Abstract
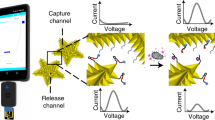
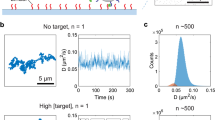
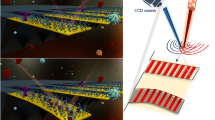
Diagnostic technologies that can provide the simultaneous detection of nucleic acids for gene expression, proteins for host response and small molecules for profiling the human metabolome will have a significant advantage in providing comprehensive patient monitoring. Molecular sensors that report changes in the electrostatics of a sensor's surface on analyte binding have shown unprecedented sensitivity in the detection of charged biomolecules, but do not lend themselves to the detection of small molecules, which do not carry significant charge. Here, we introduce the neutralizer displacement assay that allows charge-based sensing to be applied to any class of molecule irrespective of the analyte charge. The neutralizer displacement assay starts with an aptamer probe bound to a neutralizer. When analyte binding occurs the neutralizer is displaced, which results in a dramatic change in the surface charge for all types of analytes. We have tested the sensitivity, speed and specificity of this system in the detection of a panel of molecules: (deoxy)ribonucleic acid, ribonucleic acid, cocaine, adenosine triphosphate and thrombin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Xia, F. et al. Colorimetric detection of DNA, small molecules, proteins, and ions using unmodified gold nanoparticles and conjugated polyelectrolytes. Proc. Natl Acad. Sci. USA 107, 10837–10841 (2010).
Zhang, M., Win, B. C., Tan, W. & We, B. C. A versatile graphene-based fluorescence ‘on/off’ switch for multiplex detection of various targets. Biosens. Bioelectron. 26, 3260–3265 (2011).
Drummond, T. G., Hill, M. G. & Barton, J. K. Electrochemical DNA sensors. Nature Biotechnol. 21, 1192–1199 (2008).
Clack, N. G., Asalaita, K. & Groves, J. T. Electrostatic readout of DNA microarrays with charged microspheres. Nature Biotechnol. 26, 825–830 (2008).
Morrow, T. J., Li, M., Kim, J., Mayer, T. S. & Keating, C. D. Programmed assembly of DNA-coated nanowire devices. Science 323, 352 (2009).
Zheng, G., Patolsky, F., Cui, Y., Wang, U. W. & Lieber, C. M. Multiplexed electrical detection of cancer markers with nanowire sensors arrays. Nature Biotechnol. 23, 1294–1300 (2005).
Patolsky, F., Zheng, G. & Lieber, C. M. Fabrication of silicon nanowire devices for ultrasensitive, label-free, real-time detection of biological and chemical species. Nature Protocols 1, 1711–1724 (2006).
Tian, B. et al. Three-dimensional, flexible nanoscale field-effect transistors as localized bioprobes. Science 13, 830–834 (2010).
Zheng. G., Gao, X. P. A. & Lieber, C. M. Frequency domain detection of biomolecules using silicon nanowire biosensors. Nano Lett. 10, 3179–3183 (2010).
Wu, G et al. Bioassay of prostate-specific antigen (PSA) using microcantilevers. Nature Biotechnol. 19, 856–860 (2001).
Fritz, J. et al. Translating biomolecular recognition into nanomechanics. Science 288, 316–318 (2000).
Xiang, Y. & Lu, Y. Using personal glucose meters and functional DNA sensors to quantify a variety of analytical targets. Nature Chem. 3, 697–670 (2011).
Cheng, A. K. H., Ge, B. X. & Yu, H. Z. Label-free voltammetric detection of lysozyme with aptamer-modified gold electrodes. Anal. Chem. 79, 5158–5164 (2007).
Baker, B. R. et al. An electronic, aptamer-based small molecule sensor for the rapid, reagentless detection of cocaine in adulterated samples and biological fluids. J. Am. Chem. Soc. 128, 3138–3139 (2006).
Xiao, Y., Lai, R. Y. & Plaxco, K. W. Preparation of electrode-immobilized, redox-modified oligonucleotides for electrochemical DNA and aptamer-based sensing. Nature Protocols 2, 2875–2880 (2007).
Cash, K. J., Ricci, F. & Plaxco, K. W. An electrochemical sensor for the detection of protein–small molecule interactions directly in serum and other complex matrices. J. Am. Chem. Soc. 131, 6955–6957 (2009).
Zuo, X. et al. A target-responsive electrochemical aptamer switch (TREAS) for reagentless detection of nanomolar ATP. J. Am. Chem. Soc. 129, 1042–1043 (2007).
Das, J., Aziz, M. A. & Yang, H. A nanocatalyst-based assay for proteins: DNA-free ultrasensitive electrochemical detection using catalytic reduction of p-nitrophenol by gold-nanoparticle labels. J. Am. Chem. Soc. 128, 16022–16023 (2006).
Das, J., Jo, K., Lee, J. W. & Yang, H. Electrochemical immunosensor using p-aminophenol redox cycling by hydrazine combined with a low background current. Anal. Chem. 79, 2790–2796 (2007).
Das, J., Lee, J-A. & Yang, H. Ultrasensitive detection of DNA in diluted serum using NaBH4 electrooxidation mediated by [Ru(NH3)6)]3+ at indium–tin oxide electrodes. Langmuir 26, 6804–6808 (2010).
Xiang, Y., Xie, M., Bash, R., Chen, J. J. L. & Wang, J. Ultrasensitive label-free aptamer-based electronic detection. Angew. Chem. Int. Ed. 119, 9212–9214 (2007).
Plaxco, K. W. & Soh, H. T. Switch-based biosensors: a new approach towards real-time, in vivo molecular detection. Trends Biotechnol. 29, 1–5 (2011).
Lai, R. L. et al. Rapid, sequence-specific detection of unpurified PCR amplicons via a reusable, electrochemical sensor. Proc. Natl Acad. Sci. USA 103, 4017–4021 (2006).
Lapierre, M. A., O'Keefe, M. M., Taft, B. J. & Kelley, S. O. Electrocatalytic detection of pathogenic DNA sequences and antibiotic resistance markers. Anal. Chem. 75, 6327–6333 (2003).
Soleymani, L., Fang, Z., Sargent, E. H. & Kelley, S. O. Programming the detection limits of biosensors through controlled nanostructuring. Nature Nanotechnol. 4, 844–848 (2009).
Soleymani, L. et al. Nanostructuring of patterned microelectrodes to enhance the sensitivity of electrochemical nucleic acids detection. Angew. Chem. Int. Ed. 48, 8457–8460 (2009).
Yang, H. et al. Direct, electronic microRNA detection reveals differential expression profiles in 30 minutes. Angew. Chem. Int. Ed. 48, 8461–8464 (2009).
Fang, Z. et al. Direct profiling of cancer biomarkers in tumour tissue using a multiplexed nanostructured microelectrode integrated circuit. ACS Nano 3, 3207–3213 (2009).
Das, J. & Kelley, S. O. Protein detection using arrayed microsensor chips: tuning sensor footprint to achieve ultrasensitive readout of CA-125 in serum and whole blood. Anal. Chem. 83, 1167–1172 (2011).
Soleymani, L. et al. Hierarchical nanotextured microelectrodes overcome the molecular transport barrier to achieve rapid, direct bacterial detection. ACS Nano 5, 3360–3366 (2011).
Vasilyeva, E., Fang, Z., Minden, M., Sargent, E. H. & Kelley, S. O. Direct genetic analysis of ten cancer cells: tuning molecular probe design for efficient mRNA capture Angew. Chem. Int. Ed. 50, 4137–4141 (2011).
Su, L., Sankar, C. G., Sen, D. & Yu, H-Z. Kinetics of ion-exchange binding of redox metal cations to thiolate–DNA monolayers on gold. Anal. Chem. 76, 5953–5959 (2004).
Bernstein, J. A., Khodursky, A. B., Lin, P. H., Lin-Chao, S. & Cohen, S. N. Global analysis of mRNA decay and abundance in Escherichia coli at single-gene resolution using two-color fluorescent DNA microarrays Proc. Natl Acad. Sci. USA 99, 9697–9702 (2002).
Lam, B., Fang, Z., Sargent, E. H. & Kelley, S. O. Polymerase chain reaction-free, sample-to-answer bacterial detection in 30 minutes with integrated cell lysis. Anal. Chem. 84, 21–25 (2012).
Taft, B. J., O'Keefe, M., Fourkas, J. T. & Kelley, S. O. Engineering DNA-electrode connectivities: manipulation of linker length and structure. Anal. Chim. Acta 496, 81–91 (2003).
Acknowledgements
We acknowledge the Natural Sciences and Engineering Research Council (Discovery Grant to S.O.K.), Mitacs (Elevate Fellowship to J.D.) and the Defense Advanced Research Projects Agency (DXoD programme funding to S.O.K. and E.H.S.) for their support of this work.
Author information
Authors and Affiliations
Contributions
S.O.K. and J.D. conceived the experiments. J.D., K.B.C., A.Z. and P.L. performed the experiments. J.D., K.B.C., A.Z., P.L., E.H.S. and S.O.K. discussed the results. J.D., K.B.C., E.H.S. and S.O.K. co-wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 1059 kb)
Rights and permissions
About this article
Cite this article
Das, J., Cederquist, K., Zaragoza, A. et al. An ultrasensitive universal detector based on neutralizer displacement. Nature Chem 4, 642–648 (2012). https://doi.org/10.1038/nchem.1367
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1367
This article is cited by
-
DNA binding by the antimalarial compound artemisinin
Scientific Reports (2022)
-
A multiplexed, electrochemical interface for gene-circuit-based sensors
Nature Chemistry (2020)
-
Expanding detection windows for discriminating single nucleotide variants using rationally designed DNA equalizer probes
Nature Communications (2020)
-
Graphene/aptamer probes for small molecule detection: from in vitro test to in situ imaging
Microchimica Acta (2020)
-
Uniform small metal nanoparticles anchored on CeO2 nanorods driven by electroless chemical deposition
Rare Metals (2020)