Abstract
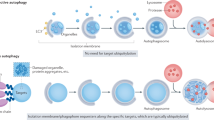
Autophagy is a conserved process that delivers components of the cytoplasm to lysosomes for degradation. The E1 and E2 enzymes encoded by Atg7 and Atg3 are thought to be essential for autophagy involving the ubiquitin-like protein Atg8. Here, we describe an Atg7- and Atg3-independent autophagy pathway that facilitates programmed reduction of cell size during intestine cell death. Although multiple components of the core autophagy pathways, including Atg8, are required for autophagy and cells to shrink in the midgut of the intestine, loss of either Atg7 or Atg3 function does not influence these cellular processes. Rather, Uba1, the E1 enzyme used in ubiquitylation, is required for autophagy and reduction of cell size. Our data reveal that distinct autophagy programs are used by different cells within an animal, and disclose an unappreciated role for ubiquitin activation in autophagy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Mizushima, N. & Komatsu, M. Autophagy: renovation of cells and tissues. Cell 147, 728–741 (2011).
Levine, B. & Yuan, J. Autophagy in cell death: an innocent convict? J. Clin. Invest. 115, 2679–2688 (2005).
Baehrecke, E. H. Autophagy: dual roles in life and death? Nat. Rev. Mol. Cell Biol. 6, 505–510 (2005).
Berry, D. L. & Baehrecke, E. H. Growth arrest and autophagy are required for salivary gland cell degradation in Drosophila. Cell 131, 1137–1148 (2007).
Hou, Y. C., Chittaranjan, S., Barbosa, S. G., McCall, K. & Gorski, S. M. Effector caspase Dcp-1 and IAP protein Bruce regulate starvation-induced autophagy during Drosophila melanogaster oogenesis. J. Cell Biol. 182, 1127–1139 (2008).
Nezis, I. P. et al. Autophagic degradation of dBruce controls DNA fragmentation in nurse cells during late Drosophila melanogaster oogenesis. J. Cell Biol. 190, 523–531 (2010).
Mizushima, N., Levine, B., Cuervo, A. M. & Klionsky, D. J. Autophagy fights disease through cellular self-digestion. Nature 451, 1069–1075 (2008).
Tsukada, M. & Ohsumi, Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 333, 169–174 (1993).
Thumm, M. et al. Isolation of autophagocytosis mutants of Saccharomyces cerevisiae. FEBS Lett. 349, 275–280 (1994).
Harding, T. M., Morano, K. A., Scott, S. V. & Klionsky, D. J. Isolation and characterization of yeast mutants in the cytoplasm to vacuole protein targeting pathway. J. Cell Biol. 131, 591–602 (1995).
Harding, T. M., Hefner-Gravink, A., Thumm, M. & Klionsky, D. J. Genetic and phenotypic overlap between autophagy and the cytoplasm to vacuole protein. J. Biol. Chem. 271, 17621–17624 (1996).
Schulman, B. A. & Harper, J. W. Ubiquitin-like protein activation by E1 enzymes: the apex for downstream signalling pathways. Nat. Rev. Mol. Cell Biol. 10, 319–331 (2009).
Denton, D. et al. Autophagy, not apoptosis, is essential for midgut cell death in Drosophila. Curr. Biol. 19, 1741–1746 (2009).
Lee, C-Y., Cooksey, B. A. K. & Baehrecke, E. H. Steroid regulation of midgut cell death during Drosophila development. Dev. Biol. 250, 101–111 (2002).
Micchelli, C. A., Sudmeier, L., Perrimon, N., Tang, S. & Beehler-Evans, R. Identification of adult midgut precursors in Drosophila. Gene Expr. Patter. 11, 12–21 (2011).
Scott, R. C., Juhász, G. & Neufeld, T. P. Direct induction of autophagy byAtg1 inhibits cell growth and induces apoptotic cell death. Curr. Biol. 17, 1–11 (2007).
Juhász, G., Erdi, B., Sass, M. & Neufeld, T. P. Atg7-dependent autophagy promotes neuronal health, stress tolerance, and longevity but is dispensable for metamorphosis in Drosophila. Genes Dev. 21, 3061–3066 (2007).
Belote, J. M. & Fortier, E. Targeted expression of dominant negative proteasome mutants in Drosophila melanogaster. Genesis 34, 80–82 (2002).
Pandey, U. B. et al. HDAC6 rescues neurodegeneration and provides an essential link between autophagy and the UPS. Nature 447, 859–863 (2007).
Fujita, N. et al. An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol. Biol. Cell 19, 4651–4659 (2008).
Pircs, K. et al. Advantages and limitations of different p62-based assays for estimating autophagic activity in Drosophila. PLoS ONE 7, e44214 (2012).
Jiang, C., Baehrecke, E. H. & Thummel, C. S. Steroid regulated programmedcell death during Drosophila metamorphosis. Development 124, 4673–4683 (1997).
Itakura, E. & Mizushima, N. p62 targeting to the autophagosome formation site requires self-oligomerization but not LC3 binding. J. Cell Biol. 192, 17–27 (2011).
Fujita, K., Maeda, D., Xiao, Q. & Srinivasula, S. M. Nrf2-mediated induction of p62 controls Toll-like receptor-4-driven aggresome-like induced structure formation and autophagic degradation. Proc. Natl Acad. Sci. USA 108, 1427–1432 (2011).
Lum, J. J. et al. Growth factor regulation of autophagy and cell survival in the absence of apoptosis. Cell 120, 237–248 (2005).
Hosokawa, N., Hara, Y. & Mizushima, N. Generation of cell lines with tetracycline-regulated autophagy and a role for autophagy in controlling cell size. FEBS Lett. 580, 2623–2629 (2006).
Nishida, Y. et al. Discovery of Atg5/Atg7-independent alternative macroautophagy. Nature 461, 654–658 (2009).
Tian, Y. et al. C. elegans screen identifies autophagy genes specific to multicellular organisms. Cell 141, 1042–1055 (2010).
McPhee, C. K., Logan, M. A., Freeman, M. R. & B., E. H. Activation of autophagy during cell death requires the engulfment receptor Draper. Nature 465, 1093–1096 (2010).
Johansen, T. & Lamark, T. Selective autophagy mediated by autophagic adapter proteins. Autophagy 7, 279–296 (2011).
Lenk, S. E., Dunn, W. A. J., Trausch, J. S., Ciechanover, A. & Schwartz, A. L. Ubiquitin-activating enzyme, E1, is associated with maturation of autophagic vacuoles. J. Cell Biol. 118, 301–308 (1992).
Taillebourg, E. et al. The deubiquitinating enzyme USP36 controls selective autophagy activation by ubiquitinated proteins. Autophagy 8, 767–779 (2012).
Scott, R. C., Schuldiner, O. & Neufeld, T. P. Role and regulation of starvation-induced autophagy in the Drosophila fat body. Dev. Cell 7, 167–178 (2004).
Chang, Y. Y. & Neufeld, T. P. An Atg1/Atg13 complex with multiple roles in TOR-mediated autophagy regulation. Mol. Biol. Cell 20, 2004–2014 (2009).
Shravage, B. V., Hill, J. H., Powers, C. M., Wu, L. & Baehrecke, E. H. Atg6 is required for multiple vesicle trafficking pathways and hematopoiesis in Drosophila. Development 140, 1321–1329 (2013).
Juhász, G. et al. The class III PI(3)K Vps34 promotes autophagy andendocytosis but not TOR signaling in Drosophila. J. Cell Biol. 181, 2347–2360 (2008).
Lee, T. V. et al. The E1 ubiquitin-activating enzyme Uba1 in Drosophila controls apoptosis autonomously and tissue growth non-autonomously. Development 135, 43–52 (2008).
Denton, D. et al. Relationship between growth arrest and autophagy inmidgut programmed cell death in Drosophila. Cell Death Differ. 19, 1299–1307 (2012).
Yamashita, S. The post-embedding method for immunoelectron microscopy of mammalian tissues: a standardized procedure based on heat-induced antigen retrieval. Methods Mol. Biol. 657, 237–248 (2010).
Dutta, S. & Baehrecke, E. H. Warts is required for PI3K-regulated growth arrest, autophagy, and autophagic cell death in Drosophila. Curr. Biol. 18, 1466–1475 (2008).
Acknowledgements
We thank A. Bergmann (University of Massachusetts Medical School, USA), M. Brodsky (University of Massachusetts Medical School, USA), S. Cherry (University of Pennsylvania, USA), G. Juhasz (Eotvos Lorand University, Hungary), P. Meier (The Institute of Cancer Research, UK), T. P. Neufeld (University of Minnesota, USA), H. D. Ryoo (New York University, USA), H. Stenmark (Oslo University Hospital, Norway), the Bloomington Stock Center, the VDRC and the Developmental Studies Hybridoma Bank for flies and antibodies, G. Juhasz for advice about immuno-electron microscopy, T. Fortier for technical support, and M. Freeman, S. Doxsey and the Baehrecke laboratory for constructive comments. This work was supported by NIH grants GM079431 to E.H.B., GM095567 to J.W.H., and S10RR027897 to the UMass EM Core, and by Millennium Pharamceuticals to J.W.H. E.H.B. is an Ellison Medical Foundation Scholar and a member of the UMass DERC (DK32520).
Author information
Authors and Affiliations
Contributions
T-K.C., B.V.S., S.D.H., J.W.H. and E.H.B. designed the experiments. All experiments were performed by T-K.C., except Fig. 4d,e by B.V.S., TEM by T-K.C., B.V.S., C.M.P. and R.T.S. and E1 charging assays by S.D.H. T-K.C., B.V.S. and E.H.B. wrote the manuscript and all authors commented on it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Knockdown of Atg1 does not alter cell growth, and Atg13, Atg6, Atg5, and Atg16 (CG31033), but not caspases, are required for midgut cell size reduction in Drosophila.
(a) Midguts dissected from early third instar larvae that mis-express Atg1IR (GFP in nucleus and cytoplasm) and are stained to detect Discs large (green) on the cortex of all cells. Representative images are shown. (b) Cell size quantification (μm2) from a, n = 10 animal intestines/genotype with 1-5 cells measured/intestine. (c) Midguts dissected from animals at puparium formation that contain Atg13Δ74 loss-of-function mutant cell clones (lacking GFP) and analyzed by fluorescence and DIC microscopy. Wild-type (+/+) control cells possess stronger GFP and heterozygous Atg13Δ74/wild-type (Δ74/+) cells have weaker GFP. Representative images are shown. (d) Cell size quantification (μm2) from c, nn = 6 animal intestines/genotype with 1-5 cells measured/intestine. (e) Midguts dissected from animals at puparium formation that contain Atg61 loss-of-function mutant cell clones (lacking RFP) and analyzed by fluorescence and DIC microscopy. Wild-type (+/+) control cells possess stronger RFP and heterozygous Atg61/wild-type (1/+) cells have weaker RFP. Representative images are shown. (f) Cell size quantification (μm2) from e, n = 8 animal intestines/genotype with 1-5 cells measured/intestine. (g) Midguts dissected from animals expressing Atg5IR specifically in GFP-marked clones of cells at puparium formation and analyzed by fluorescence and DIC microscopy. Representative images are shown. (h) Cell size quantification (μm2) from g, n = 16 animal intestines/genotype with 1-5 cells measured/intestine. (i) Midguts dissected from animals expressing Atg16IR specifically in DsRed-marked clones of cells at puparium formation and analyzed by fluorescence and DIC microscopy. Representative images are shown. (j) Cell size quantification (μm2) from i, n = 7 animal intestines/genotype with 1–5 cells measured/intestine. (k) Midguts dissected from animals expressing p35 in DsRed-marked clones of cells at puparium formation and analyzed by fluorescence and DIC microscopy. Representative images are shown. (l) Cell size quantification (μm2) from k, n = 13 animal intestines/genotype with 1-5 cells measured/intestine. Quantification is shown as mean ±s.d.. N.S.: no significance. Scale bars represent 20 μm.
Supplementary Figure 2 Atg7 is required for nutrient deprivation-induced autophagy in starved larval fat body and Vps34 is required for GFP–Atg5 puncta formation in the midgut at puparium formation.
(a,b) Fat body dissected from early 3rd instar larvae and subjected to 4 h of starvation. Representative images are shown. (a) Fat body expressing mCherry-Atg8a in all cells, and Atg7IR in GFP-marked clones of cells analyzed by fluorescence microscopy. (b) Atg7d4 loss-of-function mutant cells are shown lacking of myrRFP. Wild-type and heterozygous Atg7d4/wild-type control cells possess myrRFP. All fat body cells express GFP–Atg8a. (c) Midguts dissected from animals expressing Atg7IR specifically in DsRed-marked clones of cells at puparium formation and analyzed by fluorescence and DIC microscopy. Representative images are shown. (d) Cell size quantification (μm2) from c, n = 10 animal intestines/genotype with 1-5 cells measured/intestine. (e) Midguts expressing GFP–Atg5 in enterocytes (larger nuclei) and with Vps34Δm22 loss-of-function clone cells (lacking RFP) at puparium formation. Wild-type control cells possess stronger RFP and heterozygous cells have weaker RFP. Representative images are shown. Quantification is shown as mean ±s.d.. N.S.: no significance. Scale bars represent 20 μm.
Supplementary Figure 3 Screen for ubiquitin-like activating enzyme genes that function in midgut programmed cell reduction in size in Drosophila.
Representative images of midgut cells from animals expressing RNAi in GFP-marked clones of cells against (a) Uba2 (CG7528-1, n = 11, VDRC TID 110173, n = 14, and TRiP HM05055, n = 9), (b) Aos1 (TID 47256, n = 8, and HM05183, n = 9), (c) CG13343 (CG13343-3, n = 12, CG13343-4, n = 14, and TID 105141, n = 8), (d) APP-BP1 (TID 7728, n = 13), (e) CG13090 (TID 110326, n = 13, and TID 43558, n = 10), and (f) CG1749 (TID 110395, n = 14, and HMS01352, n = 12). n represents number of animal intestines analyzed/genotype. Scale bars represent 20 μm.
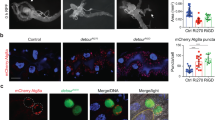
Supplementary Figure 4 Uba1 influences autophagy and programmed reduction of midgut cell size.
(a) Midguts expressing mCherry-Atg8a in all cells, and expressing Uba1IR specifically in GFP-marked clones of cells at puparium formation. Representative images are shown. (b) Midguts dissected from early third instar larvae that mis-express Uba1IR (GFP in nucleus and cytoplasm) and were stained to detect Discs large (green) on the cortex of all cells. Representative images are shown. (c) Cell size quantification (μm2) from b, n = 10 animal intestines/genotype with 1-5 cells measured/intestine. (d) Midguts from animals expressing CL1-GFP in all cells and temperature sensitive Dts7 specifically in DsRed-marked clones of cells at puparium formation. Representative images are shown. Quantification is shown as mean ±s.d.. N.S.: no significance. Scale bars represent 20 μm.
Supplementary Figure 5 Uba1 is not required for autophagy that is induced by stress in different tissues.
(a) Fat body dissected from early third instar larvae and subjected to 4 h of starvation that contain Uba1H33 loss-of-function MARCM mutant cell clones (GFP-positive) that also have mCherry-Atg8a expressed in all cells and analyzed by fluorescence microscopy. Control wild-type and heterozygous cells have no GFP. Representative images are shown. (b-f) Early third instar midguts expressing mCherry-Atg8a in all cells and expressing either Uba1IR (b,d,f) or Atg1IR (c,e) in GFP-marked clones of cells in control untreated animals (b), 4 h after starvation (c,d), or 7 h after 1.5 % H2O2 treatment (e,f). Representative images are shown. Scale bars represent 20 μm.
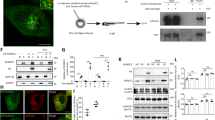
Supplementary Figure 6 Uba1, but not loss of Atg3, influences autophagy and programmed reduction of midgut cell size.
(a,b) Loading controls for the E1 charging assays, n = 2. Representative images are shown. (c,d) Representative TEM images of intestine cells 2 h after puparium formation. (c) Atg310/wild-type control (Atg3/+) and (d) Atg310/Df(3L)cat mutant (Atg3/Df) cells both possess autophagic structures. (e) Starved third instar larval fat body expressing mCherry-Atg8a in all cells and Atg3IR in GFP-marked clones of cells and analyzed with fluorescence microscopy. Representative images are shown. (f,g) Midguts dissected from animals at puparium formation that mis-express Uba1IR (GFP in nucleus and cytoplasm) and stained to detect either BrC (f, red) or Hid (g, red). Representative images are shown. Scale bars represent 1 μm (c,d) and 20 μm (e-g).
Supplementary Figure 7 Full scans of blots in Fig. 7a (a), Fig. 7b (b), Supplementary Fig. S6a (c), Supplementary Fig. S6b (d), and Fig. 7c (e,f).
Supplementary information
Supplementary Information
Supplementary Information (PDF 1719 kb)
Rights and permissions
About this article
Cite this article
Chang, TK., Shravage, B., Hayes, S. et al. Uba1 functions in Atg7- and Atg3-independent autophagy. Nat Cell Biol 15, 1067–1078 (2013). https://doi.org/10.1038/ncb2804
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2804
This article is cited by
-
GTP energy dependence of endocytosis and autophagy in the aging brain and Alzheimer’s disease
GeroScience (2023)
-
The evolutionary and functional divergence of the Atg8 autophagy protein superfamily
Biologia Futura (2022)
-
Ferulic acid alleviates lipotoxicity-induced hepatocellular death through the SIRT1-regulated autophagy pathway and independently of AMPK and Akt in AML-12 hepatocytes
Nutrition & Metabolism (2021)
-
Acetoacetate protects macrophages from lactic acidosis-induced mitochondrial dysfunction by metabolic reprograming
Nature Communications (2021)
-
Cadmium Impairs Autophagy Leading to Apoptosis by Ca2+-Dependent Activation of JNK Signaling Pathway in Neuronal Cells
Neurochemical Research (2021)