Abstract
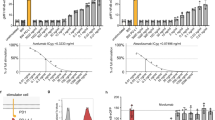
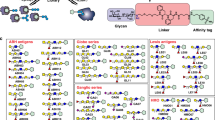
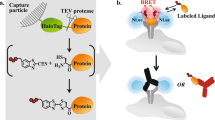
Many cellular responses are triggered by proteins, drugs or pathogens binding to cell-surface receptors, but it can be challenging to identify which receptors are bound by a given ligand. Here we describe TRICEPS, a chemoproteomic reagent with three moieties—one that binds ligands containing an amino group, a second that binds glycosylated receptors on living cells and a biotin tag for purifying the receptor peptides for identification by quantitative mass spectrometry. We validated this ligand-based, receptor-capture (LRC) technology using insulin, transferrin, apelin, epidermal growth factor, the therapeutic antibody trastuzumab and two DARPins targeting ErbB2. In some cases, we could also determine the approximate ligand-binding sites on the receptors. Using TRICEPS to label intact mature vaccinia viruses, we identified the cell surface proteins AXL, M6PR, DAG1, CSPG4 and CDH13 as binding factors on human cells. This technology enables the identification of receptors for many types of ligands under near-physiological conditions and without the need for genetic manipulations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hubner, N.C. et al. Quantitative proteomics combined with BAC TransgeneOmics reveals in vivo protein interactions. J. Cell Biol. 189, 739–754 (2010).
Glatter, T., Wepf, A., Aebersold, R. & Gstaiger, M. An integrated workflow for charting the human interaction proteome: insights into the PP2A system. Mol. Syst. Biol. 5, 237 (2009).
Bantscheff, M. & Drewes, G. Chemoproteomic approaches to drug target identification and drug profiling. Bioorg. Med. Chem. 20, 1973–1978 (2012).
Lenz, T., Fischer, J.J. & Dreger, M. Probing small molecule-protein interactions: A new perspective for functional proteomics. J. Proteomics 75, 100–115 (2011).
Barglow, K.T. & Cravatt, B.F. Activity-based protein profiling for the functional annotation of enzymes. Nat. Methods 4, 822–827 (2007).
Elschenbroich, S., Kim, Y., Medin, J.A. & Kislinger, T. Isolation of cell surface proteins for mass spectrometry-based proteomics. Expert Rev. Proteomics 7, 141–154 (2010).
Helbig, A.O., Heck, A.J.R. & Slijper, M. Exploring the membrane proteome–challenges and analytical strategies. J. Proteomics 73, 868–878 (2010).
Savas, J.N., Stein, B.D., Wu, C.C. & Yates, J.R. Mass spectrometry accelerates membrane protein analysis. Trends Biochem. Sci. 36, 388–396 (2011).
Lee, A. How lipids affect the activities of integral membrane proteins. Biochim. Biophys. Acta. 1666, 62–87 (2004).
Zeng, Y., Ramya, T.N.C., Dirksen, A., Dawson, P.E. & Paulson, J.C. High-efficiency labeling of sialylated glycoproteins on living cells. Nat. Methods 6, 207–209 (2009).
Wollscheid, B. et al. Mass-spectrometric identification and relative quantification of N-linked cell surface glycoproteins. Nat. Biotechnol. 27, 378–386 (2009).
Hofmann, A. et al. Proteomic cell surface phenotyping of differentiating acute myeloid leukemia cells. Blood 116, e26–e34 (2010).
Frei, A., Jeon, O.Y., Carreira, E. & Wollscheid, B. Trifunctional crosslinking reagents. European Patent Application No. 11000731 (2012).
Mädler, S., Bich, C., Touboul, D. & Zenobi, R. Chemical cross-linking with NHS esters: a systematic study on amino acid reactivities. J. Mass Spectrom. 44, 694–706 (2009).
Sletten, E.M. & Bertozzi, C.R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Edn Engl. 48, 6974–6998 (2009).
Ferguson, K.M. Structure-based view of epidermal growth factor receptor regulation. Annu. Rev. Biophys. 37, 353–373 (2008).
Cho, H.-S. et al. Structure of the extracellular region of HER2 alone and in complex with the Herceptin Fab. Nature 421, 756–760 (2003).
Steiner, D., Forrer, P. & Plückthun, A. Efficient selection of DARPins with sub-nanomolar affinities using SRP phage display. J. Mol. Biol. 382, 1211–1227 (2008).
Marsh, M. & Helenius, A. Virus entry: open sesame. Cell 124, 729–740 (2006).
Chung, C., Hsiao, J., Chang, Y. & Chang, W. A27L protein mediates vaccinia virus interaction with cell surface heparan sulfate. J. Virol. 72, 1577–1585 (1998).
Hsiao, J.C., Chung, C.S. & Chang, W. Vaccinia virus envelope D8L protein binds to cell surface chondroitin sulfate and mediates the adsorption of intracellular mature virions to cells. J. Virol. 73, 8750–8761 (1999).
Chiu, W.-L., Lin, C.-L., Yang, M.-H., Tzou, D.-L.M. & Chang, W. Vaccinia virus 4c (A26L) protein on intracellular mature virus binds to the extracellular cellular matrix laminin. J. Virol. 81, 2149–2157 (2007).
Mercer, J. & Helenius, A. Vaccinia virus uses macropinocytosis and apoptotic mimicry to enter host cells. Science 320, 531–535 (2008).
Mercer, J. et al. Vaccinia virus strains use distinct forms of macropinocytosis for host-cell entry. Proc. Natl. Acad. Sci. USA 107, 9346–9351 (2010).
Schroeder, N., Chung, C.-S., Chen, C.-H., Liao, C.-L. & Chang, W. The lipid raft-associated protein CD98 is required for vaccinia virus endocytosis. J. Virol. 86, 4868–4882 (2012).
Morizono, K. et al. The soluble serum protein Gas6 bridges virion envelope phosphatidylserine to the TAM receptor tyrosine kinase Axl to mediate viral entry. Cell Host Microbe 9, 286–298 (2011).
Steu, S. et al. A procedure for tissue freezing and processing applicable to both intra-operative frozen section diagnosis and tissue banking in surgical pathology. Virchows Arch. 452, 305–312 (2008).
Clough, T. et al. Protein quantification in label-free LC-MS experiments. J. Proteome Res. 8, 5275–5284 (2009).
Acknowledgements
We greatly acknowledge T. Clough and O. Vitek at Purdue University for help with statistical data analysis. We are grateful to A. Hofmann, T. Bock, D. Bausch-Fluck, F. Cerciello, A. Jacobs and A. Leitner for suggestions and support at all stages of the project. We acknowledge S. Dettwiler, P. Schraml, M. Tinguely, H. Moch and the Laboratory for In situ Technologies, University Hospital Zurich, for preparation and staining of breast cancer tissues. This work was supported by funding from National Center of Competence in Research (NCCR) Neural Plasticity and Repair (to B.W.), Swiss National Science Foundation (SNSF) (to B.W.), SystemsX.ch/InfectX (to B.W.), SNSF Ambizione (to J.M.), SystemsX.ch and European Research Council (ERC) (to R.A.) and SystemsX.ch/InfectX and ERC (to S.K. on behalf of A. Helenius). Immortalized murine pre-adipocytes were kindly provided by M. Rosenwald and C. Wolfrum (ETH Zurich). ErbB2-negative breast carcinoma tissue cut into 50 μm slices was kindly provided by the tissue biobank of the Institute of Surgical Pathology, University Hospital Zurich.
Author information
Authors and Affiliations
Contributions
A.P.F. and B.W. designed the project and wrote the paper. A.P.F. performed experiments and analyzed all data. A.P.F., B.W., O.-Y.J. and E.M.C. designed TRICEPS and O.-Y.J. synthesized the reagents. J.M. and S.K. designed and performed vaccinia virus experiments and J.M. edited the manuscript. C.J. and A.P. designed DARPin experiments and performed ELISAs. R.A., H.M. and L.M.H. contributed ideas and performed experiments. All authors discussed the results and implications and commented on the manuscript at all stages.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Note 1 (PDF 1786 kb)
Supplementary Table 1
LRC with human insulin on murine adipocytes (XLS 57 kb)
Supplementary Table 2
LRC competition experiment with human insulin on Jurkat T lymphocytes (XLS 204 kb)
Supplementary Table 3
LRC with transferrin and apelin on U-2 OS cells (XLS 90 kb)
Supplementary Table 4
LRC with EGF and trastuzumab on U251 cells (XLS 54 kb)
Supplementary Table 5
LRC with DARPin 9.01 and DARPin H14 on BT-474 cells (XLS 50 kb)
Supplementary Table 6
LRC with trastuzumab on primary breast cancer tissue (XLS 207 kb)
Supplementary Table 7
LRC with vaccinia virus on HeLa CCL2 cells (XLS 57 kb)
Supplementary Table 8
siRNA sequences used for target protein depletion (XLS 26 kb)
Rights and permissions
About this article
Cite this article
Frei, A., Jeon, OY., Kilcher, S. et al. Direct identification of ligand-receptor interactions on living cells and tissues. Nat Biotechnol 30, 997–1001 (2012). https://doi.org/10.1038/nbt.2354
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2354
This article is cited by
-
Profiling the interactome of oligonucleotide drugs by proximity biotinylation
Nature Chemical Biology (2024)
-
The vaccinia chondroitin sulfate binding protein drives host membrane curvature to facilitate fusion
EMBO Reports (2024)
-
The NERP-4–SNAT2 axis regulates pancreatic β-cell maintenance and function
Nature Communications (2023)
-
Non-invasive mapping of systemic neutrophil dynamics upon cardiovascular injury
Nature Cardiovascular Research (2023)
-
Identification of lamprey variable lymphocyte receptors that target the brain vasculature
Scientific Reports (2022)