Abstract
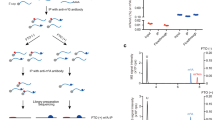
An extensive repertoire of modifications is known to underlie the versatile coding, structural and catalytic functions of RNA, but it remains largely uncharted territory. Although biochemical studies indicate that N6-methyladenosine (m6A) is the most prevalent internal modification in messenger RNA, an in-depth study of its distribution and functions has been impeded by a lack of robust analytical methods. Here we present the human and mouse m6A modification landscape in a transcriptome-wide manner, using a novel approach, m6A-seq, based on antibody-mediated capture and massively parallel sequencing. We identify over 12,000 m6A sites characterized by a typical consensus in the transcripts of more than 7,000 human genes. Sites preferentially appear in two distinct landmarks—around stop codons and within long internal exons—and are highly conserved between human and mouse. Although most sites are well preserved across normal and cancerous tissues and in response to various stimuli, a subset of stimulus-dependent, dynamically modulated sites is identified. Silencing the m6A methyltransferase significantly affects gene expression and alternative splicing patterns, resulting in modulation of the p53 (also known as TP53) signalling pathway and apoptosis. Our findings therefore suggest that RNA decoration by m6A has a fundamental role in regulation of gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Data have been deposited in NCBI’s Gene Expression Omnibus (GEO) and are accessible through GEO series accession number GSE37005 (http://ncbi.nlm.nih.gov/geo/query/acc.cgi?acc5GSE37005).
References
He, C. Grand challenge commentary: RNA epigenetics? Nature Chem. Biol. 6, 863–865 (2010)
Cantara, W. A. et al. The RNA Modification Database, RNAMDB: 2011 update. Nucleic Acids Res. 39, D195–D201 (2011)
Bokar, J. A. in Fine-Tuning of RNA Functions by Modification and Editing Vol. 12 (ed. Grosjean, H. ) 141–177 (Springer, 2005)
Bokar, J. A., Shambaugh, M. E., Polayes, D., Matera, A. G. & Rottman, F. M. Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3, 1233–1247 (1997)
Zhong, S. et al. MTA is an Arabidopsis messenger RNA adenosine methylase and interacts with a homolog of a sex-specific splicing factor. Plant Cell 20, 1278–1288 (2008)
Clancy, M. J., Shambaugh, M. E., Timpte, C. S. & Bokar, J. A. Induction of sporulation in Saccharomyces cerevisiae leads to the formation of N6-methyladenosine in mRNA: a potential mechanism for the activity of the IME4 gene. Nucleic Acids Res. 30, 4509–4518 (2002)
Hongay, C. F. & Orr-Weaver, T. L. Drosophila Inducer of MEiosis 4 (IME4) is required for Notch signaling during oogenesis. Proc. Natl Acad. Sci. USA 108, 14855–14860 (2011)
Desrosiers, R., Friderici, K. & Rottman, F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc. Natl Acad. Sci. USA 71, 3971–3975 (1974)
Horowitz, S., Horowitz, A., Nilsen, T. W., Munns, T. W. & Rottman, F. M. Mapping of N6-methyladenosine residues in bovine prolactin mRNA. Proc. Natl Acad. Sci. USA 81, 5667–5671 (1984)
Kane, S. E. & Beemon, K. Precise localization of m6A in Rous sarcoma virus RNA reveals clustering of methylation sites: implications for RNA processing. Mol. Cell. Biol. 5, 2298–2306 (1985)
Harper, J. E., Miceli, S. M., Roberts, R. J. & Manley, J. L. Sequence specificity of the human mRNA N6-adenosine methylase in vitro. Nucleic Acids Res. 18, 5735–5741 (1990)
Wei, C. M. & Moss, B. Nucleotide sequences at the N6-methyladenosine sites of HeLa cell messenger ribonucleic acid. Biochemistry 16, 1672–1676 (1977)
Dai, Q. et al. Identification of recognition residues for ligation-based detection and quantitation of pseudouridine and N6-methyladenosine. Nucleic Acids Res. 35, 6322–6329 (2007)
Levanon, E. Y. et al. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nature Biotechnol. 22, 1001–1005 (2004)
Perry, R. P. & Scherrer, K. The methylated constituents of globin mRNA. FEBS Lett. 57, 73–78 (1975)
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nature Chem. Biol. 7, 885–887 (2011)
Bringmann, P. & Luhrmann, R. Antibodies specific for N6-methyladenosine react with intact snRNPs U2 and U4/U6. FEBS Lett. 213, 309–315 (1987)
Dante, R. & Niveleau, A. Inhibition of in vitro translation by antibodies directed against N6-methyladenosine. FEBS Lett. 130, 153–157 (1981)
Munns, T. W., Liszewski, M. K., Oberst, R. J. & Sims, H. F. Antibody nucleic acid complexes. Immunospecific retention of N6-methyladenosine-containing transfer ribonucleic acid. Biochemistry 17, 2573–2578 (1978)
Munns, T. W., Liszewski, M. K. & Sims, H. F. Characterization of antibodies specific for N6-methyladenosine and for 7-methylguanosine. Biochemistry 16, 2163–2168 (1977)
Munns, T. W., Oberst, R. J., Sims, H. F. & Liszewski, M. K. Antibody-nucleic acid complexes. Immunospecific recognition of 7-methylguanine- and N6-methyladenine-containing 5′-terminal oligonucleotides of mRNA. J. Biol. Chem. 254, 4327–4330 (1979)
Munns, T. W., Sims, H. F. & Liszewski, M. K. Immunospecific retention of oligonucleotides possessing N6-methyladenosine and 7-methylguanosine. J. Biol. Chem. 252, 3102–3104 (1977)
Czerwoniec, A. et al. MODOMICS: a database of RNA modification pathways. 2008 update. Nucleic Acids Res. 37, D118–D121 (2009)
Perry, R. P., Kelley, D. E., Friderici, K. & Rottman, F. The methylated constituents of L cell messenger RNA: evidence for an unusual cluster at the 5′ terminus. Cell 4, 387–394 (1975)
Keith, J. M., Ensinger, M. J. & Mose, B. HeLa cell RNA (2′-O-methyladenosine-N6-)-methyltransferase specific for the capped 5′-end of messenger RNA. J. Biol. Chem. 253, 5033–5039 (1978)
Wei, C., Gershowitz, A. & Moss, B. N6, O2′-dimethyladenosine a novel methylated ribonucleoside next to the 5′ terminal of animal cell and virus mRNAs. Nature 257, 251–253 (1975)
Zilberman, D., Gehring, M., Tran, R. K., Ballinger, T. & Henikoff, S. Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nature Genet. 39, 61–69 (2007)
Chan, C. T. et al. A quantitative systems approach reveals dynamic control of tRNA modifications during cellular stress. PLoS Genet. 6, e1001247 (2010)
Schaefer, M. et al. RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev. 24, 1590–1595 (2010)
Lenos, K. & Jochemsen, A. G. Functions of MDMX in the modulation of the p53-response. J. Biomed. Biotechnol. 2011, 876173 (2011)
Klose, R. J. & Bird, A. P. Genomic DNA methylation: the mark and its mediators. Trends Biochem. Sci. 31, 89–97 (2006)
Mokrejs, M. et al. IRESite—a tool for the examination of viral and cellular internal ribosome entry sites. Nucleic Acids Res. 38, D131–D136 (2009)
Kislauskis, E. H., Zhu, X. & Singer, R. H. Sequences responsible for intracellular localization of β-actin messenger RNA also affect cell phenotype. J. Cell Biol. 127, 441–451 (1994)
Bernstein, P. L., Herrick, D. J., Prokipcak, R. D. & Ross, J. Control of c-myc mRNA half-life in vitro by a protein capable of binding to a coding region stability determinant. Genes Dev. 6, 642–654 (1992)
Zhang, Z. et al. The YTH domain is a novel RNA binding domain. J. Biol. Chem. 285, 14701–14710 (2010)
Harigaya, Y. et al. Selective elimination of messenger RNA prevents an incidence of untimely meiosis. Nature 442, 45–50 (2006)
Rafalska, I. et al. The intranuclear localization and function of YT521-B is regulated by tyrosine phosphorylation. Hum. Mol. Genet. 13, 1535–1549 (2004)
Brennan, C. M. & Steitz, J. A. HuR and mRNA stability. Cell. Mol. Life Sci. 58, 266–277 (2001)
Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 79, 321–349 (2010)
Keren, H., Lev-Maor, G. & Ast, G. Alternative splicing and evolution: diversification, exon definition and function. Nature Rev. Genet. 11, 345–355 (2010)
Sterner, D. A., Carlo, T. & Berget, S. M. Architectural limits on split genes. Proc. Natl Acad. Sci. USA 93, 15081–15085 (1996)
Camper, S. A., Albers, R. J., Coward, J. K. & Rottman, F. M. Effect of undermethylation on mRNA cytoplasmic appearance and half-life. Mol. Cell. Biol. 4, 538–543 (1984)
Carroll, S. M., Narayan, P. & Rottman, F. M. N6-methyladenosine residues in an intron-specific region of prolactin pre-mRNA. Mol. Cell. Biol. 10, 4456–4465 (1990)
Stoltzfus, C. M. & Dane, R. W. Accumulation of spliced avian retrovirus mRNA is inhibited in S-adenosylmethionine-depleted chicken embryo fibroblasts. J. Virol. 42, 918–931 (1982)
Barash, Y. et al. Deciphering the splicing code. Nature 465, 53–59 (2010)
Tuck, M. T., Wiehl, P. E. & Pan, T. Inhibition of 6-methyladenine formation decreases the translation efficiency of dihydrofolate reductase transcripts. Int. J. Biochem. Cell Biol. 31, 837–851 (1999)
Hamburger, A. W. & Pinnamaneni, G. Interferon-induced enhancement of transforming growth factor-α expression in a human breast cancer cell line. Proc. Soc. Exp. Biol. Med. 202, 64–68 (1993)
Mujoo, K., Donato, N. J., Lapushin, R., Rosenblum, M. G. & Murray, J. L. Tumor necrosis factor α and γ-interferon enhancement of anti-epidermal growth factor receptor monoclonal antibody binding to human melanoma cells. J. Immunother. Emphasis Tumor Immunol. 13, 166–174 (1993)
Hsu, F. et al. The UCSC Known Genes. Bioinformatics 22, 1036–1046 (2006)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Kinsella, R. J. et al. Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database 2011, bar030 (2011)
Llorian, M. et al. Position-dependent alternative splicing activity revealed by global profiling of alternative splicing events regulated by PTB. Nature Struct. Mol. Biol. 17, 1114–1123 (2010)
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. ISMB 2, 28–36 (1994)
Zuker, M. & Stiegler, P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 9, 133–148 (1981)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Anders, S. HTSeq: analysing high-throughput sequencing data with Python http://www-huber.embl.de/users/anders/HTSeq/doc/overview.html (2010)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Anders, S., Reyes, A. & Huber, W. Detecting differential usage of exons from RNA-Seq data. Available from Nature Precedingshttp://hdl.handle.net/10101/npre.2012.6837.1 (2012)
Roberts, A., Pimentel, H., Trapnell, C. & Pachter, L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 27, 2325–2329 (2011)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Salmon-Divon, M., Dvinge, H., Tammoja, K. & Bertone, P. PeakAnalyzer: genome-wide annotation of chromatin binding and modification loci. BMC Bioinformatics 11, 415 (2010)
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009)
Bembom, O. seqLogo: Sequence Logos for DNA Sequence Alignments (Division of Biostatistics, University of California, Berkeley, 2011)
Acknowledgements
We thank H. Cedar (The Hebrew University, Jerusalem) for his comments. We thank the Kahn Family Foundation for their support. This work was supported in part by grants from the Flight Attendant Medical Research Institute (FAMRI), Bio-Med Morasha ISF (grant no. 1942/08), and The Israel Ministry for Science and Technology (Scientific Infrastructure Program). R.S. was supported by the ERC-StG program (grant 260432). G.R. holds the Djerassi Chair in Oncology at the Sackler Faculty of Medicine, Tel Aviv University. This work was performed in partial fulfilment of the requirements for a PhD degree to D.D., Sackler Faculty of Medicine, Tel Aviv University.
Author information
Authors and Affiliations
Contributions
D.D., S.M.-M. and G.R. conceived and designed the experiments; D.D., S.M.-M., L.U., K.C., S.O. and J.J.-H. performed the experiments; S.S., M.S.-D. and R.S. performed the bioinformatic analysis; D.D., S.M.-M., M.S.-D., N.A., M.K., S.S., R.S. and G.R. analysed and interpreted results, and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-18, Supplementary References, Supplementary Tables 1-5, Supplementary Notes and legends for Supplementary Tables 6-8. (PDF 5060 kb)
Supplementary Table 6
This file contains Supplementary Table 6 in USCS format. (TXT 2738 kb)
Supplementary Table 6
This file contains Supplementary Table 6, which shows the dataset of all identified m6A peaks in HepG2 cell line and normal human brain. (XLS 21053 kb)
Supplementary Table 7
This file contains Supplementary Table 7 in UCSC format. (TXT 224 kb)
Supplementary Table 7
This file contains Supplementary Table 7, which shows the dataset of all identified m6A peaks in mouse liver. (XLS 8054 kb)
Supplementary Table 8
This file contains Supplementary Table 8, which shows differential m6A peaks between various experimental conditions. (XLS 242 kb)
Rights and permissions
About this article
Cite this article
Dominissini, D., Moshitch-Moshkovitz, S., Schwartz, S. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012). https://doi.org/10.1038/nature11112
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11112
This article is cited by
-
The role of the methyltransferase METTL3 in prostate cancer: a potential therapeutic target
BMC Cancer (2024)
-
N6-methyladenosine levels in peripheral blood RNA: a potential diagnostic biomarker for colorectal cancer
Cancer Cell International (2024)
-
m6A-TCPred: a web server to predict tissue-conserved human m6A sites using machine learning approach
BMC Bioinformatics (2024)
-
Lactylation-driven FTO targets CDK2 to aggravate microvascular anomalies in diabetic retinopathy
EMBO Molecular Medicine (2024)
-
Astrocytic ALKBH5 in stress response contributes to depressive-like behaviors in mice
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.