Abstract
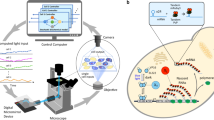
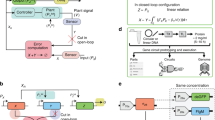
We show that difficulties in regulating cellular behavior with synthetic biological circuits may be circumvented using in silico feedback control. By tracking a circuit's output in Saccharomyces cerevisiae in real time, we precisely control its behavior using an in silico feedback algorithm to compute regulatory inputs implemented through a genetically encoded light-responsive module. Moving control functions outside the cell should enable more sophisticated manipulation of cellular processes whenever real-time measurements of cellular variables are possible.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Lim, W.A. Nat. Rev. Mol. Cell Biol. 11, 393–403 (2010).
Andrianantoandro, E., Basu, S., Karig, D.K. & Weiss, R. Mol. Syst. Biol. 2 2006.0028 (2006).
Benner, S.A. & Sismour, A.M. Nat. Rev. Genet. 6, 533–543 (2005).
Khalil, A.S. & Collins, J.J. Nat. Rev. Genet. 11, 367–379 (2010).
Purnick, P.E. & Weiss, R. Nat. Rev. Mol. Cell Biol. 10, 410–422 (2009).
Haynes, K.A. & Silver, P.A. J. Cell Biol. 187, 589–596 (2009).
Levskaya, A., Weiner, O.D., Lim, W.A. & Voigt, C.A. Nature 461, 997–1001 (2009).
Tyszkiewicz, A.B. & Muir, T.W. Nat. Methods 5, 303–305 (2008).
Leung, D.W., Otomo, C., Chory, J. & Rosen, M.K. Proc. Natl. Acad. Sci. USA 105, 12797–12802 (2008).
Shimizu-Sato, S., Huq, E., Tepperman, J.M. & Quail, P.H. Nat. Biotechnol. 20, 1041–1044 (2002).
Kalman, R.E. J. Basic Eng. 82, 35–45 (1960).
Morari, M. Comput. Chem. Eng. 23, 667–682 (1999).
Sorokina, O. et al. J. Biol. Eng. 3, 15 (2009).
Acknowledgements
We would like to thank P. Quail (UC, Berkeley) for the generous gift of the PIF3 construct and W. Lim (UCSF) and J. Stelling (ETH, Zurich) for providing PCB. The work was supported by National Science Foundation grant CCF-0943385 (H.E.-S.), ECCS-0835847 (M.K.), and MoVeS FP7-ICT-2009-257005 (J.L.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Figures 1–4 (PDF 651 kb)
Rights and permissions
About this article
Cite this article
Milias-Argeitis, A., Summers, S., Stewart-Ornstein, J. et al. In silico feedback for in vivo regulation of a gene expression circuit. Nat Biotechnol 29, 1114–1116 (2011). https://doi.org/10.1038/nbt.2018
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2018
This article is cited by
-
Maximizing protein production by keeping cells at optimal secretory stress levels using real-time control approaches
Nature Communications (2023)
-
Multidimensional characterization of inducible promoters and a highly light-sensitive LOV-transcription factor
Nature Communications (2023)
-
Decentralized PI Controller Design for Robust Perfect Adaptation in Noisy Time-Delayed Genetic Regulatory Networks
Neural Processing Letters (2023)
-
Dynamic cybergenetic control of bacterial co-culture composition via optogenetic feedback
Nature Communications (2022)
-
CyberSco.Py an open-source software for event-based, conditional microscopy
Scientific Reports (2022)