et al. An engineered eukaryotic protein glycosylation pathway in Escherichia coli. Nat. Chem. Biol. 8, 434–436 (2012).
Mally, M. et al. Glycoengineering of host mimicking type-2 LacNAc polymers and Lewis X antigens on bacterial cell surfaces. Mol. Microbiol. 87, 112–131 (2013).
Fisher, A.C. et al. Production of secretory and extracellular N-linked glycoproteins in Escherichia coli. Appl. Environ. Microbiol. 77, 871–881 (2011).
Wacker, M. et al. N-linked glycosylation in Campylobacter jejuni and its functional transfer into E. coli. Science 298, 1790–1793 (2002).
Zhang, G., Brokx, S. & Weiner, J.H. Extracellular accumulation of recombinant proteins fused to the carrier protein YebF in Escherichia coli. Nat. Biotechnol. 24, 100–104 (2006).
Feldman, M.F. et al. Engineering N-linked protein glycosylation with diverse O antigen lipopolysaccharide structures in Escherichia coli. Proc. Natl. Acad. Sci. USA 102, 3016–3021 (2005).
Linton, D., Allan, E., Karlyshev, A.V., Cronshaw, A.D. & Wren, B.W. Identification of N-acetylgalactosamine-containing glycoproteins PEB3 and CgpA in Campylobacter jejuni. Mol. Microbiol. 43, 497–508 (2002).
Kowarik, M. et al. N-linked glycosylation of folded proteins by the bacterial oligosaccharyltransferase. Science 314, 1148–1150 (2006).
Gavel, Y. & von Heijne, G. Sequence differences between glycosylated and non-glycosylated Asn-X-Thr/Ser acceptor sites: implications for protein engineering. Protein Eng. 3, 433–442 (1990).
Schwarz, F. et al. Relaxed acceptor site specificity of bacterial oligosaccharyltransferase in vivo. Glycobiology 21, 45–54 (2011).
Ielmini, M.V. & Feldman, M.F. Desulfovibrio desulfuricans PglB homolog possesses oligosaccharyltransferase activity with relaxed glycan specificity and distinct protein acceptor sequence requirements. Glycobiology 21, 734–742 (2011).
Gerber, S. et al. Mechanism of bacterial oligosaccharyltransferase: in vitro quantification of sequon binding and catalysis. J. Biol. Chem. 288, 8849–8861 (2013).
Haitjema, C.H. et al. Universal Genetic Assay for Engineering Extracellular Protein Expression. ACS Synth. Biol. 3, 74–82 (2013).
Shanks, R.M., Caiazza, N.C., Hinsa, S.M., Toutain, C.M. & O'Toole, G.A. Saccharomyces cerevisiae-based molecular tool kit for manipulation of genes from gram-negative bacteria. Appl. Environ. Microbiol. 72, 5027–5036 (2006).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1–I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Zhang, S. et al. Comparative characterization of the glycosylation profiles of an influenza hemagglutinin produced in plant and insect hosts. Proteomics 12, 1269–1288 (2012).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006).
Crooks, G.E., Hon, G., Chandonia, J.M. & Brenner, S.E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
We thank C. Guarino (Cornell University) for plasmid pSF-ClPglB and pBS-scFv13-R4DQNAT, J. Merritt (Glycobia, Inc.) for plasmid pMW07-pglΔB, B. Clemons (California Institute of Technology) for plasmid pET33b-ClStt3, M. Aebi (ETH Zürich) for providing antiserum used in this work, and R. Sherwood for his technical assistance acquiring the LC-MS/MS raw data files. This material is based on work supported by the US National Science Foundation grant CBET 1159581 (to M.P.D.) and National Institutes of Health (NIH) grant R44 GM088905-01 (to A.C.F. and M.P.D.) and NIH Shared Instrumentation Grant grant 1S10RR025449-01 (to S.Z.).
Author information
Authors and Affiliations
Contributions
A.A.O. designed research, performed research, analyzed data and wrote the paper. S.Z. performed MS analysis, analyzed MS data and wrote the paper. A.C.F. conceptualized project, designed research and analyzed data. M.P.D. conceptualized project, designed research, analyzed data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
A.C.F. is an employee of Glycobia, Inc. A.C.F. and M.P.D. have a financial interest in Glycobia, Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1–8. (PDF 26657 kb)
Rights and permissions
About this article
Cite this article
Ollis, A., Zhang, S., Fisher, A. et al. Engineered oligosaccharyltransferases with greatly relaxed acceptor-site specificity. Nat Chem Biol 10, 816–822 (2014). https://doi.org/10.1038/nchembio.1609
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1609
This article is cited by
-
Rapid biosynthesis of glycoprotein therapeutics and vaccines from freeze-dried bacterial cell lysates
Nature Protocols (2023)
-
Strategies for efficient production of recombinant proteins in Escherichia coli: alleviating the host burden and enhancing protein activity
Microbial Cell Factories (2022)
-
Genetic and process engineering strategies for enhanced recombinant N-glycoprotein production in bacteria
Microbial Cell Factories (2021)
-
Engineering orthogonal human O-linked glycoprotein biosynthesis in bacteria
Nature Chemical Biology (2020)
-
Application of Genetic Engineering in Biotherapeutics Development
Journal of Pharmaceutical Innovation (2020)