Abstract
Wild relatives of crops are an important source of genetic diversity for agriculture, but their gene repertoire remains largely unexplored. We report the establishment and analysis of a pan-genome of Glycine soja, the wild relative of cultivated soybean Glycine max, by sequencing and de novo assembly of seven phylogenetically and geographically representative accessions. Intergenomic comparisons identified lineage-specific genes and genes with copy number variation or large-effect mutations, some of which show evidence of positive selection and may contribute to variation of agronomic traits such as biotic resistance, seed composition, flowering and maturity time, organ size and final biomass. Approximately 80% of the pan-genome was present in all seven accessions (core), whereas the rest was dispensable and exhibited greater variation than the core genome, perhaps reflecting a role in adaptation to diverse environments. This work will facilitate the harnessing of untapped genetic diversity from wild soybean for enhancement of elite cultivars.
Similar content being viewed by others
Main
Annual wild soybean (Glycine soja Sieb. & Zucc.) is the closest relative and antecedent of cultivated soybean (Glycine max (L.) Merr.), one of the world's primary sources of plant protein and vegetable oil. Whereas G. max has lost substantial genetic diversity through successive bottlenecks owing to domestication and selection for traits to increase yield under intensive human cultivation, G. soja is distributed across a broad geographical range (24–53° N, 97–143° E) and has adapted to a variety of ecological conditions1, providing a promising source of novel genes for soybean improvement needed to respond to rapid population growth and environmental changes.
Since the publication of the soybean genome (var. Williams 82 (GmaxW82))2, attempts have been made to characterize G. soja through genome resequencing3,4,5. However, resequencing is limited in terms of capturing many types of structural variation, especially for crop wild relatives, which are more genetically diverse than their domesticated counterparts. High levels of sequence similarity are required to map resequencing short reads to a reference genome at the expense of losing information from more diverged genomic regions. Presence-absence variation (PAV) and copy number variation (CNV), which are often associated with agronomic traits6, may also be missed7. Moreover, a single genome is insufficient to represent the genomic content of a predominantly selfing (autogamous) species, such as G. soja, in which individuals are distinct from one another due to low levels of genetic exchange and recombination8. Thus, the de novo construction of a pan-genome for a species, consisting of a core genome shared among individuals and individual-specific or partially shared dispensable genome, is necessary to capture the majority of genetic diversity within a species9.
To fully characterize the genomic content and molecular evolutionary history of G. soja, the undomesticated gene pool of soybean8, we constructed a pan-genome by sequencing and assembling seven G. soja accessions de novo. Within this pan-genome, the dispensable gene set was found to have evolved more rapidly and be more variable than the core gene set, and genes under selection exhibited relatively little overlap between the seven G. soja lineages, indicating that local adaptation may have affected nonoverlapping sets of genes. Some of the genes with structural variation between G. soja and GmaxW82, based on homology and co-localization with mapped quantitative trait loci (QTLs), may be associated with adaptation to various abiotic and biotic stresses. Our data also push the divergence between G. max and G. soja back to ∼0.8 million years ago (mya), from a recent estimate of ∼0.27 mya4. This work illustrates the value of de novo assemblies in building and characterizing large eukaryotic pan-genomes, and provides candidate genes for introgression into cultivated soybean for crop improvement.
Results
De novo genome assemblies of seven G. soja accessions
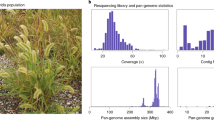
Seven G. soja accessions, representing the geographical adaptation within the species, were selected and named GsojaA to GsojaG (Supplementary Figs. 1 and 2 and Supplementary Table 1). Six accessions represent major genetically distinct clusters (North, Huanghuai and South regions of China, and Japan, Korea and Russia) of G. soja and 87.1% of the genetic diversity8. Another was selected from the Northeast region (NER) of China, a predicted soybean domestication center10, to better represent the geographic distribution of G. soja. Each genome was sequenced with Illumina HiSeq2000 using a combination of libraries with insert sizes of 180 bp, 500 bp and 2 Kbp (Supplementary Table 2), for an average of 111.9 fold coverage (Table 1). The estimated genome sizes ranged from 889.33 Mbp for GsojaG (93.6% of the GmaxW82) to 1,118.34 Mbp for GsojaD (117.7% of GmaxW82). Variation in genome size could be partially explained by differences in abundance of repetitive sequences (Supplementary Fig. 3).
Each genome was assembled using SOAPdenovo11, a de novo genome assembler based on a de Bruijn graph algorithm, and resulted in contig N50 sizes ranging from 8 to 27 Kbp and scaffold N50 sizes ranging from 17 to 65.1 Kbp (Table 1). The assemblies were validated by alignment to GmaxW82, covering more than 94% of the 54,175 genes (Glyma1.1 annotation, http://www.phytozome.net, Table 1). The coverage of GmaxW82 genes by assembled genomes was either higher than (5 G. soja genomes) or comparable to (2 G. soja genomes) the coverage by aligning resequencing reads. As such, the current assemblies enabled accurate detection of variation and comparative analyses within genic regions. Additional mate-pair libraries with large inserts would further improve assembly size, thereby resolving variation in nongenic regions.
Protein-coding genes were annotated for each genome by integrating homology searches, mRNA expression evidence, and ab initio prediction (Supplementary Table 3). After correction for gene fragments due to incomplete assembly, we estimated an average of 55,570 genes per genome (Table 1), slightly more than the 54,175 genes in GmaxW82. The quality of the annotation was supported by the observation that 89.24–91.93% of genes had at least one ortholog in the GmaxW82 genome and 63.94–71.19% of the predicted genes were expressed, as determined by RNA-seq of mixed tissues of corresponding accessions. In addition, a majority (96.15–97.16%) had protein homologs in other plant genomes (Supplementary Table 4). Underestimation of gene number may result from lack of evidence to support prediction. More likely, however, the gene number may be inflated as it includes genes split across contigs and genes on separately assembled haplotypes.
Variation between G. soja and GmaxW82
To identify variation between G. soja genomes and GmaxW82, we designed a computational pipeline that takes advantage of the sequencing reads as well as assembled genomes to catalog variation including single-nucleotide polymorphisms (SNPs), insertions or deletions (indels), CNV and PAV (Supplementary Fig. 4). There were 3.63–4.72 million SNPs in individual samples, with 0.12–0.15 million in coding sequence regions (CDS). This included 1,764 loci where G. soja had a stop codon and GmaxW82 did not, and 2,286 loci where GmaxW82 had a stop codon and G. soja did not (Supplementary Table 5). The assembly-based method enabled us to find more SNPs than by resequencing alone, especially in divergent regions where unassembled short sequencing reads are difficult to be mapped (Fig. 1a and Supplementary Figs. 5 and 6). Even though G. soja is a predominantly selfing species, three of the seven (GsojaA, GsojaC and GsojaD) had relatively high heterozygous SNP rates (Supplementary Table 5). The high rate of heterozygosity may have complicated de novo assembly and resulted in inflated gene numbers due to divergent haplotypes being represented as separate scaffolds in the assembly.
(a) A diverged region on chromosome 15. Top panel, coverage of GmaxW82 (blue) and GsojaA genome (red) by GsojaA reads. Comparison between GmaxW82 and G. soja sequences show nine SNPs in a 62-bp fragment at bottom. This highly diverged region could not be aligned by mapping GsojaA reads to GmaxW82. (b) Indels in Glyma02g25230. GmaxW82 gene structure is shown at top (exons in blue) and five amino acid changes caused by three indels in G. soja are shown in red at bottom. (c) G. soja–specific PAV. Syntenic genomic region on chromosome 14 (Block A) compared to its homolog on chromosome 02 (Block B). Genes in each block are numbered A1 to A7 and B1 to B8, respectively. An 8-Kbp G. soja–specific region is outlined in red, in which the three genes absent in the G. max region but with homologs on block A are in yellow, and the two genes with no homologs either on block A or elsewhere in G. max are in purple. Genes exhibiting relatively conserved synteny between species are marked in blue. The anchor point of each block is shown in black. Homologous genes are connected by magenta shading and the anchor points of each block are connected by gray shading. A phylogenetic tree was built using genes in red (Glyma02g38310). (d) G. max–specific PAV. Depth distribution of G. soja genomes on GmaxW82 chromosome 15 (46–46.5 Mbp) is shown with y axis representing the depth ratio on top and GC content below. The sequence depth ratio was calculated using the average depth in 10-Kbp windows for each G. soja accession over the genome-wide average. Seven of the 19 genes with low read depth in G. soja accessions were identified as G. max–specific genes, for example, Glyma15g39573. (e) Distribution of genetic variation across the soybean genome. A, chromosome ideograms for G. max, pericentromeric regions represented in gray; B, gene density; C, distribution of genes containing large-effect mutations; D, CNV (red for gain, green for loss); and E, G. max–specific PAV. For circle C, D and E, numbers represent percentages of genes with corresponding mutations over total genes in 1-Mbp window.
We detected 0.50–0.77 million indels in G. soja as compared to GmaxW82, of which 93.3% (70/75) of randomly selected indel alleles were validated by Sanger sequencing (Supplementary Table 6 and Supplementary Figs. 7 and 8). Although many indels were trinucleotides, 2,989–4,181 resulted in frameshifts (Supplementary Fig. 9 and Supplementary Tables 7 and 8). For example, three indels were identified in one of the two homologs of Spiral2 (spr2) (AT4G27060), a key microtubule gene for directional cell elongation that is associated with the right-handed helical growth in Arabidopsis12 (Fig. 1b). In Glyma02g25230, the Spiral2-homolog, three guanines were deleted at positions 695, 707 and 714 in all seven G. soja genomes, resulting in five amino acid changes in one of the HEAT-repeat motifs. Thus, Glyma02g25230 is a candidate gene that may be responsible for the change from the twining growth habit found in G. soja to erect growth found in G. max (Supplementary Fig. 10).
We identified CNV in genic regions and found hundreds of genes that had either gained or lost copies in individual G. soja accessions compared to GmaxW82. Of 1,978 genes affected in the G. soja accessions, 1,179 had CNV loss, 726 had CNV gain and 73 had both CNV loss and gain (Supplementary Tables 9 and 10). Gene Ontology (GO) analysis indicated that genes related to abiotic and biotic stress tolerance, such as resistance (R)-genes with nucleotide-binding site (NBS) or NBS-leucine-rich repeat (LRR) domains and transcription factors, were significantly (P < 0.01, chi-squared test) enriched in genes affected by CNV. This indicates that these genes are evolutionarily labile and may be involved in adaptation to local environments and/or interaction with pathogens.
In total, 2.3–3.9 Mbp of G. soja–specific PAV (defined as >100 bp and <95% identity) was present in the G. soja genomes (Supplementary Table 11), less than the 8.3 Mbp reported for a single previously resequenced G. soja genome4. Our lower PAV discovery rate may be due to the use of more stringent filtering criteria, including the removal of unassembled sequences in G. max and putative microbial sequences. There were 338 genes found to have at least 50% of their coding sequences composed of G. soja–specific sequences (Supplementary Table 12 and Supplementary Fig. 11). Of these genes or genomic loci affected by putative PAV, 37 were selected for PCR-based genotyping across GmaxW82 and the seven G. soja accessions (using 43 primer pairs). All 37 loci were concordant with the predicted PAV that was present in G. soja and absent in G. max (Supplementary Tables 13 and 14 and Supplementary Fig. 12). Of the G. soja–specific PAV genes, 34 were inferred to be involved in defense response, 33 in cell growth and 15 in photosynthesis, by GO category enrichment analysis (Supplementary Table 15).
As an example of genic PAV, an 8-Kbp G. soja–specific region was identified in all seven accessions (Block A) on chromosome 14 and validated by PCR genotyping (Supplementary Fig. 13). A paralogous region (Block B) on chromosome 02 in G. soja and G. max contained three genes missing in Block A that were related to biotic and abiotic stress tolerance or plant development (Glyma02g38290, homolog of ZRZ/ZF14 or BCD1; Glyma02g38300, homolog of SUT4 and Glyma02g38310, homolog of GLY1)13,14 (Fig. 1c). Of the 56 resequenced Glycine accessions3,4,5, 17 of 27 G. soja accessions and three of 29 G. max accessions contained this putative G. soja–specific sequence, indicating that this PAV is segregating in both G. soja and G. max but the frequency has been reduced in G. max, perhaps due to the domestication bottleneck.
The total length of G. max–specific PAV (1.86 Mbp) was less than that of G. soja–specific PAV. The GC content for these regions (32.9%) was not significantly different from the whole genome average (34.1%), rejecting the possibility of sequencing bias. A total of 16 genes were identified to contain G. max–specific sequences (>50% of the length of the CDS affected), including four genes that were entirely G. max–specific (Fig. 1d). This number was less than the 712 found in an earlier comparison of a single G. soja accession and GmaxW82 (ref. 4), which may be due to increased sampling of G. soja diversity and more stringent criteria. Based on KEGG analysis of the four G. max–specific genes, Glyma01g37051 is involved in energy metabolism (ko00190 and ko00194), Glyma13g36351 and Glyma15g39541 in lipid metabolism (ko00590, ko00591 and ko00592), and Glyma15g39596 in cell growth and death (ko04210). Glyma01g37051 is a homolog of AtLFNR1 (ferredoxin-NADP+-oxidoreductase, AT5G66190), which in Arabidopsis is involved in photosystem I–dependent cyclic electron now and affects rosette size15.
Variation was distributed unevenly throughout the genome (Fig. 1e). Genes in recombination-suppressed pericentromeric regions were more significantly (P < 0.01, chi-squared test) affected than those in chromosome arms (Supplementary Table 16), as was also observed in Zea mays (maize)16. The majority of genes affected by large-effect variations (including SNPs or indels causing stop codon gain or loss and frameshift), CNV or genome-specific sequences were rare events usually found in only one of the seven G. soja accessions. These rare variants may be useful in improving soybean cultivars through breeding, as has been indicated by studies in maize and wheat, in which rare alleles were found to be associated with QTLs for agronomic traits17.
Evolution of the G. max and G. soja species complex
We next constructed a phylogenetic tree using 670 conserved, single-copy, gene orthologs from G. soja, G. max and three other sequenced legume species (Fig. 2). Using the divergence time between Medicago and Glycine as a calibration point18, we estimated that pigeon pea and soybean diverged ∼23 mya, and common bean and soybean ∼29.5 mya, similar to previous estimates18,19. We also estimated the sequenced G. soja accessions and G. max diverged ∼0.8 mya, earlier than previously estimated (0.27 mya) based on a single G. soja accession4. This difference is likely due to the inclusion of more diverged G. soja accessions in the current study.
The divergence time between G. soja and G. max predates the estimated domestication time of soybean of ∼5,500 ya20, therefore divergent selection may have contributed to the differentiation of the two subspecies before domestication of G. max. Using the branch-site likelihood ratio test, we identified 682 genes that showed evidence of positive selection in the GmaxW82 genome (Supplementary Table 17). These genes were enriched in diverse biological functional categories related to abiotic stress regulation, including stomatal complex development, voltage-gated potassium channel activity, proline metabolism and nitrate transport (Supplementary Table 18). In the G. soja accessions, 175–408 such genes were identified in individual wild soybean accessions and none was shared among all accessions. Only ten positive-selection genes were shared by at least three G. soja accessions and 132 such genes were shared by two accessions. Genes that underwent positive selection may have contributed to the adaptation of G. soja accessions to different environments, as evidenced by the species' wide range of geographical distribution and lack of shared positive-selection genes.
Changes within the G. soja pan-genome (total gene set) and core genome (genes shared among all seven genomes) were analyzed (Supplementary Fig. 14). The number of total genes increased as additional genomes were added and, in contrast, the number of shared genes decreased with additional G. soja genomes (Fig. 3a). The average pan-genome size of any two accessions accounted for 78.2% of that found using all seven accessions, confirming that a single genome does not adequately represent the diversity contained within G. soja. The pan-genome size was asymptotic with size of the seven genomes and an initial G. soja pan-genome with 59,080 gene families and an overall size 986.3 Mbp was constructed (Supplementary Table 19 and Supplementary Fig. 15). Nearly half (48.6%) of the gene families and 80.1% of sequences conserved across all seven G. soja genomes were core genomic units (Supplementary Figs. 14 and 15). Approximately half of the gene families (51.4% or 30,364) were present in more than one, but not all seven G. soja genomes, and represent the dispensable genome. As might be expected, unique gene families, those found only in one sample, were the least frequent and, among the seven accessions, GsojaD had the greatest amount of accession-specific gene families (Supplementary Fig. 14).
Owing to recent acquisition and deletion events21, the dispensable gene set was more variable than the core gene set. The frequency of SNPs in the core gene set was estimated to be 2.67 sites per Kbp, significantly less than 4.12/Kbp in the dispensable gene set (t-test, P < 0.01). The same bias was observed in the frequency of indels (Fig. 3b and Supplementary Table 20). Moreover, the dispensable genes had significantly higher dn, ds and dn/ds ratio values than did core genes (P < 0.01, t-test; Fig. 3b), indicating that the dispensable genes have undergone weaker purifying selection and/or greater positive selection than core genes. Core genes were enriched in biological processes including growth, immune system processes, reproduction, reproductive processes, cellular processes, and cellular component organization or biogenesis (Supplementary Table 21). Three categories were enriched in dispensable genes: receptor activity, structural molecule activity and antioxidant activity in molecular function. The core genes were more functionally conserved than the dispensable genes: 58.3% of the dispensable genes and 33.9% for the core genes set could not be assigned any functional annotation. Moreover, 95.5% of core genes had homologs in other species based on BLAST searches to 32 plant genomes (excluding soybean), significantly more than the dispensable gene set (83.5%, chi-squared test, P < 0.01). These results confirm that lineage-specific genes evolve faster than genes that are shared between species, either by means of a higher evolutionary rate22 or a higher gene loss rate23.
G. soja versus G. max: genomic basis of agronomic traits
R-genes with NBS domains mediate effector-triggered immunity acting as detectors for pathogen virulence proteins24. The quantity of R-genes with a given domain architecture and the number of unique domain architectures varied between the two species. G. soja contained more R-gene domain architectures (25) than G. max (14) (Fig. 4a). The 11 lineage-specific R-gene domain architectures found in G. soja accessions were part of the dispensable genome, possibly reflecting adaptation to biotic stresses. The G. soja genome contained a broader range of NBS R-gene domain architectures than did G. max, but G. max had more NBS-encoding genes (460 versus 334 in GsojaA to 382 in GsojaD) (Supplementary Table 22). R-genes showed more copy number loss in G. soja accessions (88 lost versus 33 gained and 8 copy number gain/loss, Supplementary Table 23), several of which were known to confer resistance to soybean mosaic virus25, Asian soybean rust26 and Phytophthora root rot27, and be involved in legume-rhizobia symbiosis28. In addition, five structurally diverged regions (located on chromosomes 03, 06, 07 and 18) within cultivated G. max29, had a high frequency of CNV in the pan-genome (Supplementary Fig. 16).
(a) Heatmap for CNV of resistance-related PFAM gene categories across all accessions. Copy number ratio of gene categories in each box was calculated as the number of genes in each G. soja accession over number of genes in GmaxW82 within the same gene category. Red represents greater gene numbers in G. soja than GmaxW82, whereas green indicates fewer. Gene numbers for GmaxW82 for the first 14 gene categories are shown on left. In categories where GmaxW82 does not have any genes (gene number = 0), black indicates the absence of genes in a given G. soja accession, while red indicates the presence of genes in G. soja vs. none in GmaxW82. (b) Large-effect mutations and new gene copy in flowering time–related genes. In PHYA4, E1 and LFY, amino acid changes resulting from large-effect SNPs and indels are shown. In the FT gene family, a new copy named FT2c was found in all seven G. soja accessions.
Genes controlling flowering time were likely involved in adaptation to new climatic regions. Comparing the G. soja genomes and GmaxW82, flowering time-related genes with large-effect variation were identified, including light receptor PHYA, floral integrator FLOWERING LOCUS T (FT) and meristem identity protein LEAFY (LFY) genes (Fig. 4b). Two of the four homologs of PHYA in soybean, GmPHYA2 and GmPHYA3, correspond to major flowering loci, E4 (ref. 30) and E3 (ref. 31), respectively. We found mutations in GmPHYA4 (Glyma03g38620), resulting in changes and deletions of amino acid (AA) in the N-terminal P3/GAF domain (Supplementary Figs. 17 and 18). Among ten FT homologs in soybean, GmFT2c showed the closest relationship to GmFT2a, a candidate gene that coordinates flowering32. In GmaxW82, GmFT2c was missing ∼50% of the N-terminal peptides (Fig. 4b and Supplementary Fig. 19); in G. soja, however, GmFT2c had a complete gene structure and was expressed. LFY, which regulates the timing of floral induction in Arabidopsis, had two putative orthologous copies in GmaxW82 [GmLFY1 (Glyma04g37900) and GmLFY2 (Glyma06g17170)]. Two 3-bp indels and nine nonsynonymous SNPs were identified in GmLFY1 among the seven G. soja accessions, all in exons 1 and 2, forming variation hotpots (Fig. 4b). Both GmFT2c and GmLFY1 were located in or near QTL, controlling the period of reproductive growth stage and maturity time33, thus providing gene candidates that may underlie flowering time in soybean.
Oil and fatty acid content of soybean has been intensely selected such that G. max produces nearly twice as much oil as G. soja34. A total of 1,332 genes were annotated in GmaxW82 based on homology to genes in 24 acyl lipid subpathways in Arabidopsis (Supplementary Table 24)35. Of these genes, 15.9% contained CNV and/or large-effect SNP and/or indels between the G. soja accessions and GmaxW82. Although five subpathways exhibited a significantly low frequency of variation (P < 0.05, Fisher's exact test; Supplementary Table 24), a number of gene variants were identified. Fatty acids are precursors for jasmonate production in the oxylipin pathway in plants following stress36. SAG101 (senescence-associated gene101) in Arabidopsis, which encodes a triacylglycerol lipase to hydrolyze triacylglycerol to monoacylglycerol and produce free fatty acid, plays important roles in plant innate immunity against biotrophic pathogens37. In this study, large-effect SNP and/or indels were found in three clustered homologs of SAG101 (Glyma13g04561, Glyma13g04651 and Glyma13g04571) and are candidate genes for nearby QTL that affect oil content and confer resistance to Phytophthora sojae38 (Supplementary Fig. 20).
Triacylglycerol is a major storage lipid in soybean and acyl-CoA:diacylglycerol acyltransferase (DGAT) is a key enzyme that catalyzes diacylglycerol to triacylglycerol, determining seed oil content in Arabidopsis35. Among 113 potential triacylglycerol biosynthesis genes, 10 had putative loss-of-function frameshifts due to large-effect SNP and/or indels, including the DGAT homolog (Glyma16g21960, GmDGAT2B) and a homolog of Abscisic Acid Insensitive 4 (ABI4) (Glyma14g06290), a transcription factor that regulates triacylglycerol synthesis in Arabidopsis by activating DGAT1 expression during nitrogen deficiency (Supplementary Fig. 21)39. A frame-shift mutation in GmDGAT2B was previously found to be associated with reduced seed oil concentration40. Another DGAT homolog, DGAT1B (Glyma17g06120), negatively associated with seed protein content40, was found to have been positively selected during domestication (protein and oil content are negatively correlated).
In contrast to G. soja, cultivated soybean has larger organ sizes, thicker stems and shorter plant height (Supplementary Fig. 10). A QTL region controlling plant height, lodging and yield spans ∼1.2Mbp on chromosome 9 (ref. 41). In this region, four genes were identified with large-effect SNP and/or indels (Supplementary Table 25). GO analysis implicated all four genes in plant development, and work in other species has shown these genes to be either directly or indirectly involved in cytokinin metabolism42,43. The identification of large-effect SNP and/or indels between G. soja and GmaxW82 can facilitate discovery and cloning of candidate genes related to the domestication and improvement of soybean.
Discussion
Our work illustrates the advantage of de novo assembly in detecting genetic variation that would not have been found by resequencing alone. The majority (94%) of G. max genes were covered by our G. soja assemblies. Thus, we were able to identify structural variation, including 338 PAV, 1,978 CNV, and a series of SNPs and indels in highly divergent genic regions, where only assembled sequences could be mapped. Using this data set, we estimated that the divergence time between G. soja and G. max was nearly three times earlier than a previous estimate based on a single G. soja accession4.
The individual genomes and the pan-genome could be improved by additional sequencing of larger insert libraries to increase contig and scaffold sizes, which would allow full exploration of the nongenic parts of the genome. These improvements would also enable further interrogation of chromosome-scale structural variation, such as inversion and translocation events. Given the structural complexity of plant genomes, further advances in sequencing technology and bioinformatics tools are needed to overcome difficulties in genome assembly and annotation to deal with highly similar transposable elements and other abundant repetitive sequences, heterozygosity and polyploidy44. Furthermore, the concept of a pan-genome should be expanded beyond the DNA level to include variation in gene regulation and expression.
This study confirms that a single genome does not adequately represent the diversity contained within a species8. We constructed a de novo assembly-based pan-genome of a crop or crop wild relatives, by sequencing seven representative wild soybean accessions from East Asia8, the center of G. soja diversity and domestication center of G. max. The G. soja pan-genome was 30.2 Mbp larger than the genome of a single resequenced accession4. The number of accessions required to represent the majority of genetic variation within a species is case-dependent45, and cataloguing a pan-genome will never be complete. The observation that the G. soja pan-genome size began to level off suggests that we have captured a representative portion of the species' gene pool. Sequencing additional samples may further reveal allelic structure and variation within and between populations, thereby allowing investigation into local adaptation and migration. Increasing the sample size may also add to the dispensable genome, which is characterized by greater variation than the core genome and is potentially involved in environmental adaptation and organismal interactions46. Approximately 20% of the G. soja sequences and 51.4% of gene families were found to be dispensable, an interesting juxtaposition to the recently published pan-transcriptome of 503 maize samples where 82.7% of the representative transcript assemblies were dispensable47, supporting the prediction that an outbred species such as maize would have a larger dispensable genome than an autogamous species45.
The pan-genome of G. soja shows the extent of novel genes and alleles in wild relatives that can be introgressed into crops, which typically have lower genetic diversity as the result of domestication and breeding48,49. Candidate genes associated with adaptation to various abiotic and biotic stresses may contribute to increased resilience to climate variability in cultivated soybean. Genes with structural variation and large-effect variation between G. soja and G. max (GmaxW82) that associate with agronomic phenotypes, as inferred from homology and comparison with mapped QTLs, can be used to develop molecular markers for these segments and test new allelic combinations. Newly identified genetic variation for genomic regions that have been fixed in G. max can be used to design crosses to determine if these fixed regions underlie phenotypes of agricultural value, providing additional candidate genes for the development of new varieties. Further, these data should enable the crop breeding community to more effectively use molecular approaches, such as genomic selection, to reduce the yield drag often associated with introgression from wild species.
Methods
DNA and RNA sequencing of seven G. soja accessions.
Seven G. soja accessions representing the array of geographical adaptation across the species, named GsojaA to GsojaG, respectively, were selected to construct pan-genome of G. soja species (Supplementary Figs. 1 and 2 and Supplementary Table 1). Of these, six represent the six major distinct clusters of G. soja species (North, Huanghuai and South regions of China, and Japan, Korea and Russia) and 87.1% of gene diversity as revealed by analysis of 99 simple sequence repeats8. To maximally represent the species' wide geographic distribution, an additional accession was selected from the Northeast region (NER) of China, one of predicted domestication centers10. Four Chinese G. soja accessions were from the Chinese National Soybean GeneBank (CNSGB), and the other three were obtained from the USDA-ARS Soybean Germplasm Collection (University of Illinois, Urbana, IL), provided by Randall Nelson. Short-insert (180 bp and 500 bp) and 2 Kbp mate-pair genomic DNA libraries were constructed for each soybean sample. RNA was extracted from eight tissues, including young leaf, flower, young pod (1 cm), pod shell (5 mm), pod shell (2 cm), seed (3 mm, half full and full). RNAs were purified and RNAs from different tissues for each accession were mixed equally to construct sequencing libraries. The libraries were paired-end sequenced on the Illumina HiSeq 2000 platform. The following types of reads were filtered out from subsequent analyses:
(a) Reads with ≥10% unidentified nucleotides (N);
(b) Reads with >10 nt aligned to the adaptor, allowing ≤10% mismatches;
(c) Reads with >50% bases having phred quality <5; and
(d) Putative PCR duplicates generated by PCR amplification in the library construction process (i.e., read 1 and read 2 of two paired-end reads that were completely identical).
A total of 779.2 Gb DNA (average 111.9× coverage per sample) and 35.47 Gb (average 5.07 Gb per sample) of RNA sequence data were obtained.
De novo assembly.
First, we generated a 17-mer depth distribution of short-insert paired-end reads using Meryl50 and applied GCE51 to estimate the genome sizes of individual G. soja accessions. Reads were preprocessed by ALLPATHS-LG52 error correction module to remove base calling errors. We also used ErrorCorrection in SOAPdenovo11 package to connect 180-bp library pair end reads and to generate longer sequences for assembly. Reads of 180-bp and 500-bp library were used for contig building, and all pair-end reads libraries were used to provide links for scaffold construction. GapCloser (v1.12) from SOAPdenovo11 package was used for gap filling within assembled scaffolds using all pair-end reads. Finally, scaffold sequences, which can be aligned to bacterial genomes with identity ≥95% and e-value ≤1e-5, were filtered.
Genomic alignment and short read mapping.
Assembled scaffolds for all G. soja accessions were aligned to the reference genome (GmaxW82) using the NUCmer program from the MUMmer package53. The parameters used for the genome alignment were “-maxmatch -c 90 -l 40”. The alignment results were further filtered to retain only one-to-one alignment regions using the “delta-filter” program incorporated in MUMmer package53. Short reads were mapped to both GmaxW82 and G. soja genomes using BWA54. After BWA alignment, putative PCR duplicates were filtered using the 'rmdup' utility of SAMtools55.
Gene prediction and annotation.
Genome assemblies were scanned for putative gene coding regions using the Augustus56 package. Then protein sequences from Glycine max, Lotus japonicas, Medicago truncatula and Arabidopsis thaliana were mapped to our assemblies using TblastN57 (1e-5) and refined by GeneWise58. RNA-seq reads were mapped to G. soja assemblies using TopHat (v2.0.7)59 with default parameters to identify exon regions and splice positions, and transcriptome-based gene structure predictions were made by Cufflinks (v2.0.2)60 with default parameters. Results by ab initio, homology-based and transcriptome-based predictions were combined with the Evidence Modeler (EVM) package61 (EVM set). A second GeneWise58 prediction was done by using only proteins from G. max to identify the best gene model. To exclude putative false positive predictions, protein sequences from the final gene models were aligned against protein annotations from 18 plant species using BLAST57 (1e-5), including Glycine max, Cajanus cajan, Phaseolus vulgaris, Lotus japonicas, Medicago truncatula, Populus trichocarpa, Ricinus communis, Cucumis sativus, Malus domestica, Arabidopsis thaliana, Brassica rapa, Carica papaya, Vitis vinifera, Solanum lycopersicum, Musa acuminate, Sorghum bicolor, Zea mays and Oryza sativa. Gene models with <50% alignment coverage of their best homologs were filtered out. Gene models with >20% CDS region covered by repeat contents or corresponding protein sequence matched to TE-related domains by HMMPfam62 were also filtered out. Finally, we checked gene models that were split into small fragments due to the discontinuity of scaffolds and revised our prediction of genes in each of the seven G. soja genomes. Putative biological functions of individual genes were assigned according to the best BLAST57 hits in the Arabidopsis thaliana proteome, and by searching publicly available databases including Pfam62, PRINTS, PROSITE, ProDom and SMART with InterProScan63. Gene Ontology (GO)64 terms for individual genes were retrieved from the corresponding InterPro descriptions. We also mapped these genes to the KEGG pathway65 to identify their best matched categories.
Gene clustering.
The core and dispensable gene sets were estimated based on gene family clustering. Based on the OrthoMCL66 clustering results, we extracted gene families that were shared between G. soja samples, which were defined as core gene families. Gene families that are missed in one or more soybean samples were defined as dispensable gene families.
SNP and indel identification.
Homozygous SNPs and small indels (less than 100 bp) were extracted from the one-to-one genomic alignment results using MUMmer53. We detected heterozygous SNPs using SAMtools55 based on alignments of short reads onto assembled G. soja genomes. Then we located these heterozygous SNP sites on GmaxW82 genome according to the one-to-one genome alignment results. For indels larger than 100 bp, we searched for all alignment gaps (>100 bp) in both G. soja genomes and GmaxW82 genome. Indels identified as gaps surrounded by one-to-one alignments with at least 100 bp in both ends were retained. We used EMBOSS Water67 based on the standard Smith-Waterman algorithm68 to revise the retained indel set by redoing the alignments of regions that contain multiple indels. Putative functional effects of SNPs and indels were annotated using the ANNOVAR package69. SNPs/indels causing stop codon gain, stop codon loss and frameshift were defined as large-effect mutations. Gene Ontology category enrichments for genes containing large-effect mutations were found using the FUNC package70. To validate our results, we randomly selected 51 SNPs and 75 indel events, which is, 173 SNP sites and 20 indel sites in seven G. soja accessions, for PCR-Sanger sequencing using the ABI 3730XL. Additional 73 SNP events at 25 SNP sites were selected for mass-spectrometric assay (Sequenom). Among these sites, 81 SNP and 64 indel events were only detected with the current approach and could not be found via resequencing approach alone.
CNV detection.
To avoid abnormal depths in non-coding regions owing to repeat elements, we focused on CNV in gene regions. Only CDS regions were sampled as background for CNV detection. The mean sequencing depth (d0) and s.d. (s0) were calculated for the background data set. Sequencing depth of each gene's CDS was extracted, and the mean value (dg) and s.d. (sg) were calculated. A standard t-test was applied to genes with dg below 0.2d0 or above 1.8d0, and those with P-value ≤ 1e-5 were defined as CNV. Genes with high similarity with GmaxW82 chloroplast and mitochondrion (50% length covered by e-value ≤ 1e-5 and identity ≥95% BLAST57 hits) were not included.
PAV detection.
For sequences that could not be aligned to GmaxW82, we used BLAST57 to realign them to the GmaxW82 genome, all whole-genome sequencing trace reads and bacterial artificial clone sequences of G. max from GenBank, and filtered sequence stretches with an identity larger than 95%. G. soja–specific sequences were obtained after excluding potential bacterial contamination based on BLAST57 to the NT database. After filtering for putative repeat elements (80% coverage) through RepeatMasker (http://www.repeatmasker.org/)71 using an in-house repeat database72, genes with >50% CDS regions covered by G. soja–specific sequences were defined as G. soja–specific genes. Based on the short reads alignment results, blocks with no mapped reads by G. soja were defined as GmaxW82-specific sequences. Regions with distance less than 500 bp were merged into one block. Genes that overlapped these blocks with 50% length were considered as GmaxW82-specific sequences.
G. soja–specific sequence validation.
G. soja–specific sequences were validated using multiplex PCR amplification. Each amplification included two pairs of primers, G. soja–specific sequence (Supplementary Table 12) and a control sequence shared by G. soja and G. max. For the control sequence (C1, 218 bp in GmaxW82), the forward and reverse primers were GTCATTGTAACAGGTGGGGAGA and ACTGCGACTTTATTTAAGATAG (C1). For each sequence, one or two (different locations) multiplexed PCRs were tested.
Phylogenetic analysis.
Protein coding genes from G. max, Cajanus cajan, Phaseolus vulgaris, Medicago truncatula, Populus trichocarpa, Cucumis sativus, Arabidopsis thaliana and the seven sequenced G. soja accessions were used for gene family construction. For genes with alternative splicing isoforms, the longest isoform for each gene was used. Protein sequences for genes were compared by using all-by-all BLASTP57 (1e-5), and then OrthoMCL66 was used to cluster genes into orthologous gene families. From the clustering results, 670 orthologous gene families, with exactly one copy from each genome, were classified and defined as conserved single-copy gene families. Protein sequences from the 670 gene families were aligned by MUSCLE73. Fourfold degenerative sites from the CDS alignments were extracted and concatenated for phylogenetic analysis. Phylogenetic trees were built by the Neighbor-joining method incorporated in MEGA package74. The divergence time between legume genomes were estimated using the 'mcmctree' program incorporated in the PAML75 package. We identified the SNPs in 57 published soybean accessions3,4 together with our seven G. soja accessions using SAMtools55 and constructed a separate phylogenetic tree for these 64 soybean accessions.
Construction of the G. soja pan-genome.
First, we defined the core genome, which is shared by all seven G. soja accessions. Based on genomic alignment results, the aligned regions on GmaxW82 genome for each G. soja accession contributed to the first portion of the G. soja core genome. For sequences that could not be aligned to GmaxW82, Mugsy76 was used to make multiple alignments using the default parameters. From the multiple alignments, we identified sequences that were shared in seven samples, which contributed to the second portion of the core genome. Sequences shared by less than seven samples, plus those presented in only one sample, were defined as the G. soja dispensable genome.
Identification of positive selection and domestication related genes.
The syntenic orthologous gene families were used to estimate selection pressure by using the branch-site model incorporated in the PAML package. Based on a maximum likelihood ratio test (LRT), we identified genes under positive selection in GmaxW82 and in each G. soja lineage. These genes were identified as positively selected according to the chi-squared test (P < 0.01, FDR < 0.05, df = 1), and containing amino acid sites that were selected with a Bayes probability higher than 95%.
R-gene classification.
All genes from the G. max and G. soja samples were annotated using the HMM model against the Pfam-A database62. Genes with the Pfam domain of PF00931 were defined as R-genes, classified by different Pfam domains with E-value of 0.001. Domains of PF00931 (NB-ARC) and PF01582 (TIR) were defined as NBS and TIR domains, respectively, whereas domains with leucine-rich repeat, including PF00560 (LRR_1), PF07725 (LRR_3), PF12799 (LRR_4), PF13306 (LRR_5), PF13516 (LRR_6), PF13504 (LRR_7) and PF13855 (LRR_8), were defined as LRR domains. TIR-only and TIR-X sequences were not included in this study. For R-genes with CNV, the expression analysis result and related traits reported in previous studies25,26,27,28,77,78,79 were listed in Supplementary Table 23.
Accession codes.
Genome assembly data, BioProject: PRJNA195632. Unassembled sequencing reads and raw sequencing reads of the transcriptome, SRA: SRP040255.
Accession codes
References
Qiu, L.J. et al. A platform for soybean molecular breeding: the utilization of core collections for food security. Plant Mol. Biol. 83, 41–50 (2013).
Schmutz, J. et al. Genome sequence of the palaeopolyploid soybean. Nature 463, 178–183 (2010).
Lam, H.M. et al. Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat. Genet. 42, 1053–1059 (2010).
Kim, M.Y. et al. Whole-genome sequencing and intensive analysis of the undomesticated soybean (Glycine soja Sieb. and Zucc.) genome. Proc. Natl. Acad. Sci. USA 107, 22032–22037 (2010).
Li, Y.H. et al. Molecular footprints of domestication and improvement in soybean revealed by whole genome re-sequencing. BMC Genomics 14, 579 (2013).
Hufford, M.B. et al. Comparative population genomics of maize domestication and improvement. Nat. Genet. 44, 808–811 (2012).
Li, R. et al. Building the sequence map of the human pan-genome. Nat. Biotechnol. 28, 57–63 (2010).
Li, Y.H. et al. Genetic diversity in domesticated soybean (Glycine max) and its wild progenitor (Glycine soja) for simple sequence repeat and single-nucleotide polymorphism loci. New Phytol. 188, 242–253 (2010).
Tettelin, H. et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial 'pan-genome'. Proc. Natl. Acad. Sci. USA 102, 13950–13955 (2005).
Fukuda, Y. Cytological studies on the wild and cultivated Manchurian soybeans. Jap. J. Bot. 6, 489–506 (1933).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Shoji, T. et al. Plant-specific microtubule-associated protein SPIRAL2 is required for anisotropic growth in Arabidopsis. Plant Physiol. 136, 3933–3944 (2004).
Chanda, B. et al. Glycerol-3-phosphate is a critical mobile inducer of systemic immunity in plants. Nat. Genet. 43, 421–427 (2011).
Weise, A. et al. A new subfamily of sucrose transporters, SUT4, with low affinity/high capacity localized in enucleate sieve elements of plants. Plant Cell 12, 1345–1355 (2000).
Lintala, M. et al. Structural and functional characterization of ferredoxin-ADP+-xidoreductase using knock-out mutants of Arabidopsis. Plant J. 49, 1041–1052 (2007).
Swanson-Wagner, R.A. et al. Pervasive gene content variation and copy number variation in maize and its undomesticated progenitor. Genome Res. 20, 1689–1699 (2010).
Jiao, Y. et al. Genome-wide genetic changes during modern breeding of maize. Nat. Genet. 44, 812–815 (2012).
Arakaki, M. et al. Contemporaneous and recent radiations of the world's major succulent plant lineages. Proc. Natl. Acad. Sci. USA 108, 8379–8384 (2011).
Varshney, R.K. et al. Draft genome sequence of pigeonpea (Cajanus cajan), an orphan legume crop of resource-poor farmers. Nat. Biotechnol. 30, 83–89 (2012).
Lee, G.A., Crawford, G.W., Liu, L., Sasaki, Y. & Chen, X. Archaeological soybean (Glycine max) in East Asia: does size matter? PLoS ONE 6, e26720 (2011).
Donati, C. et al. Structure and dynamics of the pan-genome of Streptococcus pneumoniae and closely related species. Genome Biol. 11, R107 (2010).
Cai, J.J. & Petrov, D.A. Relaxed purifying selection and possibly high rate of adaptation in primate lineage-specific genes. Genome Biol. Evol. 2, 393–409 (2010).
Krylov, D.M., Wolf, Y.I., Rogozin, I.B. & Koonin, E.V. Gene loss, protein sequence divergence, gene dispensability, expression level, and interactivity are correlated in eukaryotic evolution. Genome Res. 13, 2229–2235 (2003).
Innes, R.W. et al. Differential accumulation of retroelements and diversification of NB-LRR disease resistance genes in duplicated regions following polyploidy in the ancestor of soybean. Plant Physiol. 148, 1740–1759 (2008).
Wen, R.H., Khatabi, B., Ashfield, T., Maroof, M.S. & Hajimorad, M. The HC-Pro and P3 cistrons of an avirulent soybean mosaic virus are recognized by different resistance genes at the complex Rsv1 locus. Mol. Plant Microbe Interact. 26, 203–215 (2013).
Monteros, M.J., Ha, B.-K., Phillips, D.V. & Boerma, H.R. SNP assay to detect the 'Hyuuga'red-brown lesion resistance gene for Asian soybean rust. Theor. Appl. Genet. 121, 1023–1032 (2010).
Zhang, J. et al. Genetic characterization and fine mapping of the novel Phytophthora resistance gene in a Chinese soybean cultivar. Theor. Appl. Genet. 126, 1555–1561 (2013).
Yang, S., Tang, F., Gao, M., Krishnan, H.B. & Zhu, H. R gene-controlled host specificity in the legume–rhizobia symbiosis. Proc. Natl. Acad. Sci. USA 107, 18735–18740 (2010).
McHale, L.K. et al. Structural variants in the soybean genome localize to clusters of biotic stress-response genes. Plant Physiol. 159, 1295–1308 (2012).
Liu, B. et al. Genetic redundancy in soybean photoresponses associated with duplication of the phytochrome A gene. Genetics 180, 995–1007 (2008).
Watanabe, S. et al. Map-based cloning of the gene associated with the soybean maturity locus E3. Genetics 182, 1251–1262 (2009).
Kong, F. et al. Two coordinately regulated homologs of FLOWERING LOCUS T are involved in the control of photoperiodic flowering in soybean. Plant Physiol. 154, 1220–1231 (2010).
Xin, D.W. et al. Analysis of quantitative trait loci underlying the period of reproductive growth stages in soybean (Glycine max [L.] Merr.). Euphytica 162, 155–165 (2008).
Xu, B. et al. A study on fat content and fatty acid composition of wild soybean (G. soja) in China. Jinlin Agric. Sci. 2, 1–6 (1993).
Li-Beisson, Y. et al. Acyl-lipid metabolism. The Arabidopsis Book 11, e0161 (2013).
Kachroo, A. & Kachroo, P. Fatty acid-derived signals in plant defense. Annu. Rev. Phytopathol. 47, 153–176 (2009).
Feys, B.J. et al. Arabidopsis SENESCENCE-ASSOCIATED GENE101 stabilizes and signals within an ENHANCED DISEASE SUSCEPTIBILITY1 complex in plant innate immunity. Plant Cell 17, 2601–2613 (2005).
Qi, Z.M. et al. Soybean oil content QTL mapping and integrating with meta-analysis method for mining genes. Euphytica 179, 499–514 (2011).
Yang, Y., Yu, X., Song, L. & An, C. ABI4 activates DGAT1 expression in Arabidopsis seedlings during nitrogen deficiency. Plant Physiol. 156, 873–883 (2011).
Eskandari, M., Cober, E.R. & Rajcan, I. Using the candidate gene approach for detecting genes underlying seed oil concentration and yield in soybean. Theor. Appl. Genet. 126, 1839–1850 (2013).
Wang, D., Graef, G., Procopiuk, A. & Diers, B. Identification of putative QTL that underlie yield in interspecific soybean backcross populations. Theor. Appl. Genet. 108, 458–467 (2004).
Ashikari, M. et al. Cytokinin oxidase regulates rice grain production. Science 309, 741–745 (2005).
Johnson, K. & Lenhard, M. Genetic control of plant organ growth. New Phytol. 191, 319–333 (2011).
Claros, M.G. et al. Why assembling plant genome sequences is so challenging. Biology 1, 439–459 (2012).
Medini, D., Donati, C., Tettelin, H., Masignani, V. & Rappuoli, R. The microbial pan-genome. Curr. Opin. Genet. Dev. 15, 589–594 (2005).
Read, B.A. et al. Pan genome of the phytoplankton Emiliania underpins its global distribution. Nature 499, 209–213 (2013).
Hirsch, C.N. et al. Insights into the maize pan-genome and pan-transcriptome. Plant Cell 26, 121–135 (2014).
Hyten, D.L. et al. Impacts of genetic bottlenecks on soybean genome diversity. Proc. Natl. Acad. Sci. USA 103, 16666–16671 (2006).
Doebley, J.F., Gaut, B.S. & Smith, B.D. The molecular genetics of crop domestication. Cell 127, 1309–1321 (2006).
Myers, E.W. et al. A whole-genome assembly of Drosophila. Science 287, 2196–2204 (2000).
Liu, B. et al. Estimation of genomic characteristics by analyzing k-mer frequency in de novo genome projects. Preprint at http://arxiv.org/abs/1308.2012 (2012).
Butler, J. et al. ALLPATHS: de novo assembly of whole-genome shotgun microreads. Genome Res. 18, 810–820 (2008).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19, ii215–ii225 (2003).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Birney, E., Clamp, M. & Durbin, R. GeneWise and genomewise. Genome Res. 14, 988–995 (2004).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Haas, B.J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7 (2008).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Res. 40, D290–D301 (2012).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res. 33, W116–W120 (2005).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Li, L., Stoeckert, C.J. & Roos, D.S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European molecular biology open software suite. Trends Genet. 16, 276–277 (2000).
Smith, T.F. & Waterman, M.S. Identification of common molecular subsequences. J. Mol. Biol. 147, 195–197 (1981).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Prüfer, K. et al. FUNC: a package for detecting significant associations between gene sets and ontological annotations. BMC Bioinformatics 8, 41 (2007).
Smit, A.F., Hubley, R. & Green, P. RepeatMasker Open-3.0 (1996).
Du, J. et al. SoyTEdb: a comprehensive database of transposable elements in the soybean genome. BMC Genomics 11, 113 (2010).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Yang, Z. PAML: a program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13, 555–556 (1997).
Angiuoli, S.V. & Salzberg, S.L. Mugsy: fast multiple alignment of closely related whole genomes. Bioinformatics 27, 334–342 (2011).
Kang, Y. et al. Genome-wide mapping of NBS-LRR genes and their association with disease resistance in soybean. BMC Plant Biol. 12, 139 (2012).
Kim, K.H. et al. RNA-Seq analysis of a soybean near-isogenic line carrying bacterial leaf pustule-resistant and-susceptible alleles. DNA Res. 18, 483–497 (2011).
Suh, S.J. et al. The Rsv3 locus conferring resistance to soybean mosaic virus is associated with a cluster of coiled-coil nucleotide-binding leucine-rich repeat genes. Plant Genome 4, 55–64 (2011).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (no. 31271753), the State Key Basic Research and Development Plan of China (973) (no. 2010CB125903 and 2009CB118404), the International Science and Technology Cooperation and Exchanges Projects (no. 2008DFA30550), the Platform of National Crop Germplasm Resources of China (nos. 2012-004 and 2013-004) and US National Science Foundation (no. DBI 0822258). We thank R. Stupar for critical reading of this manuscript and valuable suggestions. We also thank H. Zhu, Y. Xin, X. Meng and many additional staff at Novogene Bioinformatics Institute who contributed to this teamwork.
Author information
Authors and Affiliations
Contributions
L.Q., R.L., S.A.J., Ying-hui Li, G.Z., and J.M. conceived the study and jointly wrote the paper. Ying-hui Li, S.Z., X.S., Yan-fei Li, H.H., Z.L., K.W., Li-juan Zhang and R.C. provided DNAs and performed PCR validation and Sanger sequencing. G.Z., R.L., W.J., Z.Z., J.Z., Q.Z., Y.W., H.R., H.W., J.L. and Z.J. performed sequencing, genome assembly and genome annotation. Ying-hui Li, G.Z., W.J., J.Z., S.A.J., J.M., J.L., Y.G., Y.S., Z.Z., B.T., J.S., L.J., Liangtao Zheng, Y.H., G.K., C.K.L.Y., W.Z., R.G., W.L., S.C. and L.Q. performed comparative, population and evolutionary biology analyses.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–21 and Supplementary Tables 1–25 (PDF 9993 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike license (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This license does not permit commercial exploitation, and derivative works must be licensed under the same or similar license.
About this article
Cite this article
Li, Yh., Zhou, G., Ma, J. et al. De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits. Nat Biotechnol 32, 1045–1052 (2014). https://doi.org/10.1038/nbt.2979
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2979
This article is cited by
-
A chromosome-level genome assembly of the Echiura Urechis unicinctus
Scientific Data (2024)
-
Pan-genome of Citrullus genus highlights the extent of presence/absence variation during domestication and selection
BMC Genomics (2023)
-
Genetic diversity, population genetic structure and gene flow in the rare and endangered wild plant Cypripedium macranthos revealed by genotyping-by-sequencing
BMC Plant Biology (2023)
-
Whole-genome resequencing and transcriptome analyses of four generation mutants to reveal spur-type and skin-color related genes in apple (Malus domestica Borkh. Cv. Red delicious)
BMC Plant Biology (2023)
-
A review of the pangenome: how it affects our understanding of genomic variation, selection and breeding in domestic animals?
Journal of Animal Science and Biotechnology (2023)