Abstract
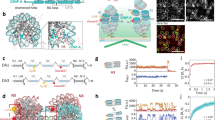
Epigenetic mechanisms regulate genome activation in diverse events, including normal development and cancerous transformation. Centromeres are epigenetically designated chromosomal regions that maintain genomic stability by directing chromosome segregation during cell division. The histone H3 variant CENP-A resides specifically at centromeres, is fundamental to centromere function and is thought to act as the epigenetic mark defining centromere loci. Mechanisms directing assembly of CENP-A nucleosomes have recently emerged, but how CENP-A is maintained after assembly is unknown. Here, we show that a small GTPase switch functions to maintain newly assembled CENP-A nucleosomes. Using functional proteomics, we found that MgcRacGAP (a Rho family GTPase activating protein) interacts with the CENP-A licensing factor HsKNL2. High-resolution live-cell imaging assays, designed in this study, demonstrated that MgcRacGAP, the Rho family guanine nucleotide exchange factor (GEF) Ect2, and the small GTPases Cdc42 and Rac, are required for stability of newly incorporated CENP-A at centromeres. Thus, a small GTPase switch ensures epigenetic centromere maintenance after loading of new CENP-A.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Goldberg, A. D., Allis, C. D. & Bernstein, E. Epigenetics: a landscape takes shape. Cell 128, 635–638 (2007).
Karpen, G. H. & Allshire, R. C. The case for epigenetic effects on centromere identity and function. Trends Genet. 13, 489–496 (1997).
Palmer, D. K., O'Day, K., Trong, H. L., Charbonneau, H. & Margolis, R. L. Purification of the centromere-specific protein CENP-A and demonstration that it is a distinctive histone. Proc. Natl Acad. Sci. USA 88, 3734–3738 (1991).
Choo, K. H. Centromerization. Trends Cell Biol. 10, 182–188 (2000).
Bernad, R., Sanchez, P. & Losada, A. Epigenetic specification of centromeres by CENP-A. Exp. Cell Res. 315, 3233–3241 (2009).
Sullivan, K. F., Hechenberger, M. & Masri, K. Human CENP-A contains a histone H3 related histone fold domain that is required for targeting to the centromere. J. Cell Biol. 127, 581–592 (1994).
Cleveland, D. W., Mao, Y. & Sullivan, K. F. Centromeres and kinetochores: from epigenetics to mitotic checkpoint signaling. Cell 112, 407–421 (2003).
Schuh, M., Lehner, C. F. & Heidmann, S. Incorporation of Drosophila CID/CENP-A and CENP-C into centromeres during early embryonic anaphase. Curr. Biol. 17, 237–243 (2007).
Jansen, L. E., Black, B. E., Foltz, D. R. & Cleveland, D. W. Propagation of centromeric chromatin requires exit from mitosis. J. Cell Biol. 176, 795–805 (2007).
Hayashi, T. et al. Mis16 and Mis18 are required for CENP-A loading and histone deacetylation at centromeres. Cell 118, 715–729 (2004).
Fujita, Y. et al. Priming of centromere for CENP-A recruitment by human hMis18α, hMis18β, and M18BP1. Dev. Cell 12, 17–30 (2007).
Maddox, P. S., Hyndman, F., Monen, J., Oegema, K. & Desai, A. Functional genomics identifies a Myb domain-containing protein family required for assembly of CENP-A chromatin. J. Cell Biol. 176, 757–763 (2007).
Foltz, D. R. et al. Centromere-specific assembly of CENP-A nucleosomes is mediated by HJURP. Cell 137, 472–484 (2009).
Dunleavy, E. M. et al. HJURP is a cell-cycle-dependent maintenance and deposition factor of CENP-A at centromeres. Cell 137, 485–497 (2009).
Silva, M. C. & Jansen, L. E. At the right place at the right time: novel CENP-A binding proteins shed light on centromere assembly. Chromosoma 118, 567–574 (2009).
Perpelescu, M., Nozaki, N., Obuse, C., Yang, H. & Yoda, K. Active establishment of centromeric CENP-A chromatin by RSF complex. J. Cell Biol. 185, 397–407 (2009).
Etienne-Manneville, S. & Hall, A. Rho GTPases in cell biology. Nature 420, 629–635 (2002).
Izuta, H. et al. Comprehensive analysis of the ICEN (interphase centromere complex) components enriched in the CENP-A chromatin of human cells. Genes Cells 11, 673–684 (2006).
Kawashima, T. et al. MgcRacGAP is involved in the control of growth and differentiation of hematopoietic cells. Blood 96, 2116–2124 (2000).
Minoshima, Y. et al. Phosphorylation by aurora B converts MgcRacGAP to a RhoGAP during cytokinesis. Dev. Cell 4, 549–560 (2003).
Toure, A. et al. MgcRacGAP, a new human GTPase-activating protein for Rac and Cdc42 similar to Drosophila rotundRacGAP gene product, is expressed in male germ cells. J. Biol. Chem. 273, 6019–6023 (1998).
Yuce, O., Piekny, A. & Glotzer, M. An ECT2-centralspindlin complex regulates the localization and function of RhoA. J. Cell Biol. 170, 571–582 (2005).
Canman, J. C. et al. Inhibition of Rac by the GAP activity of centralspindlin is essential for cytokinesis. Science 322, 1543–1546 (2008).
Miller, A. L. & Bement, W. M. Regulation of cytokinesis by Rho GTPase flux. Nat. Cell Biol. 11, 71–77 (2009).
Hirose, K., Kawashima, T., Iwamoto, I., Nosaka, T. & Kitamura, T. MgcRacGAP is involved in cytokinesis through associating with mitotic spindle and midbody. J. Biol. Chem. 276, 5821–5828 (2001).
Jantsch-Plunger, V. et al. CYK-4: A Rho family gtpase activating protein (GAP) required for central spindle formation and cytokinesis. J. Cell Biol. 149, 1391–1404 (2000).
Pavicic-Kaltenbrunner, V., Mishima, M. & Glotzer, M. Cooperative assembly of CYK-4/MgcRacGAP and ZEN-4/MKLP1 to form the centralspindlin complex. Mol. Biol. Cell 18, 4992–5003 (2007).
Maddox, A. S. & Oegema, K. Closing the GAP: a role for a RhoA GAP in cytokinesis. Mol. Cell 11, 846–848 (2003).
Black, B. E. et al. Centromere identity maintained by nucleosomes assembled with histone H3 containing the CENP-A targeting domain. Mol. Cell 25, 309–322 (2007).
Foltz, D. R. et al. The human CENP-A centromeric nucleosome-associated complex. Nat. Cell Biol. 8, 458–469 (2006).
von Dassow, G., Verbrugghe, K. J., Miller, A. L., Sider, J. R. & Bement, W. M. Action at a distance during cytokinesis. J. Cell Biol. 187, 831–845 (2009).
Yasuda, S. et al. Cdc42 and mDia3 regulate microtubule attachment to kinetochores. Nature 428, 767–771 (2004).
Oceguera-Yanez, F. et al. Ect2 and MgcRacGAP regulate the activation and function of Cdc42 in mitosis. J. Cell Biol. 168, 221–232 (2005).
Sanders, L. C., Matsumura, F., Bokoch, G. M. & de Lanerolle, P. Inhibition of myosin light chain kinase by p21-activated kinase. Science 283, 2083–2085 (1999).
Rando, O. J., Zhao, K. & Crabtree, G. R. Searching for a function for nuclear actin. Trends Cell Biol. 10, 92–97 (2000).
Glotzer, M. The molecular requirements for cytokinesis. Science 307, 1735–1739 (2005).
Maddox, A. S. & Oegema, K. Deconstructing cytokinesis. Nat. Cell Biol. 5, 773–776 (2003).
Field, C., Li, R. & Oegema, K. Cytokinesis in eukaryotes: a mechanistic comparison. Curr. Opin. Cell Biol. 11, 68–80 (1999).
Chen, M. & Shen, X. Nuclear actin and actin-related proteins in chromatin dynamics. Curr. Opin. Cell Biol. 19, 326–330 (2007).
Tomonaga, T. et al. Overexpression and mistargeting of centromere protein-A in human primary colorectal cancer. Cancer Res. 63, 3511–3516 (2003).
Collins, K. A., Furuyama, S. & Biggins, S. Proteolysis contributes to the exclusive centromere localization of the yeast Cse4/CENP-A histone H3 variant. Curr. Biol. 14, 1968–1972 (2004).
Ye, J., Zhao, J., Hoffmann-Rohrer, U. & Grummt, I. Nuclear myosin I acts in concert with polymeric actin to drive RNA polymerase I transcription. Genes Dev. 22, 322–330 (2008).
Theriot, J. A., Mitchison, T. J., Tilney, L. G. & Portnoy, D. A. The rate of actin-based motility of intracellular Listeria monocytogenes equals the rate of actin polymerization. Nature 357, 257–260 (1992).
Stelzer, E. H., Wacker, I. & De Mey, J. R. Confocal fluorescence microscopy in modern cell biology. Semin. Cell Biol. 2, 145–152 (1991).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Transactions Syst. Man Cybern. 9, 62–66 (1979).
Lindeberg, T. & Eklundh, J. O. Scale detection and region extraction from a scale-space primal sketch. Proc. 3rd International Conference on Computer Vision (Osaka, Japan) 416–426 (1990).
Dorn, J. F., Danuser, G. & Yang, G. Computational processing and analysis of dynamic fluorescence image data. Methods Cell Biol. 85, 497–538 (2008).
Jaqaman, K. et al. Kinetochore alignment within the metaphase plate is regulated by centromere stiffness and microtubule depolymerases. J. Cell Biol. 188, 665–679 (2010).
Dorn, J. F. et al. Yeast kinetochore microtubule dynamics analyzed by high-resolution three-dimensional microscopy. Biophys. J. 89, 2835–2854 (2005).
Carroll, C. W., Silva, M. C., Godek, K. M., Jansen, L. E. & Straight, A. F. Centromere assembly requires the direct recognition of CENP-A nucleosomes by CENP-N. Nat. Cell Biol. 11, 896–902 (2009).
Cheeseman, I. M. & Desai, A. A combined approach for the localization and tandem affinity purification of protein complexes from metazoans. Sci STKE 2005, pl1 (2005).
Cheeseman, I. M. et al. A conserved protein network controls assembly of the outer kinetochore and its ability to sustain tension. Genes Dev. 18, 2255–2268 (2004).
Acknowledgements
The authors would like to thank J.-C. Labbé, A. Desai, K. Bloom and S. Meloche for helpful comments and discussions, the IRIC proteomics facility (specifically É. Bonneil and P. Thibault) for LC–MS/MS analysis, the IRIC genomics facility for sequencing and qPCR analyses, and A. Straight and B. Moree for the generous gift of the CENP-A SNAP-tag cell line. J.F.D. was supported by a postdoctoral fellowship from the Swiss National Science Foundation; V.D.R. was supported by a student fellowship from the CIHR. A.S.M. receives salary support from the FRSQ. P.S.M. holds the Canada Research Chair in Chromosome Organization and Mitotic Mechanisms. This work was funded by grants #018450 awarded to P.S.M. and #019162 awarded to A.S.M. from the Terry Fox Foundation and the CCSRI. V.D.R., A.S.M. and P.S.M. participate in the Systems biology option in the graduate program in Molecular Biology, Université de Montréal.
Author information
Authors and Affiliations
Contributions
A.L. conducted all immunoprecipitation and mass spectrometry experiments. A.L. and J.F.D. performed shRNA experiments and analysis, respectively. J.F.D. performed live-cell experiments. A.L. evaluated all shRNA experiments (qPCR, western blot). V.D.R. performed the small GTPases localization experiments. All authors performed essential tasks in generating figures. P.S.M., A.L., J.D., A.-M.L., and A.S.M. conceived the experiments. P.S.M. wrote the manuscript, assisted by all authors, in particular A.S.M. and J.F.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 986 kb)
Supplementary Movie 1
Supplementary Information (MOV 1642 kb)
Supplementary Movie 2
Supplementary Information (MOV 235 kb)
Rights and permissions
About this article
Cite this article
Lagana, A., Dorn, J., De Rop, V. et al. A small GTPase molecular switch regulates epigenetic centromere maintenance by stabilizing newly incorporated CENP-A. Nat Cell Biol 12, 1186–1193 (2010). https://doi.org/10.1038/ncb2129
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2129