Abstract
Fibroblasts are non-haematopoietic structural cells that define the architecture of organs, support the homeostasis of tissue-resident cells and have key roles in fibrosis, cancer, autoimmunity and wound healing1. Recent studies have described fibroblast heterogeneity within individual tissues1. However, the field lacks a characterization of fibroblasts at single-cell resolution across tissues in healthy and diseased organs. Here we constructed fibroblast atlases by integrating single-cell transcriptomic data from about 230,000 fibroblasts across 17 tissues, 50 datasets, 11 disease states and 2 species. Mouse fibroblast atlases and a DptIRESCreERT2 knock-in mouse identified two universal fibroblast transcriptional subtypes across tissues. Our analysis suggests that these cells can serve as a reservoir that can yield specialized fibroblasts across a broad range of steady-state tissues and activated fibroblasts in disease. Comparison to an atlas of human fibroblasts from perturbed states showed that fibroblast transcriptional states are conserved between mice and humans, including universal fibroblasts and activated phenotypes associated with pathogenicity in human cancer, fibrosis, arthritis and inflammation. In summary, a cross-species and pan-tissue approach to transcriptomics at single-cell resolution has identified key organizing principles of the fibroblast lineage in health and disease.
Similar content being viewed by others
Main
Fibroblasts populate all tissues, delineate the topography of organs by producing and remodelling extracellular matrix proteins (ECMs)2 and support other tissue-resident cell types1,3. Fibroblasts perform functions associated with their lineage and specialized programs suited to the needs of specific tissue contexts to maintain organ homeostasis. Macrophages achieve generalized function and specialization via a lineage-wide core transcriptomic signature and tissue-specific programming driven by microenvironmental cues4,5,6. It is unclear how fibroblasts execute functions both common to their lineage and required by their organ of residence.
Technologies such as single-cell RNA-sequencing (scRNA-seq) have revealed intra-tissue fibroblast heterogeneity1. Elucidating the inter-tissue population structure of fibroblasts has clinical relevance, as subtypes of fibroblasts drive disease in arthritis7,8,9, cancer10,11,12 and fibrotic indications such as idiopathic pulmonary fibrosis (IPF)2. Emerging paradigms7,10,13,14 suggest that discrete fibroblast subtypes within tissues govern distinct aspects of tissue homeostasis and disease. Understanding whether fibroblast phenotypes across indications are context-specific or more broadly conserved may inform therapeutic approaches. We hypothesized that fibroblast heterogeneity was promoted by tissue type in the steady-state and disease context during perturbation.
Fibroblasts in steady-state mouse tissues
To investigate this hypothesis, we first performed bulk RNA-seq and assay for transposase-accessible chromatin with sequencing (ATAC–seq) on fluorescence-activated cell sorting (FACS)-sorted fibroblasts from multiple mouse tissues15. These data identified regions of open chromatin and transcriptional networks driven by tissue type, similar to recent reports15,16 (Extended Data Figs. 1, 2, Supplementary Tables 1, 2). However, bulk sequencing cannot discriminate gene signatures that represent a single, homogeneous cell population from those that reflect the average of heterogeneous populations. To resolve this issue, we collected mouse scRNA-seq datasets enriched for non-haematopoietic cells from our laboratory and from public repositories. We removed non-fibroblast cells and corrected for cross-laboratory batch effects to produce a fibroblast-specific single-cell atlas composed of 28 datasets across 16 unperturbed tissues (n = 120,583 cells; Fig. 1a, b, Extended Data Fig. 3a–c, Supplementary Table 3). An interactive data browser for the atlases is publicly available (see ‘Data availability’). Notably, our bulk RNA-seq and single-cell data were highly concordant, which indicates that our single-cell analytical approach did not introduce technical bias (Extended Data Fig. 3d).
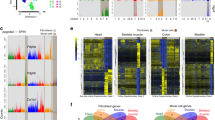
a, Twenty-eight datasets for steady-state fibroblast atlas from 16 mouse tissues. b, Uniform manifold approximation and projection (UMAP) embedding of 120,583 single cells in the steady-state atlas. Ten clusters identified through graph-based clustering are indicated by colour. c, Heat map of the relative average expression of the most strongly enriched genes for each cluster (log(fold change) of one cluster versus all others, z-score by row) with key genes, functional annotation and tissue distributions listed. MSC, mesenchymal lineage cell; OLC, osteolineage cell. d, Pseudotime(s) visualized using principal curves representing trajectories of fibroblast differentiation across steady-state atlas with Pi16+ cluster set as root.
In the steady-state atlas, ten clusters were identified on the basis of differential gene expression (Fig. 1b, c). The exact number of clusters in analyses of this type is data-driven but still somewhat subjective; some clusters were well-separated from others, while in other instances it is likely that an expression continuum, with intermediate states, exists. We identified more than 200 differentially expressed genes (DEGs) for each cluster and annotated clusters according to the dominant cluster-specific gene: Pi16+, Col15a1+, Ccl19+, Coch+, Comp+, Cxcl12+, Fbln1+, Bmp4+, Npnt+ and Hhip+ (Supplementary Table 4, Extended Data Fig. 3e). Known fibroblast-associated genes showed distinct expression across the clusters, confirming heterogeneity within the fibroblast lineage (Extended Data Fig. 3f). We were able to use tissue-distribution patterns and hallmark genes to ascribe functional identities to most clusters, including Ccl19+ fibroblastic reticular cells (FRCs)3, Coch+ red pulp fibroblasts17, Cxcl12+ mesenchymal stromal cells and osteolineage cells18, Fbln1+ and Bmp4+ intestinal fibroblasts19, Comp+ fibroblasts20, Npnt+ alveolar fibroblasts and Hhip+ peribronchial fibroblasts21 (Fig. 1c, Extended Data Figs. 3g, 4a–p). The specialization of these clusters was reflected in differential enrichment of gene expression in core signalling pathways, including NFκB and TNF in the Ccl19+ cluster and WNT signalling in the Fbln1+ and Bmp4+ clusters19 (Extended Data Fig. 3h).
Notably, nearly all tissues contributed to the Pi16+ and Col15a1+ clusters, which suggests that these clusters are universal (Fig. 1c, Extended Data Fig. 3g). Genes that defined these two clusters differentiated fibroblasts from mesothelial cells in bulk RNA-seq data (Extended Data Fig. 4r–t). DEGs in the Pi16+ cluster (Pi16, Dpp4 and Ly6c1) suggested an identity similar to adventitial stromal cells22, which are found in vascular niches and can produce ECMs and acquire gene expression profiles consistent with specialized fibroblasts23,24. The Col15a1+ cluster exhibited an association with the basement membrane, evidenced by expression of Col4a1, Hspg2 and Col15a1 (Extended Data Fig. 3i, Supplementary Table 4). The ubiquity of the universal Pi16+ and Col15a1+ subtypes across tissues and the elevated level of stemness-associated genes (Cd34 and Ly6a (which encodes SCA1), Extended Data Fig. 3j), led us to investigate the potential for a developmental relationship among clusters. Slingshot lineage inference identified trajectories that emerged from the Pi16+ cluster, passed through the Col15a1+ cluster, and ended at specialized clusters (Fig. 1d).
Collectively, our analysis showed that in steady-state mouse tissues, universal (Pi16+ and Col15a1+) and specialized fibroblast subtypes exist, and that these may be developmentally linked. The roles of universal fibroblasts include ECM secretion, with Col15a1+ universal fibroblasts exhibiting the capacity to secrete basement membrane proteins and the Pi16+ subtype potentially serving as a resource cell that can develop into specialized fibroblasts.
Dpt IRESCreERT2 mouse validates scRNA-seq
To validate our scRNA-seq analysis, we used the surface markers SCA1 and LY6C (encoded by Ly6c1) to distinguish Pi16+ (LY6C+SCA1+) and Col15a1+ (LY6C−SCA1+) universal fibroblasts from specialized fibroblasts (LY6C−SCA1−; Extended Data Figs. 5a, 6a). Flow cytometry revealed that PDGFRα+ fibroblasts could be sorted into these three groups across 11 tissues (Extended Data Fig. 6b, c). This approach confirmed the existence of bona fide universal and specialized phenotypes: expression of Pi16 was enriched in LY6C+SCA1+ fibroblasts, whereas markers of more specialized fibroblasts such as Ccl19 (lymph node) and Npnt (lung) were enriched in LY6C− fibroblasts. Expression of dermatopontin (Dpt) was inversely correlated with specialization (Extended Data Fig. 6d, e). Dpt was diffusely expressed across the steady-state fibroblast atlas but was significantly enriched in Pi16+ and Col15a1+ universal fibroblasts, with highest expression in the Pi16+ cluster. The intestine-specific Fbln1+ cluster also showed Dpt expression at a level similar to the Col15a1+ cluster (Extended Data Fig. 6e, Supplementary Table 4). Expression of Dpt and Pi16 RNA in lung and small intestine was assayed histologically using RNAscope. Dpt+Pi16+ cells were enriched near vascular structures in both tissues, as expected22. Dpt+Pi16− cells were observed in the lung parenchyma, in alignment with our predicted distribution of Col15a1+ cells, and at the base of the villi in the small intestine, consistent with a lack of functional specialization19 (Extended Data Fig. 6f).
We generated a genetically modified mouse model for tracking Dpt-expressing cells to test whether Dpt preferentially marked universal fibroblasts. An IresCreERT2 cassette, which requires the oestrogen receptor modulator tamoxifen for activity, was inserted downstream of the stop codon in exon 4 of Dpt. We crossed this strain with the Rosa26LSLYFP mouse line. The resulting DptIresCreERT2Rosa26LSLYFP mouse irreversibly marks DptIresCreERT2-positive cells and their progeny with yellow fluorescent protein (YFP) after tamoxifen administration (Extended Data Fig. 6g).
DptIresCreERT2Rosa26LSLYFP mice were given tamoxifen in their chow for 14 days. In the 11 tissues examined, PDGFRα+ fibroblasts robustly expressed YFP (Fig. 2a, Extended Data Fig. 6h). Recombination efficiency varied across tissues: lymph nodes exhibited the lowest proportion of YFP+ fibroblasts (28.7 ± 5.5% (mean ± s.e.m.)) whereas the heart exhibited the highest (83.7 ± 3.1%; Fig. 2b). Other cell types expressed little or no YFP (Extended Data Fig. 6i, j). FACS-sorted YFP-positive fibroblasts (sorted 14 days after DptIresCreERT2 recombination) expressed higher levels of Pi16 and Col15a1 and lower levels of Ccl19 and Npnt in the lymph node and lung, respectively, than YFP-negative fibroblasts. This suggests that Dpt-expressing cells are transcriptionally distinct from lymph node FRCs and lung alveolar fibroblasts (Extended Data Fig. 6k). YFP+ and YFP− fibroblasts expressed equivalent levels of Csf1, suggesting that both Dpt+ and Dpt− fibroblasts are involved in macrophage homeostasis25 (Extended Data Fig. 6l).
a, Representative gating for YFP (Dpt) in pancreatic fibroblasts (live, EpCAM−CD45−CD31−PDPN+PDGFRα+). Red, DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP; grey, DptIRESCreERT2wt/wtRosa26LSLYFPwt/loxP mouse. Horizontal bar denotes the percentage of positive cells. b, Quantification of YFP+ (Dpt+) cells. Frequency (blue dots with mean, top x-axis) and cell number per milligram tissue (red boxplots, bottom x-axis). Percentage YFP+ determined by subtracting fluorescence in DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP mice from fluorescence in DptIRESCreERT2wt/wtRosa26LSLYFPwt/loxP mice. c, YFP expression in LY6C+SCA1+, LY6C−SCA1+ and SCA1− fibroblasts from the lymph node in Ccl19YFP mice. d, YFP expression in LY6C+SCA1+, LY6C−SCA1+ and SCA1− fibroblasts from lymph node in Grem1CreERT2wt/ki Rosa26LSLYFPwt/loxP mice treated with tamoxifen via intraperitoneal injection for 5 consecutive days, assayed on days 14–16 after first injection. Bars in c, d represent mean. e, YFP expression in LY6C+SCA1+, LY6C−SCA1+ and SCA1− fibroblasts from lymph node, omentum, lung, inguinal adipose, pancreas and skin (flank) in DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP animals. Mean ± s.e.m. n = 4 (b) or 3 (c–e) or representative of 4 (a) biologically independent experiments with 4 (b), 7 (c), 10 (d) or 3 (e) mice. In b, centre lines in boxplots show median; box spans interquartile range, whiskers denote10th–90th percentile. Each dot represents one mouse; mice received tamoxifen chow for 14 days and were analysed on days 14–16 (b–d). Subcut., subcutaneous adipose.
The distinction between universal and specialized fibroblasts was also observed at the protein level via YFP expression in Ccl19YFP (ref. 26) and Gremlin1CreERT2Rosa26LSLYFP mice27. Here, the fraction of YFP-expressing cells (denoting an FRC phenotype) increased from LY6C+SCA1+ (Pi16+) fibroblasts to LY6C−SCA1+ (Col15a1+) fibroblasts and further to SCA1− (specialized) fibroblasts (Fig. 2c, d, Extended Data Fig. 5b, c). YFP expression in DptIresCreERT2;Rosa26LSLYFP mice showed the opposite trend in lymph node, lung, and other tissues (Fig. 2e, Extended Data Fig. 5d). This evidence suggested that Dpt expression encompassed both of the universal fibroblast subtypes identified by scRNA-seq; therefore, we refer to these as Dpt+Pi16+ and Dpt+Col15a1+ universal fibroblasts.
Fibroblasts in perturbed mouse tissues
We next investigated how fibroblasts are affected by infection, injury, cancer, fibrosis, metabolic changes and arthritis (Fig. 3a). We integrated 17 publicly available scRNA-seq datasets across 13 tissues to generate a perturbed-state fibroblast atlas (n = 99,596 cells; Fig. 3b, c, Extended Data Fig. 7a–c, Supplementary Table 3). This approach showed ten clusters: Pi16+, Col15a1+, Ccl19+, Cxcl12+, Comp+, Npnt+, Hhip+, Adamdec1+, Cxcl5+ and Lrrc15+ (Fig. 3b, c, Extended Data Fig. 7d, Supplementary Table 5). Clusters in the perturbed-state atlas displayed heterogeneous expression of common fibroblast-associated genes (Extended Data Fig. 7e). In nearly all perturbed tissues and for all types of inflammation, some fibroblasts occupied the universal Pi16+ and Col15a1+ clusters, and these clusters expressed the highest levels of Dpt (Extended Data Figs. 7f, 8a–o, r). The Cxcl12+, Ccl19+, Comp+, Npnt+, and Hhip+ clusters were similar to the analogous steady-state clusters with respect to gene expression and tissue distribution (Fig. 3b, c, Extended Data Fig. 7g). Conversely, the Cxcl5+, Adamdec1+ and Lrrc15+ clusters appeared to represent perturbation-specific, activated fibroblast states that were not observed in the steady-state atlas.
a, Seventeen datasets for perturbed atlas from 13 perturbed mouse tissues. b, UMAP embedding of 99,596 cells in the perturbed-state atlas. Ten clusters identified through graph-based clustering are indicated by colour. Coloured clusters are universal (bold) or perturbation-specific; grey clusters have steady-state analogues. c, Heat map of the relative average expression of the most strongly enriched genes for each cluster (log(fold change) of one cluster versus all others, z-score by row). d, Representative flow cytometry plots showing frequency of YFP+ cells in LRRC15+ fibroblasts from KPR3070 subcutaneous tumour at day 21 post-inoculation in DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP mouse (representative of two biologically independent experiments).
Fibroblasts from early muscle injury contributed the majority of cells to the Cxcl5+ cluster and expressed chemokine genes such as Ccl2 and Ccl7 (Fig. 3c, Extended Data Fig. 8j, p). Analysis of co-regulated genes suggested that this cluster was driven by PI3K, TNF and NFκB signalling (Extended Data Fig. 8v). Cells in the Adamdec1+ cluster were from colitis and showed upregulation of Il11 and Grem1 (Extended Data Figs. 7f, 8e, q). Gene expression in these cells was associated with MAPK signalling (Extended Data Fig. 8v). We subjected mice to DSS-induced colitis and found increased expression of Grem1 in lesions as compared to healthy areas of the colon by RNAscope, validating the colitis-specific expression of Grem1 in the mouse perturbed-state atlas (Extended Data Fig. 8w). The Lrrc15+ cluster was composed of cells from arthritis, skin wound, fibrosis and small and large pancreatic ductal adenocarcinoma (PDAC; Extended Data Figs. 7f, 8a–o). This cluster showed high expression of Cthrc1, Acta2, Postn and Adam12, and of collagens, which suggests that these cells represented myofibroblasts (Extended Data Fig. 8p–u). The increase in collagens and TGFβ signalling is consistent with previous reports10 (Extended Data Fig. 8v).
In perturbed tissues, universal Dpt+Pi16+ fibroblasts maintained the highest expression of stemness-associated genes (Extended Data Fig. 8x). Lineage inference identified trajectories from Dpt+Pi16+ through Dpt+Col15a1+ and then on to perturbation-specific, activated Cxcl5+ and Lrrc15+ clusters or the Adamdec1+ cluster (Extended Data Fig. 8y). We tested whether universal fibroblasts give rise to LRRC15+ myofibroblasts using a subcutaneous tumour model in the DptIresCreERT2;Rosa26LSLYFP mouse. We found that 52 ± 7% of LRRC15+ myofibroblasts were YFP+ in DptIresCreERT2ki/ki mice (Fig. 3d, Extended Data Fig. 8z–b′). This indicates that Dpt-expressing cells marked before tumour implantation can differentiate into LRRC15+ myofibroblasts.
Single-cell RNA-seq of human fibroblasts
We hypothesized that mice may exhibit some parity to humans in terms of steady- and perturbed-state fibroblast subtypes. We performed scRNA-seq on tumour and normal adjacent tissue (NAT) samples from three patients with pancreatic cancer and identified two subsets of fibroblasts (n = 21,262 cells; Extended Data Fig. 9a–c, Supplementary Tables 6, 7). Clusters c3 and c8 were annotated as cancer-associated fibroblasts (CAFs) and normal fibroblasts, respectively. Twelve of the 20 most upregulated genes in c8 were significantly upregulated in mouse steady-state Dpt+Pi16+ (DPT, IGFBP5, IGFBP6, C3, APOD) or Dpt+Col15a1+ (CXCL12, SMOC2, C7, FBLN5, MFAP4, LUM, FMO2) clusters (Supplementary Tables 4, 7). These 20 upregulated genes defined our human universal fibroblast expression module. To test whether c8 represented universal fibroblasts in humans, we inferred the abundance of these cells across tissues from the GTEx database (n = 5,961 samples, Extended Data Fig. 9d, e). We found strong (r > 0.5) co-expression of universal module genes in 12 human tissues, including normal pancreas (Extended Data Fig. 9f, g). At the single-cell level, the human universal module was observed in c8 pancreas and human adipose tissue (Extended Data Fig. 9h, i). We next scored samples from 122 patients with pancreatic cancer28 for the human universal module as well as a human fibroblast activation program (20 most enriched genes in c3 versus c8; Extended Data Fig. 10a). There was a strong negative correlation between the universal and activated gene programs (r = −0.54), which suggests that activation in human fibroblasts may be associated with loss of universal fibroblast gene expression (Extended Data Fig. 10b), reminiscent of the lineage relationship we observed in mice.
We investigated whether the transcriptional fibroblast subtypes we observed in humans had mouse orthologues and observed that the human c3 signature was enriched in the mouse Lrrc15+ myofibroblast cluster (Extended Data Fig. 10c). This was not restricted to pancreatic cancer14: fibroblast signatures from human rheumatoid arthritis (RA)7,9, interstitial lung diseases29, IPF30 and ulcerative colitis31 also showed localized gene expression in the mouse Lrrc15+ myofibroblast cluster (Extended Data Fig. 10c, Supplementary Table 8). We found evidence of universal fibroblasts in samples from patients with interstitial lung disease29 and ulcerative colitis31 (Extended Data Fig. 10d). Cells described as inflammatory fibroblasts across human samples and indications predominantly aligned with the universal clusters of the mouse perturbed-state atlas7,9,11 (Extended Data Fig. 10e). We also observed that mouse IL-1+ CAFs10 localized to universal clusters in the mouse perturbed-state atlas (Extended Data Fig. 10f), suggesting that cells described as inflammatory fibroblasts may represent universal fibroblasts that have not undergone sufficient transcriptional change to constitute a new cell state.
Last, we composed a human perturbed-state fibroblast atlas by integrating c3 PDAC CAFs, colon fibroblasts from patients with colitis and lung fibroblasts from individuals with non-small cell lung cancer (NSCLC), IPF or COVID-19 (Fig. 4a). This human perturbed-state atlas (n = 10,355 cells) exhibited six clusters (Fig. 4b, c, Supplementary Table 9). We observed expected clusters, including NPNT+ alveolar fibroblasts21 derived from individuals with lung disease, ADAMDEC1+ and CCL19+31 clusters primarily from colitis samples, and a PI16+ cluster, in which the human universal signature was most enriched (Fig. 4b, c, Extended Data Fig. 10g). We observed two myofibroblast clusters defined by LRRC15+ and COL3A1+ expression. LRRC15+ myofibroblasts were enriched in cells from individuals with pancreatic and lung cancer, whereas the COL3A1+ cluster was enriched in cells from patients with COVID-19 (Fig. 4d). Both myofibroblast subsets expressed high levels of collagens and ECM-modifying genes, including CTHRC1 (Extended Data Fig. 10h, i). The human perturbed-state atlas validated aspects of the mouse perturbed-state atlas, including LRRC15+ myofibroblasts and ADAMDEC1+ fibroblasts, but also suggested that indication-specific stimuli or signal duration may drive additional myofibroblast populations in humans that were not seen in our mouse atlas.
a, Datasets for perturbation atlas from three perturbed human tissues. b, UMAP embedding of 10,355 cells in the perturbed-state human fibroblast atlas. Six clusters identified through graph-based clustering are indicated by colour. c, Heat map of relative average expression of the most strongly enriched genes for each cluster (log(fold change) of one cluster versus all others, z-score by row). d, Relative abundance of each disease in UMAP clusters.
Discussion
Fibroblasts have emerged as nexus cells that define the architecture of tissues, augment the function and positioning of other cell types, and have key roles in many diseases. We initially hypothesized that fibroblasts accomplish these diverse tasks through tissue-specific transcriptional programming. While our initial hypothesis was supported by bulk sequencing modalities, scRNA-seq across mouse tissues and perturbations led to a different conclusion—that the fibroblast lineage was compartmentalized into universal and specialized (steady-state) or activated (perturbed-state) subtypes. It is possible that fibroblast subsets may exhibit additional imprinting by their tissue of residence. Our scRNA-seq observations were supported experimentally by the DptIresCreERT2 knock-in mouse. Our data suggest a paradigm in which Dpt+ universal fibroblasts give rise to distinct subsets of fibroblasts across tissues, enabling this cell lineage to give rise to phenotypes that can execute the myriad functions ascribed to fibroblasts in the steady-state and in disease. In this paradigm, diverse tissues harbour Dpt+ universal fibroblasts that can differentiate into activated fibroblasts during inflammation and, we speculate, into specialized fibroblasts during development. In this way, Dpt+ universal fibroblasts operate as resource cells, providing functional plasticity to the fibroblast lineage.
Our data suggest concordance between fibroblast phenotypes in mice and humans. We confirmed that a subset of fibroblasts in human tissues has transcriptional similarities to mouse universal fibroblasts. Examination of human datasets representing an array of diseases identified mouse fibroblast orthologues, suggesting that our mouse perturbed-state atlas provides a basis for understanding fibroblast subtypes in human disease states. However, we observed a unique myofibroblast population that was enriched in patients with COVID-19, relative to the other human indications we analysed, which suggests that important inter-species or perturbation-specific differences may exist as well.
Our approach to understanding fibroblasts across tissues has identified both broad similarities and essential differences. Exposing the contours of fibroblast gene expression across tissues and activation states may help to clarify fibroblast subtyping and nomenclature. Open questions still remain about the spatial dynamics among fibroblast subtypes, the existence of other subtypes not captured in these datasets, and the structural or immune cells that promote specialization in the steady-state or activation during inflammation. It remains unclear why two universal Dpt+ fibroblast subtypes exist, though we speculate that this may represent a necessary division of labour within the lineage. Overall, the organizing principles described here may enable a deeper understanding of the development, evolution and behaviour of universal, specialized and activated fibroblasts, which may in turn yield dividends for human medicine.
Methods
Mice
Wild-type mice were obtained from Jackson Laboratory (JAX; colony 00064) and maintained at Genentech. Ccl19YFP (ref. 32), Grem1CreERT2 (ref. 27) and Rosa26LSLYFP mice were bred at Genentech. DptIresCreERT2 mice were designed, generated and bred at Genentech. Male and female mice aged 6–12 weeks were used for all studies. For tamoxifen-induced cre expression, mice were injected with 2 mg tamoxifen (Sigma, cat. T5648) diluted in sunflower seed oil (Sigma, cat. 88921) for 5 consecutive days intraperitoneally or were fed chow containing tamoxifen (Envigo, cat. TD.130859). All experiments were performed under protocols approved by the Institutional Animal Care and Use Committee at Genentech. Other genotypes listed in Supplementary Table 3 were not bred in our facility but used in the integrated single-cell RNA-seq fibroblast atlases. No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Mouse tissue digestion and stromal cell isolation or identification by FACS
Tissues were isolated and fibroblasts and mesothelial cells were isolated as previously described15. In brief, tissues were obtained and minced, aside from the LN, omentum (neither minced) and bone (decapped, marrow removed and crushed). To isolate flank skin, hair was shaved, adipose tissue was removed and tissue was minced. Next, tissues were placed in a 15-ml conical tube with 5 ml digestion medium (RPMI + 2% FBS with 100 mg/ml Dispase (Life Tech., cat. 17105041), 100–200 mg/ml collagenase P (Roche, cat. 11249002001), and 50 mg/ml DNase I (Roche, cat. 10104159001)) and agitated. Tubes were placed in a 37 °C water bath for 15 min, and 5-ml fractions were removed and filtered (70 μm) into RPMI supplemented with 2% FCS (VWR) three times. For skin fibroblast scRNA-seq, skin was minced and digested in the enzymatic cocktail described above for 60 min. We used 200mg/ml collagenase P to isolate cells from dense tissues such as the spleen, liver, inguinal adipose, brown adipose and pancreas. After a single-cell suspension was obtained, the cells were layered on top of a 26% optiprep (Sigma, cat. D1556; diluted in phosphate buffered saline (PBS)) gradient in 15-ml conical tubes and spun at 1,500g for 15 min with slow acceleration and the brake off. Cells in suspension were isolated with a transfer pipette. After digestion, the preparations were incubated with Ack for 2–5 min to remove red blood cells.
Cells were labelled with the following monoclonal antibodies purchased from eBioscience, BioLegend, or BD Biosciences at 1:200 for 20–30 min, unless otherwise noted. Prior to cell surface staining with the following fluorescently labelled antibodies, cells were blocked with Fc block (2.4G2; 1:500–1:1,000, cat #553142). Surface staining for experiments was performed as below, unless otherwise noted: CD45 (30-F11, cat. 564279), EPCAM (G8.8, cat. 118216), CD31 (390, cat. 612802 or 102524), PDGFRα (AP5, cat. 135906), PDPN (8.1.1; 1:800, cat. 127410), LY6C (HK1.4; 1:200, cat. 45-5932-82), SCA1 (D7; 1:200, cat. 108131), CD24 (M1/69; 1:200, cat. 612832) or LRRC1510 (1:200, in-house). Live cells were identified by washing after Fc block and incubation with Fixable Viability Dye Violet (Invitrogen, cat. L34955, 1:1,000) before surface staining or incubation with calcein blue (Invitrogen, cat. C1429, 1:1,000) after surface staining. Data were acquired on a Fortessa, Symphony or LSRII (BD Biosciences) and analysed using FlowJo (Tree Star, v9.9.6) or cells were sorted on a Fusion or Aria (BD Biosciences). Data were presented using Prism (Graphpad, v9).
qPCR, RNA extraction and cDNA synthesis
For each qPCR experiment, organs from 3–5 mice were pooled, digested and sorted by FACS. Total RNA was extracted from FACS-sorted cells using a QIACube HT with the RNeasy Plus Mini Kit (QIAGEN, cat. 74134). cDNA were synthesized using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, cat. 4368814) with an RNase Inhibitor (Applied Biosystems, cat. N8080119). Each cDNA sample was diluted 1:200–1:300 in RNase-free water before use in qPCR. qPCR was conducted using TaqMan Gene Expression Assay Probes for the genes Pi16 (Mm00470084_m1), Col15a1 (Mm00456551_m1), Dpt (Mm01273496_m1), Npnt (Mm00473794_m1), Ccl19 (Mm00839967_g1), Csf1 (Mm00432686_m1) and Gapdh (Mm99999915_g1). Each TaqMan probe was diluted 1:10 in TaqMan Fast Advanced Master Mix (ThermoFisher, cat. 4444557) to create a TaqMan probe working solution. All qPCR reactions were carried out in a MicroAmp optical 384-well reaction plate. qPCR was performed using the QuantStudio 5 (Applied Biosystems) under the following cycling conditions: 1 cycle of 50 °C for 2 min and 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. The relative expression values were determined by: 1) subtracting the average Ct value of the gene of interest (GOI) from Gapdh (Gapdh − GOI) for each sample and 2) raising 2 to the power of (Gapdh − GOI). Data were presented using Prism (Graphpad, v9). Data points were called non-detectable if Ct values were over 36 or if replicates had a standard deviation greater than 1.
Mouse scRNA-seq and cell hashing
Cells hashtagged with unique barcoded antibodies (Biolegend) were processed using Chromium Single Cell Gene Expression 3′ v3 Library and Gel Bead Kit following the manufacturer’s instructions (10X Genomics, Cat. PN-1000075). Cells were counted and checked for viability using Vi-CELL XR cell counter (Beckman Coulter), and then injected into microfluidic chips to form Gel Beads-in-Emulsion (GEMs) in the 10X Chromium instrument. Reverse transcription (RT) was performed on the GEMs, and RT products were purified and amplified. DNA from antibody-derived tags (ADTs) was separated from cDNA based on size selection using SPRIselect beads (Beckman Coulter, cat. B23318). Expression libraries and ADT libraries were generated and profiled using the Bioanalyzer High Sensitivity DNA kit (Agilent Technologies, cat. 5067-4626) and quantified with Kapa Library Quantification Kit (Roche, cat. 07960255001). Illumina HiSeq and NovaSeq (Illumina) were used to sequence the libraries.
Tissue processing for histology
The small intestine or colon was removed, flushed with 10% neutral buffered formalin, and then cut into 1-inch-long strips from the duodenum, jejunum, ileum, and colon. The lung was removed and flushed with 10% neutral buffered formalin to inflate the tissue. A suture was then tied to the trachea to ensure the tissue stayed inflated while fixing in 10% neutral buffered formalin for 24 h. After fixation, the samples were transferred to 70% ethanol for 24 h and then processed for paraffin embedding. Once embedded, the blocks were trimmed on a Leica RM-2245 microtome until the centre of tissue was reached. Five-micrometre-thick sections were then collected for downstream analysis.
RNAscope in situ hybridization
In situ hybridization (ISH) was performed using reagents and protocols from Advanced Cell Diagnostics (ACD). After sectioning, the slides were allowed to dry for 24 h before baking in an oven at 60 °C for 1 h. Sections were then rehydrated in two washes of xylene for 5 min each followed by two washes in 100% ethanol for 1 min each. After rehydration, endogenous peroxidase activity was quenched with hydrogen peroxide before the sections were boiled in antigen retrieval buffer (ACD, cat. 322500) for 15 min. After antigen retrieval, the sections were digested with proteinase (ACD, cat. 322330) for 25 min at 40 °C. After digestion, slides were washed twice for 1 min with ISH wash buffer (ACD, cat. 310091) then hybridized with probes of interest (Grem1-C1, cat. 314741; Dpt-C1 probe, 451311; Pi16-C1 and C2 probe; 451311 (C1) and 451318 (C2)) for 2 h at 40 °C. After hybridization, amplification steps were completed according to the ACD protocol. After the final amplification incubation, signal was detected with HRP-conjugated DAB (ACD, cat. 322360 and 322500), counterstained with haematoxylin for 30 s, then baked in an oven at 60 °C for 15 min before mounting with non-aqueous mounting medium.
DSS-induced colitis
We added 3% (w/w) colitis-grade DSS (MP Biosciences, cat. 9011-18-1) to the drinking water for 7 days and then euthanized the animals and processed tissues for histology. The water was weighed daily to determine the average water consumption per cage. Mouse bodyweights were recorded daily and mice that lost 20% or more of their body weights were euthanized following IACUC standards
Generation of Dpt IresCreERT2 knock-in mouse
Homologous recombination and mouse embryonic stem (ES) cell technology33,34,35 were used to generate a genetically modified mouse strain with a DptIresCreERT2 knock-in. A gene-targeting vector was constructed with a 1,928-bp 5′ arm of homology corresponding to GRCm38/mm10 chr1: 164,821,309–164,823,236 and a 1,810-bp arm of 3′ homology corresponding to chr1: 164,823,237–164,825,046. IRES-CREERT2-SV40-FRT-pgk-neo-FRT was inserted immediately after the STOP codon (TAG) of exon 4. The final vector was confirmed by DNA sequencing, linearized and used to target C2 (C57BL/6N) ES cells using standard methods (G418 positive and ganciclovir negative selection)36.
C57BL/6N C2 ES cells37 were electroporated with 20 μg linearized targeting vector DNA and cultured under drug selection essentially as described38. Positive clones were identified using long-range PCR followed by sequence confirmation. Correctly targeted ES cells were subjected to karyotyping. Euploid gene-targeted ES cell clones were treated with Adeno-FLP to remove PGK neomycin, ES cell clones were tested to identify clones with no copies of the PGK neomycin cassette, and the correct sequence of the targeted allele was verified. The presence of the Y chromosome was verified before microinjection into albino BALB/c embryos. Germline transmission was obtained after crossing the resulting chimaeras with C57BL/6N females. Genomic DNA from pups born was screened by long-range PCR to verify the desired gene targeted structure before mouse colony expansion. For genotyping, the following primers were used: 5′-CGCCAGTGGAAGTTCA -3′, 5′-TATAGGAACTTCGCTCGC -3′ and 5′-GTGCTGTGCAAGGAAG-3′ amplified 329-bp wild-type and 278-bp knock-in DNA fragments.
Tumour inoculation
Age-matched 6–10-week-old DptIresCreERT2wt/wtRosa26LSLYFP and DptIresCreERT2ki/kiRosa26LSLYFP mice received tamoxifen injections for 5 days, rested for 13–14 days and were inoculated subcutaneously in the right unilateral flank with 1 × 105 KPR3070 tumour cells suspended in Hanks’s buffered saline solution and phenol-red-free Matrigel (Corning, cat. 356237). Tumours were removed 21 days after implantation.
Human patient information
Pancreatic cancer sample collection was approved by the Ethics Committee of Beijing Cancer Hospital. All patients in this study provided written informed consent.
Human tissue digestion and stromal cell isolation
Samples were obtained and sequenced by Analytical Biosciences. Single cells were dissociated from tumour and adjacent non-cancer tissues as described previously39. In brief, tumours and adjacent non-cancer tissues were cut into approximately 1–2-mm3 pieces in RPMI-1640 medium (Gibco) and enzymatically digested with gentleMACS (Miltenyi) for 60 min on a rotor at 37 °C, according to the manufacturer’s instructions. The dissociated cells were subsequently passed through a 100-μm SmartStrainer and centrifuged at 400g for 5 min. After the supernatant was removed, the pelleted cells were suspended in red blood cell lysis buffer (TIANDZ) and incubated on ice for 1–2 min to lyse red blood cells. After being washed twice with 1 × PBS (Gibco), the cell pellets were re-suspended in sorting buffer (PBS supplemented with 1% fetal bovine serum (FBS, Gibco)).
Single-cell suspensions were stained with antibodies against CD45 and 7AAD for FACS sorting, performed on a BD Aria SORP instrument. Based on FACS analysis, single cells were sorted into 1.5-ml tubes (Eppendorf) and counted manually under the microscope. The concentration of single cell suspensions was adjusted to 500–1,200 cells per μl. Cells were loaded at between 7,000 and 15,000 cells per chip position using the 10X Chromium Single cell 5′ Library, Gel Bead & Multiplex Kit and Chip Kit (10X Genomics, V1.0 barcoding chemistry) according to the manufacturer’s instructions. All the subsequent steps were performed following the standard manufacturer’s protocols. Purified libraries were analysed using an Illumina Hiseq X Ten sequencer with 150-bp paired-end reads.
Mouse bulk RNA-seq analysis
For ex vivo bulk RNA-seq, cells were isolated and stained as described above. Each tissue was represented by 2–3 individual replicates that were each derived by pooling tissues from 3–5 mice and FACS sorting cells directly into Trizol (Invitrogen, cat. 15596026). In some cases, lysed cells from at least three independent experiments were pooled for one replicate. In total, RNA was generated from an average of 35,195 ± 7,357 (mean ± s.e.m.) fibroblasts and 17,318 ± 7,618 mesothelial cells. RNA was isolated as described40 or at Expression Analysis, Inc.
Paired-end RNA-seq libraries were constructed from at least 747 pg of RNA using the SMART-Seq v4 ULTRA Low Input RNA Kit for Sequencing (Takara, cat. 634891) and NexteraXT kits (Illumina, cats. FC-131-1096 and FC-131-2001) for Low Input RNA Kits. Libraries were then sequenced on an Illumina HiSeq yielding, on average, 35 million read pairs (2 × 50 bp) per sample. Reads were aligned to the GENCODE basic mouse transcriptome index (M14) and transcript levels quantified using salmon with parameters --type quasi -k 25. Subsequently, counts were transformed into gene-level counts in R using the tximport (https://bioconductor.org/packages/release/bioc/html/tximport.html) package. Differential expression analysis taking batches into account was carried out on the gene by sample count matrix with DESeq241, using a design of ~0 + condition + batch having a coefficient for each level of condition. For principal component analysis (PCA), log-transformed normalized counts (lengthScaledTPM) were batch corrected using Combat42 and PCA was performed in the space of variable genes (coefficient of variation >0.3). Gene set enrichment analysis (GSEA) using the fgsea method43 was performed on genes ranked by their principal component 1 loadings using the top 20 marker genes for Pi16+ and Col15a1+ clusters from the steady-state fibroblast atlas.
Mouse bulk ATAC–seq analysis
For ex vivo bulk ATAC–seq, cells were isolated and stained as described above. Each tissue was represented by 2–4 individual replicates that were each derived by pooling tissues from 3–5 mice and FACS-sorting fibroblasts. On average, 28,455 cells (± 5,325 (s.e.m.)) were sorted per tissue. These cells were then frozen in Gibco Recovery Cell Culture Freezing Medium (ThermoFisher, cat. 12648010). The cells were then thawed in a 37 °C water bath, pelleted, washed with cold PBS, and tagmented as previously described44, with some modifications45. In brief, cell pellets were resuspended in lysis buffer, pelleted, and tagmented using the enzyme and buffer provided in the Nextera Library Prep Kit (Illumina, cat. FC-121-1031). Tagmented DNA was then purified using the MinElute PCR purification kit (Qiagen, cat. 28004), amplified with 10 cycles of PCR, and purified using Agencourt AMPure SPRI beads (Beckman Coulter, cat. A63882). The resulting material was quantified using the KAPA Library Quantification Kit for Illumina platforms (Roche, 07960255001), and sequenced with PE42 sequencing on the NextSeq 500 sequencer (Illumina), with 42-bp paired-end reads. Library preparation and sequencing was performed by ActiveMotif, Inc.
Reads were aligned to the GRCm38/mm10 build of the mouse genome using GSNAP46 with parameters -M 2 -n 10 -B 2 -i 1 --pairmax-dna = 1000 --terminal-threshold = 1000 --gmap-mode = none --clip-overlap. Read pairs that aligned concordantly and uniquely to a single genomic location were retained for downstream analysis. PCR duplicates were removed using Picard MarkDuplicates (http://broadinstitute.github.io/picard/). Library depth-corrected coverage bigwig files were obtained to visualize the regions of interest.
OCR identification
Open chromatin regions (OCRs) were identified as peaks on individual replicates and pooled samples combining the replicates of a given tissue using MACS247, with parameters macs2 callpeak -f BAM --call-summits --nomodel–shift -95 --extsize 199 --keep-dup all -p 0.1 --call-summits (these choices of the shift and extsize parameters correct for the +5/−4 bp transposase insertion offset). The irreproducible discovery rate (IDR) pipeline48 was used to assess peak concordance between the individual replicates of a given tissue, and these IDR estimates were subsequently appended to the associated pooled peaks. Robust peaks per tissue were defined as pooled peaks that overlapped at least 50% of a peak from at least two individual replicates and that passed an IDR threshold of 0.1. All robust peaks across all tissues were first centred on their summits (summit ± 199 bp) and then those that overlapped mitochondrial and noncanonical chromosomes were removed. Finally, all remaining peaks were merged to obtain the final set of all accessible regions (n = 207,803). Per sample, reads that overlapped each region in the atlas were counted using the bedtools command multiBamCov49. To find tissue-specific OCRs, differential accessibility analysis was conducted on the count matrix using DESeq241, where the accessibility (that is, overlapping read count) of a given region in each tissue was compared against the count for that region in all other tissues. In this setting, the tissue-specific OCRs were defined according to the following criteria: log2(fold change) ≥ 2, adjusted P value ≤ 0.01. In addition, for each tissue a nondifferential/insignificant OCR set was defined according to: −0.585 ≤ log2(fold change) ≤ 0.585, q value > 0.05.
Motif enrichment analysis
For motif enrichment analysis, for each tissue-specific OCR set, an equally sized matched background set was selected on the basis of region length and GC content from among the nondifferential/insignificant OCRs, using MatchIt50. AME51 from the MEME suite was used with default settings to assess the enrichment of a set of 321 position weight matrices (PWMs) from Homer (http://homer.ucsd.edu/homer/) in the tissue-specific OCR sets versus the background sets. Specifically, Fisher’s exact test was used to compare the number of matches to a given PWM in the specific set versus the background set, and to assess statistical significance. Enriched PWMs were reported based on an adjusted P value threshold of 0.05.
ATAC–seq and RNA-seq concordance
To compute the correlation of log2(fold changes) inferred from the ATAC–seq and RNA-seq differential analyses, the ATAC–seq final atlas peaks were assigned to the gene with the closest transcription start site (TSS), using Gencode mouse M14 annotations and a distance threshold of 50 kb. Following the assignment, genes and atlas peaks with absolute log2(fold change) ≥ 1 and q value ≤ 0.05 in a given tissue were used in the correlation calculation.
An additional analysis to infer concordance between ATAC–seq and RNA-seq datasets was the BETA52 analysis, which takes a set of peaks (tissue-specific OCRs from ATAC–seq) and differential gene expression results from RNA-seq. In short, BETA calculates a regulatory potential score based on the number of peaks in a fixed window (100 kb by default) around each gene TSS and ranks the genes on the basis of this score. For each top gene set based on that rank, it calculates the percentage of the total up- and downregulated genes, as well as unregulated background genes, to provide P values for the overall up- or downregulation potential of the whole peak set. BETA was used with parameters -k BSF -g mm10 -n basic --df 0.1, for all pairwise tissue combinations, so for both matching and non-matching tissues.
Mouse scRNA-seq meta-analysis
The steady-state fibroblast atlas was composed of 28 datasets10,18,19,20,21,31,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70 and the perturbed-state fibroblast atlas was composed of 17 datasets10,18,21,31,53,54,55,56,57,58,61,69,70,71. Integrated fibroblast atlases at steady and perturbed states were generated and analysed using the following steps: (1) processing and filtering individual scRNA-seq datasets from healthy and diseased tissues; (2) integrating healthy and diseased datasets separately to generate steady- and perturbed-state atlases; (3) clustering and annotation; and (4) trajectory inference. The aforementioned steps are described in detail in the following sections.
(1) Processing and filtering individual scRNA-seq datasets
Single cell transcriptomics datasets, enriched in non-haematopoietic cells, generated using 10X Genomics and available as processed CellRanger files, were collected from public repositories and in-house lab datasets (Supplementary Table 3). For public datasets where processed files were not made available, we analysed raw data using cellranger count (CellRanger 2.1.0, 10X Genomics) using a custom reference package based on mouse reference genome GRCm38. A total of 32 scRNA-seq datasets representing multiple tissues and perturbations were analysed individually. In order to ensure comparability, for every individual dataset, we retained genes found in the Ensembl mouse (GRCm38) gene model, followed by implementing the Seurat single-cell analysis pipeline (version 3.9.9.9010)72,73 in R (version 4.0.0). Specifically, for each dataset we filtered low quality cells with <500 measured genes and a high percentage of mitochondrial contamination (>~5–20%, depending on the dataset). After filtering, data in each cell were normalized to log(CPM/100+1), the 2,000 most variable genes were identified, and the expression levels of these genes were scaled before performing PCA in variable gene space. Next, 20 principal components were used for graph-based clustering (resolution = 0.1) and UMAP dimensionality reduction was computed. All steps were performed using functions implemented in the Seurat package (NormalizeData, FindVariableFeatures, ScaleData, RunPCA, FindNeighbours, FindClusters, RunUMAP) with default parameters, except where mentioned. Cell clusters marked by the canonical marker gene for immune cells, Ptprc (Cd45), were discarded. All individual datasets devoid of Cd45+ cells were then used for integration to create two main atlases: (1) a steady-state fibroblast atlas comprising data from healthy tissues; and (2) a perturbed-state fibroblast atlas comprising data from diseased and inflamed tissues.
(2) Dataset integration for steady- and perturbed-state atlases
Before dataset integration, we imported the aforementioned filtered, non-processed Seurat objects (not scaled) of healthy and diseased datasets, and determined a common gene space by retaining only those genes that were measured across all datasets (21,087 genes). Next, individual healthy and diseased Seurat objects were merged separately into two different steady- and perturbed-state objects, respectively. Each of these merged objects was normalized (function NormalizeData, method = ‘LogNormalize’, scale.factor = 10,000), and scaled to regress out the stress gene signature (computed using Seurat’s AddModuleScore) of subpopulations affected by tissue dissociation methods74 before we performed PCA for the most variable genes. These processed, merged objects were next used for batch effect correction and integration using Harmony75 (version 1.0). We adjusted the diversity clustering penalty parameter, theta, to 1. We then provided the top 20 harmony dimensions as an input for UMAP and visualized the first two UMAP dimensions at a clustering resolution of 0.1 for the Cd45− atlases. Next, we identified distinct cell types using canonical marker genes such as Sparc, Col3a1, Dcn (fibroblasts), Epcam (epithelial cells), Alb (hepatocytes), Pecam1 (endothelial cells), Msln (mesothelial cells), Rgs5 (mural cells: pericytes), Myh11 (mural cells: smooth muscle cells), Top2a and Mki67 (proliferating cells), and Cd24a (remnant immune cell populations) (Extended Data Figs. 3a–c, 7a–c). The computational pipeline for integration was iterated twice to generate the fibroblast atlases.
Specifically, at each of the following steps non-relevant or unwanted cell types were filtered followed by recalculation of the variable gene space, batch-effect correction and clustering to generate fibroblast-specific atlas for steady- and perturbed state: (1) a Cd45− steady- and perturbed-state atlas comprising 180,922 and 182,609 cells, respectively, and (2) a fibroblast-specific steady- and perturbed-state atlas comprising 120,697 and 99,596 cells, respectively, after selecting for Dpt+Pdgfra+ fibroblast clusters from Cd45− atlases and discarding other stromal cells including endothelial and mesothelial cells, pericytes, smooth muscle cells and clusters with <1,000 cells.
(3) Clustering and annotation of steady- and perturbed-state fibroblast atlases
Each fibroblast atlas was constructed from its Cd45− counterpart, in which the top 40 harmony dimensions were provided as an input for UMAP and visualized at a clustering resolution of 0.2. Differential gene expression was computed using the Seurat function FindAllMarkers using a Wilcoxon rank sum test and corrected for multiple testing using the Bonferroni method.
Gene expression scores were computed using Seurat’s AddModuleScore function, visualized using VlnPlot or DotPlot. To determine markers for specialized or activated clusters relative to universal fibroblasts we used the Seurat function FindMarkers with default parameters. Next, we scored bulk tissue-specific signatures in the steady-state atlas, computed average scores per tissue signature across tissues represented in the steady-state atlas and visualized them using the ComplexHeatmap function Heatmap. We implemented the ClusterMap76 algorithm using default parameters to identify matching steady- and perturbed-state clusters. ClusterMap compares clusters among two or more datasets via hierarchical clustering based on the binary expression patterns of marker genes. Next, similarity scores of matched clusters are determined from the clustering results.
To infer the activity of signalling pathways that govern different fibroblastic cells at steady state and after perturbation, we implemented the Bioconductor package PROGENy (1.10.0)77. For both fibroblast atlases, we implemented the same strategy. First, we down-sampled each atlas using the Seurat function subset with parameters ‘WhichCells(object, downsample, seed = 1)’ followed by implementing the function progeny with default parameters ‘scale = TRUE, organism = “Mouse”, top = 100, perm = 1, return.assay = TRUE’. We then summarized the progeny scores by cell population and visualized them as a heatmap using the function pheatmap.
(4) Pseudotime reconstruction and trajectory inference
Single-cell pseudotime trajectories for both steady and perturbed state maps were computed using the algorithm slingshot (version 1.6.1), which enables computation of lineage structures in a low-dimensional space78. Specifically, slingshot was implemented in the analysis pipeline after dimensionality reduction and clustering of the integrated object. Each fibroblast object was first downsampled before trajectory inference. Pre-computed cell embeddings and clusters from the Seurat pipeline served as an input to the function slingshot (reducedDim = ‘UMAP’, clusterlabels = object$ RNA_snn_res.0.2, start.clus = “Pi16”, extend = 'n', stretch = 0). The start cluster was chosen using prior biological knowledge and the expression of genes such as Cd34 and Ly6a, known markers of progenitor-like cells. The wrapper function slingshot then performed lineage inference by treating clusters as nodes and constructing a minimum spanning tree (MST) between the nodes. Next, lineages or trajectories were defined by ordering clusters via tracing paths through the MST. Finally, individual pseudotime(s) were visualized using principal curves on the full fibroblast object.
Human scRNA-seq meta-analysis
A human perturbed-state fibroblast atlas was constructed using scRNA-seq datasets generated with the 10X technology, from disease indications such as PDAC, idiopathic pulmonary fibrosis(IPF)29, COVID-1979 and ulcerative colitis31 or SMART-seq2 technology in non-small cell lung carcinoma (NSCLC)80. Individual datasets were processed according to the description provided in ‘(1) Processing and filtering individual scRNA-seq datasets’, with the exception that LUM+ fibroblasts were retained per dataset before final integration. Specifically, for COVID-19 we used data from individuals who had died from severe COVID-19-associated pneumonia. Next, we down-sampled the COVID-19 dataset to 2,500 cells, in order to mitigate the effects of over-representation of a dataset due to high cell number contribution. We then aggregated all the datasets, chose a common gene space (~17,000) and implemented the same pipeline as described in the section ‘(2) Dataset integration for steady- and perturbed-state atlases’. We adjusted the diversity clustering penalty parameter, theta, to 1, provided the top 30 harmony dimensions as an input for UMAP, and visualized the first two UMAP dimensions at a clustering resolution of 0.2. Differential gene expression analyses and visualization of markers were performed using Seurat functions described above. Note, in this meta-analysis we used datasets generated using 10X and SMART-seq2 technologies across a common cell type, fibroblasts. This is because a benchmarking study demonstrated the superior performance of Harmony batch-effect correction using single-cell data generated on different platforms for an identical cell type81.
Bioinformatics data processing of human data
For the human pancreatic cancer single-cell data generated in our laboratory, raw sequencing data were transformed into FASTQ format with CellRanger’s (v2.1) mkfastq command, mapped to the human genome (GRCh38), and quantified with CellRanger count using default parameters. Quantified UMI count matrices from each patient were merged in R and analysed with the Seurat package (v 3.1.4). First, cells with <500 measured genes, or <2,700 UMIs, or >10% mitochondrial counts were removed from the dataset. In the resulting filtered dataset, data in each cell were normalized to log(CPM/100 + 1), the 2,000 most variable genes were identified, and the expression levels of these genes were scaled before PCA in the space of the most variable genes. Subsequently, 30 principal components were used for graph-based clustering (resolution = 0.1) and UMAP dimensionality reduction. All steps were performed with the methods implemented in the Seurat package (NormalizeData, FindVariableFeatures, ScaleData, RunPCA, FindNeighbours, FindClusters, RunUMAP) and default parameters, except for parameters mentioned above. Markers for each cluster were identified using the FindAllMarkers function, limiting the maximum number of cells per cluster to 1,000 for runtime improvement. Genes that were differentially expressed between clusters 3 and 8 were detected using the FindMarkers function and default parameters. To map human expression signatures onto the mouse perturbed state map, human gene symbols were translated to their mouse orthologues and an enrichment score for the gene signature was calculated using Seurat’s AddModuleScore function. Gene sets were identified within referenced papers (Supplementary Table 8).
Pseudo-bulk samples for co-expression analysis were generated from the human single-cell dataset using the following strategy: we randomly sampled 10% of cells from the pancreatic cancer single-cell dataset and pooled their reads into a bulk profile, which was subsequently normalized to log2(CPM). Using this strategy, we generated 100 bulk RNA-seq profiles with known proportions of cells from individual single-cell clusters. This allowed us to compare the expression of individual cluster 8 marker genes across pseudo-bulk samples both pairwise between genes and to the known cell type proportion of cluster 8 in the pseudo-bulks. Next, we generated similar bulk samples, but this time excluding cells from cluster 8 in the sampling process. On these samples we again calculated gene-by-gene correlation coefficients for C8 marker genes and compared the distributions of pairwise correlation coefficients to the distributions in the pseudo-bulk containing cells from cluster 8.
GTEx bulk RNA-seq data for normal tissues were obtained as batch-corrected, log-normalized counts from the UCSC Xenabrowser82. Pairwise correlations were visualized with the corrplot (https://cran.r-project.org/web/packages/corrplot/) package. For cross-tissue correlation analyses, only tissues with a median DPT expression >7.5 were considered. In this analysis, the top 20 marker genes for cluster 8 of the single-cell dataset ordered by log(fold change), which were found in fewer than 15% of other cells, were used. For deconvolution of microdissected PDAC stromal samples, raw expression counts per sample (n = 122) were downloaded from GEO (GSE93326). Data were normalized to log2(CPM + 1). Scores for cluster 8- and cluster 3-derived expression signatures (described above) in these bulk samples were calculated based on the average expression of the 20 most upregulated genes from the respective single-cell cluster (ordered by log(fold change), only genes expressed in at most 30% of other cells were considered).
Pseudo-bulk analytical strategy
We first generated 100 pseudo-bulk RNA-seq profiles from our single-cell dataset with varying numbers of cells from individual single-cell clusters (Extended Data Fig. 9d, top). We observed that the expression of marker genes for fibroblast cluster 8 co-varied depending on the number of cells from cluster 8 in the bulks. As a consequence, their expression profiles were strongly correlated, but only if cells from cluster 8 were added to the pseudo-bulk. Leaving cells from cluster 8 out resulted in an extensive drop in gene-wise correlations to close to 0. Therefore, co-expression of a single-cell-derived marker gene set can be used to infer the presence or absence of a particular cell population in bulk RNA-seq.
Projection of human gene sets onto mouse perturbed-state atlas
Gene expression signatures from human scRNA-seq datasets (Supplementary Table 7) corresponding to different fibroblast types were scored on the perturbed state atlas using the Seurat function AddModuleScore. The density of cells with the highest activation score (top 25th percentile for all clusters except in the Comp cluster (where we visualized the top 5th percentile)) was visualized using the function LSD::Heatscatter.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Raw and processed RNA-seq, ATAC–seq and mouse healthy skin scRNA-seq datasets are available from the ArrayExpress repository under the accession numbers E-MTAB-10324, E-MTAB-10316 and E-MTAB-10315, respectively. Supplementary Table 3 lists the studies used to generate the Cd45− maps and fibroblast atlases. These integrated scRNA-seq objects used for analysis are provided in an online resource that can be accessed at https://fibroXplorer.com. Human pancreatic cancer single-cell data are available in the EGA database under accession EGAD00001005365. Source data are provided with this paper.
Code availability
No new algorithms were developed for this manuscript. All code generated for analysis is available from the authors upon request.
References
Koliaraki, V., Prados, A., Armaka, M. & Kollias, G. The mesenchymal context in inflammation, immunity and cancer. Nat. Immunol. 21, 974–982 (2020).
Hinz, B. & Lagares, D. Evasion of apoptosis by myofibroblasts: a hallmark of fibrotic diseases. Nat. Rev. Rheumatol. 16, 11–31 (2020).
Buechler, M. B. & Turley, S. J. A short field guide to fibroblast function in immunity. Semin. Immunol. 35, 48–58 (2018).
Gautier, E. L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Croft, A. P. et al. Distinct fibroblast subsets drive inflammation and damage in arthritis. Nature 570, 246–251 (2019).
Wei, K. et al. Notch signalling drives synovial fibroblast identity and arthritis pathology. Nature 582, 259–264 (2020).
Zhang, F. et al. Defining inflammatory cell states in rheumatoid arthritis joint synovial tissues by integrating single-cell transcriptomics and mass cytometry. Nat. Immunol. 20, 928–942 (2019).
Dominguez, C. X. et al. Single-cell RNA sequencing reveals stromal evolution into LRRC15+ myofibroblasts as a determinant of patient response to cancer immunotherapy. Cancer Discov. 10, 232–253 (2020).
Elyada, E. et al. Cross-species single-cell analysis of pancreatic ductal adenocarcinoma reveals antigen-presenting cancer-associated fibroblasts. Cancer Discov. 9, 1102–1123 (2019).
Lambrechts, D. et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat. Med. 24, 1277–1289 (2018).
Öhlund, D. et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J. Exp. Med. 214, 579–596 (2017).
Rinkevich, Y. et al. Identification and isolation of a dermal lineage with intrinsic fibrogenic potential. Science 348, aaa2151 (2015).
Buechler, M. B. et al. A stromal niche defined by expression of the transcription factor WT1 mediates programming and homeostasis of cavity-resident macrophages. Immunity 51, 119–130.e5 (2019).
Krausgruber, T. et al. Structural cells are key regulators of organ-specific immune responses. Nature 583, 296–302 (2020).
Bellomo, A. et al. Reticular fibroblasts expressing the transcription factor WT1 define a stromal niche that maintains and replenishes splenic red pulp macrophages. Immunity 53, 127–142.e7 (2020).
Baryawno, N. et al. A cellular taxonomy of the bone marrow stroma in homeostasis and leukemia. Cell 177, 1915–1932.e16 (2019).
McCarthy, N. et al. Distinct mesenchymal cell populations generate the essential intestinal BMP signaling gradient. Cell Stem Cell 26, 391–402.e5 (2020).
Wirka, R. C. et al. Atheroprotective roles of smooth muscle cell phenotypic modulation and the TCF21 disease gene as revealed by single-cell analysis. Nat. Med. 25, 1280–1289 (2019).
Tsukui, T. et al. Collagen-producing lung cell atlas identifies multiple subsets with distinct localization and relevance to fibrosis. Nat. Commun. 11, 1920 (2020).
Dahlgren, M. W. & Molofsky, A. B. Adventitial cuffs: regional hubs for tissue immunity. Trends Immunol. 40, 877–887 (2019).
Sitnik, K. M. et al. Context-dependent development of lymphoid stroma from adult CD34+ adventitial progenitors. Cell Rep. 14, 2375–2388 (2016).
Merrick, D. et al. Identification of a mesenchymal progenitor cell hierarchy in adipose tissue. Science 364, eaav2501 (2019).
Zhou, X. et al. Circuit design features of a stable two-cell system. Cell 172, 744–757.e17 (2018).
Chai, Q. et al. Maturation of lymph node fibroblastic reticular cells from myofibroblastic precursors is critical for antiviral immunity. Immunity 38, 1013–1024 (2013).
Kapoor et al.
Maurer, C. et al. Experimental microdissection enables functional harmonisation of pancreatic cancer subtypes. Gut 68, 1034–1043 (2019).
Habermann, A. C. et al. Single-cell RNA sequencing reveals profibrotic roles of distinct epithelial and mesenchymal lineages in pulmonary fibrosis. Sci. Adv. 6, eaba1972 (2020).
Emblom-Callahan, M. C. et al. Genomic phenotype of non-cultured pulmonary fibroblasts in idiopathic pulmonary fibrosis. Genomics 96, 134–145 (2010).
Kinchen, J. et al. Structural remodeling of the human colonic mesenchyme in inflammatory bowel disease. Cell 175, 372–386.e17 (2018).
Cremasco, V. et al. B cell homeostasis and follicle confines are governed by fibroblastic reticular cells. Nat. Immunol. 15, 973–981 (2014).
Kuehn, M. R., Bradley, A., Robertson, E. J. & Evans, M. J. A potential animal model for Lesch-Nyhan syndrome through introduction of HPRT mutations into mice. Nature 326, 295–298 (1987).
Smithies, O., Gregg, R. G., Boggs, S. S., Koralewski, M. A. & Kucherlapati, R. S. Insertion of DNA sequences into the human chromosomal β-globin locus by homologous recombination. Nature 317, 230–234 (1985).
Thomas, K. R., Folger, K. R. & Capecchi, M. R. High frequency targeting of genes to specific sites in the mammalian genome. Cell 44, 419–428 (1986).
Newman, R. J., Roose-Girma, M. & Warming, S. Efficient conditional knockout targeting vector construction using co-selection BAC recombineering (CoSBR). Nucleic Acids Res. 43, e124 (2015).
Gertsenstein, M. et al. Efficient generation of germ line transmitting chimeras from C57BL/6N ES cells by aggregation with outbred host embryos. PLoS ONE 5, e11260 (2010).
Hughes, E. D. & Saunders, T. L. Advanced Protocols for Animal Transgenesis, An ISTT Manual 291–325 (Springer, 2011).
Zhang, L. et al. Single-cell analyses inform mechanisms of myeloid-targeted therapies in colon cancer. Cell 181, 442–459.e29 (2020).
Painter, M. W. et al. Transcriptomes of the B and T lineages compared by multiplatform microarray profiling. J. Immunol. 186, 3047–3057 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Korotkevich, G., Sukhov, V. & Sergushichev, A. Fast gene set enrichment analysis. Preprint at https://doi.org/10.1101/060012 (2021).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Li, Q., Brown, J. B., Huang, H. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ho, D. E., Imai, K., King, G. & Stuart, E. A. MatchIt: nonparametric preprocessing for parametric causal inference. J. Stat. Softw. 42, 1–28 (2011).
McLeay, R. C. & Bailey, T. L. Motif enrichment analysis: a unified framework and an evaluation on ChIP data. BMC Bioinformatics 11, 165 (2010).
Wang, S. et al. Target analysis by integration of transcriptome and ChIP–seq data with BETA. Nat. Protoc. 8, 2502–2515 (2013).
Soliman, H. et al. Pathogenic potential of Hic1-expressing cardiac stromal progenitors. Cell Stem Cell 26, 205–220.e8 (2020).
Dobie, R. et al. Single-cell transcriptomics uncovers zonation of function in the mesenchyme during liver fibrosis. Cell Rep. 29, 1832–1847.e8 (2019).
Xiong, X. et al. Landscape of intercellular crosstalk in healthy and NASH liver revealed by single-cell secretome gene analysis. Mol. Cell 75, 644–660.e5 (2019).
Dahlgren, M. W. et al. Adventitial stromal cells define group 2 innate lymphoid cell tissue niches. Immunity 50, 707–722.e6 (2019).
Peyser, R. et al. Defining the activated fibroblast population in lung fibrosis using single-cell sequencing. Am. J. Respir. Cell Mol. Biol. 61, 74–85 (2019).
Xie, T. et al. Single-cell deconvolution of fibroblast heterogeneity in mouse pulmonary fibrosis. Cell Rep. 22, 3625–3640 (2018).
Koga, S. et al. Peripheral PDGFRα+ gp38+ mesenchymal cells support the differentiation of fetal liver–derived ILC2. J. Exp. Med. 278, jem.20172310-23 (2018).
Dell’Orso, S. et al. Single cell analysis of adult mouse skeletal muscle stem cells in homeostatic and regenerative conditions. Development 146, dev181743 (2019).
Scott, R. W., Arostegui, M., Schweitzer, R., Rossi, F. M. V. & Underhill, T. M. Hic1 defines quiescent mesenchymal progenitor subpopulations with distinct functions and fates in skeletal muscle regeneration. Cell Stem Cell 25, 797–813.e9 (2019).
Buechler, M. B. & Turley, S. J. Neutrophils follow stromal omens to limit peritoneal inflammation. Immunity 52, 578–580 (2020).
Davidson, S. et al. Single-cell RNA sequencing reveals a dynamic stromal niche that supports tumor growth. Cell Rep. 31, 107628 (2020).
Cheng, H.-W. et al. Origin and differentiation trajectories of fibroblastic reticular cells in the splenic white pulp. Nat. Commun. 10, 1739 (2019).
Burl, R. B. et al. Deconstructing adipogenesis induced by β3-adrenergic receptor activation with single-cell expression profiling. Cell Metab. 28, 300–309.e4 (2018).
Schwalie, P. C. et al. A stromal cell population that inhibits adipogenesis in mammalian fat depots. Nature 559, 103–108 (2018).
Micheli, A. J. D. et al. Single-cell transcriptomics identify extensive heterogeneity in the cellular composition of mouse achilles tendons. Preprint at https://doi.org/10.1101/801266 (2020).
Hepler, C. et al. Identification of functionally distinct fibro-inflammatory and adipogenic stromal subpopulations in visceral adipose tissue of adult mice. eLife 7, 771 (2018).
Boyd, D. F. et al. Exuberant fibroblast activity compromises lung function via ADAMTS4. Nature 587, 466–471 (2020).
Perez-Shibayama, C. et al. Type I interferon signaling in fibroblastic reticular cells prevents exhaustive activation of antiviral CD8+ T cells. Sci. Immunol. 5, eabb7066 (2020).
Guerrero-Juarez, C. F. et al. Single-cell analysis reveals fibroblast heterogeneity and myeloid-derived adipocyte progenitors in murine skin wounds. Nat. Commun. 10, 650 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
van den Brink, S. C. et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat. Methods 14, 935–936 (2017).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Gao, X., Hu, D., Gogol, M. & Li, H. ClusterMap: compare multiple single cell RNA-seq datasets across different experimental conditions. Bioinformatics 35, 3038–3045 (2019).
Schubert, M. et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat. Commun. 9, 20 (2018).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Bharat, A. et al. Lung transplantation for patients with severe COVID-19. Sci. Transl. Med. 12, eabe4282 (2020).
Maynard, A. et al. Therapy-induced evolution of human lung cancer revealed by single-cell RNA sequencing. Cell 182, 1232–1251.e22 (2020).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Goldman, M. J. et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 38, 675–678 (2020).
Acknowledgements
We thank members of the Turley laboratory and individuals in the bioinformatics department for discussions; C. Dominguez for experimental assistance; S. Lianoglou, Y. Senbabaoglu and N. Lounsbury for bioinformatics assistance; facility staff at Genentech for vivarium maintenance and core facility assistance; and Turley laboratory members and A. Berkley for their careful review of this manuscript. This work was supported by Genentech.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.B.B., R.N.P., S.M., R.B. and S.J.T. Methodology: M.B.B., R.N.P., S.M., R.B. and S.J.T. Software, formal analysis and data curation: R.N.P., S.M., M.B.B. and A.K.C. Investigation: M.B.B., R.N.P., A.T.K., C.C., A.W.W., Y.A.Y., L.T., R.C. and A.K.C. Writing: M.B.B., R.N.P., S.M., R.B. and S.J.T. Visualization: M.B.B., R.N.P., A.T.K., C.C., S.M. Supervision: M.R.-G., Z.M., J.R.A., S.M., R.B. and S.J.T.
Corresponding authors
Ethics declarations
Competing interests
All authors except for R.B. are employees of Genentech. R.B. is an employee of Freenome.
Additional information
Peer review information Nature thanks Christopher Buckley, Neil Henderson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mouse fibroblast bulk RNA-seq and ATAC–seq.
a, Diagram of tissues isolated for bulk RNA-seq and ATAC–seq (adipose tissue (brown, subcutaneous, epididymal), bone, liver, lung, lymph node, mesentery, omentum and pancreas) and experimental scheme. b, Correlation plot of bulk RNA sequencing samples based on top 1,000 most differentially expressed genes. c, Fraction of ATAC–seq reads identified as PCR duplicates, mitochondrial DNA, or unique non-duplicate non-mitochondrial DNA based on genomic mapping. d, Fragment lengths of ATAC–seq samples. e, Number of called peaks across ATAC–seq samples. Atlas describes the universe of all peaks. f, Distribution of ATAC–seq peaks across genomic regions. g, Aggregate signal around the TSS. SLN_2 samples shown, representative of all other samples. h, Heatmap of pairwise correlation coefficients of ATAC–seq samples. Ing, subcutaneous adipose; Epi, epididymal adipose; Panc, pancreas; Om, omentum; SLN, lymph node.
Extended Data Fig. 2 Bulk gene expression and open chromatin.
a, Heat map depicting enriched genes per tissue by bulk RNA-seq. Adjusted P ≤ 0.05, log2(fold change) ≥ 2 (DESeq2, two-sided). Row z-scored. Top, bar plot depicting number of signature genes per tissue. b, Heat map depicting regions of open chromatin per tissue by bulk ATAC–seq. Adjusted P ≤ 0.01, log2(fold change) ≥ 2 (DESeq2, two-sided). Row z-scored. Top, bar plot depicting number of open chromatin regions per tissue. c, Correlation (top) and BETA analysis (bottom) of bulk RNA-seq and ATAC–seq samples. d, BETA analysis of SLN evaluating enriched gene expression compared to enriched SLN OCRs. These data are representative of the rest of the dataset. e, Number of transcription factor binding motifs in signature OCRs per tissue. f, Statistical inference of transcription factor motif enrichment in fibroblasts. Bar colours denote RNA expression of transcription factor. Subcut., subcutaneous adipose; LN, lymph node.
Extended Data Fig. 3 Steady-state fibroblast and Cd45− atlases.
a, UMAP embedding of 180,922 single cells in steady-state Cd45− atlas. Fourteen clusters identified through graph-based clustering are indicated by colour. b, Relative abundance of each tissue in steady-state Cd45− UMAP clusters. The sizes of bubbles indicate the contributions of cells from each tissue to a cluster, and grey bubbles indicate lack of contribution (number of cells <0.02% of tissue) of that tissue to the corresponding cluster. Graph to be read column-wise. c, Fibroblast- and other lineage-associated genes (in grey) in steady-state Cd45− atlas. The sizes of circles denote the percentages of cells from each cluster, and colour encodes the average expression across all cells within a cluster. The colour scale shows the expression level based on row z-score. d, Average bulk tissue-specific fibroblast gene signature scores across tissues represented in the steady-state atlas. Mean-centred values shown. e, Expression of cluster hallmark genes in steady-state fibroblast atlas. The sizes of circles denote the percentages of cells from each cluster, and colour encodes the average expression across all cells within a cluster. The colour scale shows the expression level based on row z-score. f, Expression of fibroblast- and other lineage-associated genes (in grey). Circle sizes denote percentages of cells from each cluster, and colour encodes average expression across all cells within cluster. Colour scale shows expression level (z-score by row). g, Relative abundance of each tissue in steady-state fibroblast UMAP clusters. The sizes of bubbles indicate the contributions of cells from each tissue to a cluster, and grey bubbles indicate lack of contribution (number of cells <0.02% of tissue) of that tissue to the corresponding cluster. Graph to be read column-wise. h, Expression of pathway-responsive genes in perturbed-state atlas clusters as assessed by PROGEN(y) analysis (z-scored per row). i, Volcano plot comparing genes enriched in Pi16+ cluster (left; blue) to Col15a1+ cluster (right; red). Select genes are labelled. Dots labelled in red or blue were determined as having a log2(fold change) of greater than 1 or less than −1 and an adjusted P value less than or equal to 0.05 (Bonferroni correction). j, Expression of Ly6a and Cd34 in steady-state clusters. Wilcoxon’s rank sum test P < 0.05.
Extended Data Fig. 4 Bulk sequencing validates single-cell atlas.
a–p, UMAPs highlighting distribution of cells from individual tissues in the steady-state fibroblast atlas. q, PCA of FACS-sorted bulk RNA-seq of fibroblasts and mesothelial cells, calculated for the 1,000 genes with the highest interquartile range. Circles represent fibroblasts and triangles are mesothelial cells. Each colour denotes a different tissue. r, Heatmap depicting top 20 Pi16+ (fibroblast) and Col15a1+ genes from steady-state fibroblast atlas in bulk RNA-seq data. Rows are z-scored. s, Gene enrichment analysis of top genes (log(fold change) > 0.5) from Pi16+ cluster and Col15a1+ cluster in loadings of PC1, which discriminates between mesothelial cells and fibroblasts (test: FGSEA, Benjamini–Hochberg correction). t, ATAC–seq traces of select genes from Pi16+ and Col15a1+ clusters genes, Ptprc (encoding CD45) and Hprt at ±2 kb of the TSS.
Extended Data Fig. 5 Supplementary FACS gating.
a, Representative FACS gating of live EpCAM−CD45−CD31−PDPN+PDGFRα+ fibroblasts in Figs. 2, 3, Extended Data Figs. 8, 9. b, c, Representative FACS gating on LY6C+SCA1+ (red), LY6C+SCA1− (blue) and LY6C−SCA1− (green) PDGFRα+ cells (as gated in Extended Data Fig. 7a) in lymph node of Ccl19YFP mice (b) and lymph node of Grem1creERT2Rosa26LSLYFP mice (c). Grem1creERT2Rosa26LSLYFP mice received tamoxifen via intraperitoneal injection for five consecutive days and were analysed on days 14–16 after first injection. d, Representative FACS gating on LY6C+SCA1+ (red), LY6C+SCA1− (blue) and LY6C−SCA1− (green) PDGFRα+ cells (as gated in Extended Data Fig. 7a) in tissues from DptIRESCreERT2Rosa26LSLYFP mice. Mice received tamoxifen chow for 14 days and were analysed on days 14–16. n = representative of 3 or more biologically independent experiments.
Extended Data Fig. 6 DptIRESCreERT2 marks universal fibroblasts.
a, Expression of Ly6c and Ly6a in the mouse steady-state fibroblast atlas. b, Representative gating for LY6C+SCA1+, LY6C−SCA1+ and SCA1− fibroblasts across indicated tissues. c, Quantification of LY6C+SCA1+, LY6C−SCA1+ and SCA1− fibroblasts across indicated tissues. Frequency (top), cell number per mg of tissue (bottom). d, Abundance of Pi16, Ccl19, Npnt, and Dpt transcripts in LY6C+SCA1+, LY6C−SCA1+ and SCA1− fibroblasts from lymph node (black) and lung (red). e, Expression of Pi16, Ccl19, Npnt and Dpt in mouse steady-state fibroblast atlas. f, RNAscope for Dpt (blue) and Pi16 (red) on lung (left) and small intestine (right). Scale bar, 50 μm. g, Schematic of generation of DptIRESCreERT2Rosa26LSLYFP mouse. YFP expression is achieved in Dpt-expressing cells following cre-mediated excision of a loxP-flanked transcriptional stop sequence. h, Representative gating for YFP (Dpt) in fibroblasts (live, EpCAM−CD45−CD31−PDPN+PDGFRα+) across indicated tissues. Red, shaded histogram is DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP and black is DptIRESCreERT2wt/wtRosa26LSLYFPwt/loxP. i, Representative gating for YFP (Dpt) in CD45+ cells (live, EpCAM−CD45+), EpCAM+ cells (live, EpCAM+), and endothelial cells (live, EpCAM−CD45−CD31+) from the pancreas. Red line is DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP and grey, shaded histogram is DptIRESCreERT2wt/wtRosa26LSLYFPwt/loxP. j, Quantification of YFP+ cells in DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP animals in indicated cell types across tissues. Percentage YFP+ was determined by subtracting fluorescence in DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP animals from DptIRESCreERT2wt/wtRosa26LSLYFPwt/loxP animals. k, Abundance of Pi16, Ccl19, Npnt, and Dpt transcripts in FACS-sorted YFP+ and YFP− cells from DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP and DptIRESCreERT2wt/kiRosa26LSLYFPwt/loxP animals 14–16 days after tamoxifen chow, in lymph node (top) and lung (bottom). l, Abundance of Csf1 transcripts in FACS-sorted YFP+ and YFP− cells from DptIRESCreERT2ki/kiRosa26LSLYFPwt/loxP and DptIRESCreERT2wt/kiRosa26LSLYFPwt/loxP animals 14–16 days after tamoxifen chow in lymph node and lung. n = 3 (c, d, k, l) or representative of 3 or more biologically independent experiments (b, f, h, i). Dot represents each mouse with bar at mean (c, j), plots show minimum and maximum (whiskers) and median (centre line). (d, k, l). h–l, Mice received tamoxifen chow for 14 days and were analysed on days 14–16.
Extended Data Fig. 7 Perturbed-state fibroblast and Cd45− atlases.
a, UMAP embedding of 182,609 cells in perturbed-state Cd45− atlas. Twelve clusters identified through graph-based clustering are indicated by colour. b, Relative abundance of each tissue in perturbed-state Cd45− UMAP clusters. The sizes of bubbles indicate the contributions of cells from each tissue to a cluster, and grey bubbles indicate lack of contribution (number of cells <0.02% of tissue) of that tissue to the corresponding cluster. Graph to be read column-wise. c, Fibroblast- and other lineage-associated genes (in grey) in Cd45− perturbed-state map. The sizes of circles denote the percentages of cells from each cluster, and colour encodes the average expression across all cells within a cluster. The colour scale shows the expression level based on row z-score. d, Expression of cluster hallmark genes in perturbed-state fibroblast atlas. The sizes of circles denote the percentages of cells from each cluster, and colour encodes the average expression across all cells within a cluster. The colour scale shows the expression level based on row z-score. e, Fibroblast- and other lineage-associated genes (in grey) in mouse perturbed-state fibroblast atlas. Circle sizes denote percentages of cells from each cluster; colour encodes average expression across all cells within cluster. Colour scale shows expression level (z-score by row). f, Relative abundance of each tissue in perturbed-state fibroblast UMAP clusters. The sizes of bubbles indicate the contributions of cells from each tissue to a cluster, and grey bubbles indicate lack of contribution (number of cells <0.02% of tissue) of that tissue to the corresponding cluster. Graph to be read column-wise. Clusters with analogues in steady state are depicted in grey, irrespective of the tissue contribution. g, Dendrogram representation of the hierarchical clustering of steady-state (blue) and perturbed-state (red) cluster marker genes. Similarity scores are indicated at the bottom of matching clusters from steady-state and perturbed-state tissues.
Extended Data Fig. 8 Validation of perturbed-state atlas.
a–o, UMAP representations of the distribution of fibroblasts across tissues and perturbations. p–u, Heat maps showing average relative gene expression in Pi16+, Col15a1+, Adamdec1+, Cxcl5+, and Lrrc15+ clusters (z-scored per row) in the following categories. p, Cytokines and chemokines. q, Wnt-associated genes. r, ECM-associated genes. s, Collagens and laminins. t, Matrix metalloproteases and cathepsins. u, Receptors and surface molecules. v, Expression of pathway-responsive genes in perturbed-state atlas clusters as assessed by PROGEN(y) analysis (z-scored per row). w, RNAscope for Dpt (blue) and Grem1 (red) in non-lesional colon (top) and lesional colon (bottom) on day 7 after induction of DSS colitis. Data are representative of three experiments. Scale bars, 50 μm (top) and 250 μm (bottom). x, Ly6a and Cd34 expression in perturbed-state clusters. Wilcoxon’s rank sum test, P < 0.05. y, Pseudotime(s) visualized using principal curves representing trajectories of fibroblast differentiation across perturbed-state fibroblast object. Blue lines show trajectory to activated clusters, grey lines show trajectory to clusters with a steady-state analogue. Pi16+ cluster set as root. z, Representative FACS strategy for subcutaneous tumour experiments. a′, Representative flow cytometry plots showing frequency of YFP+ cells in LRRC15+ fibroblasts from KPR3070 subcutaneous tumour at day 21 post-inoculation in DptIRESCreERT2wt/wtRosa26LSLYFPwt/loxP animals. b′, Quantification of FACS data (Fig. 3d, Extended Data Fig. 8z, a′). Data are from b′ or representative of 2 (z–a′) or 3 (w) experiments. Each dot represents one mouse (b′). n = 2 (b′) or representative of 2 (z–a′) or 3 (w) biologically independent experiments.
Extended Data Fig. 9 Human universal fibroblasts.
a, Top, UMAP embeddings of human pancreatic cancer tumour and normal adjacent tissue (n = 21,626 cells). Bottom, percentage of cells in each cluster coming from tumour or NAT. b, UMAP as in a, coloured by expression of indicated genes. c, Relative average expression of top 10 marker genes (sorted by log(fold change)) for each cluster in the pancreatic cancer single-cell dataset. Two representative genes highlighted per cluster. DEGs across clusters. d, Top, expression level of indicated marker genes (colour, y-axis) across 100 pseudo-bulk samples (x-axis) generated from human pancreatic cancer scRNA-seq data. The known percentage of cells from cluster 8 in each pseudo-bulk is shown by the dotted blue line. Bottom, boxplots representing the distributions of pairwise correlation coefficients of the top 20 marker genes for cluster 8 in pseudo-bulk samples containing (left) and not containing cells from cluster 8 (right). e, Boxplots summarizing DPT expression distributions across tissues from the GTEx portal. Tissues with mean above horizontal black line were included in correlation analysis (f). n = 7,851 biologically independent samples. f, Co-expression as in d, results from the gene-by-gene correlation matrices are summarized as boxplots for each individual tissue from GTEx. n = 5,957 biologically independent samples. g, Gene-by-gene correlation matrix of pairwise correlations in 205 normal pancreas bulk RNA-seq samples from GTEx. Blue indicates Pi16+ cluster signature gene, red indicates Col15a1+ signature gene. h, Human universal fibroblast score projected onto human pancreatic cancer samples. i, Human universal fibroblast score projected onto human subcutaneous adipose. d–f, Box and whisker plots show minimum and maximum (whiskers), interquartile range (box) and median (centre line).
Extended Data Fig. 10 Perturbed-state human fibroblasts.
a, Volcano plot visualizing log(fold change) (x-axis) and adjusted P value (y-axis) comparing fibroblasts from cluster 3 in Extended Data Fig. 9a compared to fibroblasts from cluster 8 (Wilcoxon’s rank sum test). b, Scatterplot visualizing the scores for a NAT (cluster 3) fibroblast expression gene set (y-axis) compared to scores for a CAF (cluster 8) expression gene set (x-axis) in 122 bulk RNA-seq samples of microdissected PDAC tissue. Each dot represents a sample, the regression line is given in blue. Error band, 95% confidence interval. c–f, UMAP representation of cells from the mouse perturbed-state atlas, each cell coloured by their score for gene sets corresponding to the following. c, Lrrc15+ myofibroblasts from human pancreatic cancer (cluster 3 CAF, ‘MyCAF’11), rheumatoid arthritis (‘Human RA F2’7), lung disease (‘Myofibroblast’29), IPF30 and ulcerative colitis (‘S2’31). d, Universal fibroblasts from human pancreatic cancer (cluster 8 NAT), lung disease (‘PLIN2+ fibroblast’29) and ulcerative colitis (‘S1’31). e, Inflammatory fibroblasts from pancreatic cancer (‘iCAF’11) and rheumatoid arthritis (‘Human RA F5’7). f, UMAP of IL-1 CAF cells derived from ref. 10, projected onto perturbed-state fibroblast atlas. g, Expression of human universal fibroblast module across human perturbed-state atlas clusters. h, Abundance of LRRC15 and CTHRC1 expression in human perturbed-state atlas. i, Heat maps showing average relative gene expression in human perturbed-state atlas (z-scored by row).
Supplementary information
Supplementary Table 1
Tissue-specific fibroblast and mesothelial cell genes by bulk RNAseq (related to Extended Data Fig. 2).
Supplementary Table 2
Tissue-specific OCRs and TFs by bulk ATACseq (related to Extended Data Fig. 2).
Supplementary Table 3
Tissue contributions to single cell RNAseq objects used in analyses of steady state and perturbed state murine tissues (related to Figs 1 and 3).
Supplementary Table 4
DEGs for steady-state mouse fibroblast atlas (related to Fig. 1).
Supplementary Table 5
DEGs for mouse perturbed-state fibroblast atlas (related to Fig. 3).
Supplementary Table 6
Patient information (related to Fig. 4 and Extended Data Figs 11 and 12).
Supplementary Table 7
DEGs for human pancreatic cancer samples (related to Fig. 4 and Extended Data Figs 11 and 12).
Supplementary Table 8
Human gene sets projected onto mouse perturbed-state fibroblast atlas (related to Extended Data Fig. 12).
Supplementary Table 9
DEGs for human perturbed-state atlas (related to Fig. 4).
Rights and permissions
About this article
Cite this article
Buechler, M.B., Pradhan, R.N., Krishnamurty, A.T. et al. Cross-tissue organization of the fibroblast lineage. Nature 593, 575–579 (2021). https://doi.org/10.1038/s41586-021-03549-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03549-5
This article is cited by
-
Evidence of steady-state fibroblast subtypes in the normal human breast as cells-of-origin for perturbed-state fibroblasts in breast cancer
Breast Cancer Research (2024)
-
Single-cell analysis of the cellular landscape of vulvar melanoma provides new insight for immunotherapy administration
BMC Cancer (2024)
-
Crosstalk with renal proximal tubule cells drives acidosis-induced inflammatory response and dedifferentiation of fibroblasts via p38-singaling
Cell Communication and Signaling (2024)
-
Agomir-122-loaded nanoparticles coated with cell membrane of activated fibroblasts to treat frozen shoulder based on homologous targeting
Journal of Nanobiotechnology (2024)
-
Fibroblast and myofibroblast activation in normal tissue repair and fibrosis
Nature Reviews Molecular Cell Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.